Week 9 HW: Cell Free System
PART 1 — General Homework Questions
1. What are the main advantages of cell-free protein synthesis over in vivo methods, particularly in terms of flexibility and control of experimental variables? Provide at least two cases where CFPS is more beneficial.
Cell-free protein synthesis (CFPS) offers significantly greater flexibility compared to in vivo systems because it is not constrained by cellular viability. In this system, biological components are extracted from cells, allowing researchers to directly manipulate reaction conditions such as ion concentrations (Mg²⁺, K⁺), energy molecules (ATP), and enzymatic composition. This enables precise experimental control without interference from complex cellular regulatory networks. Furthermore, CFPS operates as an open system, meaning components can be added or removed dynamically during the reaction, enhancing experimental tunability. CFPS is particularly advantageous in several scenarios, including:
• The production of toxic proteins (e.g., pore-forming proteins such as α-hemolysin), which would otherwise kill host cells.
• Rapid prototyping of genetic circuits in synthetic biology, as it eliminates the need for transformation and cell culture, thereby accelerating the design–build–test cycle.
2. What are the main components of a cell-free expression system and what are their respective roles?
A cell-free expression system consists of several essential components that collectively enable transcription and translation outside of living cells. The core component is the cell extract (cell lysate), typically derived from organisms such as E. coli, which contains ribosomes, transfer RNA (tRNA), transcription and translation enzymes, and other necessary factors. Additionally, a genetic template in the form of DNA or mRNA is required to encode the target protein. An energy system, including ATP and GTP, is crucial to drive biosynthetic reactions, often coupled with an energy regeneration mechanism to sustain the reaction over time. Other important components include:
• Amino acids, which serve as building blocks for protein synthesis
• Ions (Mg²⁺, K⁺), which stabilize ribosomal structure and enzymatic activity
• Cofactors such as NAD⁺ and CoA, which support auxiliary metabolic reactions
3. Why is energy regeneration critical in cell-free systems, and how can continuous ATP supply be maintained?
Energy regeneration is critical in cell-free systems because protein translation is highly energy-intensive, requiring approximately four ATP molecules per peptide bond formation. Without an efficient regeneration system, ATP is rapidly depleted, leading to premature termination of protein synthesis. Therefore, maintaining a continuous ATP supply is essential for sustaining reaction efficiency and increasing protein yield. One commonly used method is the phosphoenolpyruvate (PEP)-based system, where PEP is converted into pyruvate while generating ATP. Alternative approaches that offer improved stability include:
• Glucose-based energy systems that utilize metabolic pathways for ATP regeneration
• Creatine phosphate systems coupled with creatine kinase These strategies help prolong reaction duration and enhance overall productivity.
4. Compare prokaryotic and eukaryotic cell-free systems. Provide one protein example for each and justify your choice.
Prokaryotic cell-free systems, such as those derived from E. coli, are known for their high speed, efficiency, and relatively low cost. However, they are limited in their ability to perform post-translational modifications such as glycosylation or complex disulfide bond formation. In contrast, eukaryotic systems, including wheat germ or mammalian extracts, are capable of more accurate protein folding and post-translational modifications, albeit at higher cost and lower production speed. For example:
• Green Fluorescent Protein (GFP) is well-suited for prokaryotic systems because it does not require complex modifications.
• Antibodies (e.g., IgG) are better produced in eukaryotic systems due to their requirement for proper folding, disulfide bond formation, and glycosylation.
5. How would you design a cell-free experiment to optimize membrane protein expression? What challenges are involved and how can they be addressed?
Membrane protein expression in cell-free systems presents significant challenges because these proteins require a lipid environment to maintain proper structure and function. In the absence of such an environment, proteins are prone to misfolding and aggregation. Therefore, experimental design must incorporate strategies to mimic biological membranes. Possible solutions include:
• Incorporating liposomes or nanodiscs to simulate membrane environments
• Using mild detergents to maintain protein solubility
• Enabling co-translational insertion, allowing proteins to integrate into membranes during synthesis
• Adding molecular chaperones such as GroEL/ES to assist in protein folding
These approaches can significantly improve protein stability and yield.
6. If low protein yield is observed, what are the possible causes and how would you troubleshoot them?
Low protein yield in cell-free systems can arise from several factors. One common cause is the degradation of DNA or mRNA by nucleases present in the extract. This issue can be mitigated by using more stable DNA templates or adding nuclease inhibitors. Another cause is insufficient energy supply due to ATP depletion, which can be addressed by optimizing the energy regeneration system, such as using PEP or glucose-based systems. Additionally, improper protein folding may result in aggregation and reduced yield. This can be improved by adding chaperones or optimizing reaction conditions such as temperature and ionic strength.
PART 2 — Synthetic Minimal Cell Design
7. Design a synthetic minimal cell: what is its function and how does it work?
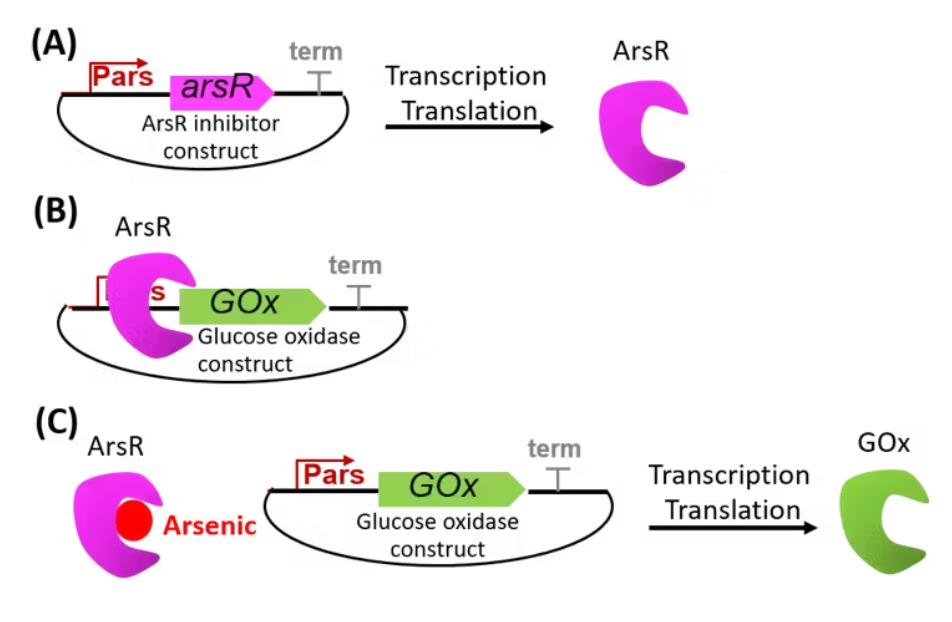
The proposed synthetic minimal cell functions as a biosensor for detecting arsenic contamination in water. The system operates by sensing arsenic ions (As³⁺) through a regulatory protein such as ArsR. In the presence of arsenic, ArsR dissociates from the DNA, allowing transcription of a reporter gene such as GFP. The output is a measurable fluorescent signal that indicates the presence of arsenic in the environment.
8. Can this function be achieved without encapsulation or using natural cells?
This function can technically be achieved using a non-encapsulated cell-free system; however, it would lack the compartmentalization characteristic of a synthetic cell. It can also be implemented using genetically engineered natural cells such as E. coli, but this approach introduces limitations related to biosafety, regulatory complexity, and reduced modularity compared to synthetic vesicle-based systems.
9. What are the main components of the synthetic cell?
The synthetic cell consists of a lipid membrane composed of phospholipids such as POPC combined with cholesterol to enhance stability. Inside the vesicle, an E. coli-based cell-free system is encapsulated along with DNA encoding the arsR regulatory gene and the gfp reporter gene. This setup enables arsenic detection and signal generation within a controlled microenvironment.
10. How does the system communicate with the environment and how is the output measured?
Communication with the environment occurs through the diffusion of arsenic ions across the lipid membrane, as small molecules can permeate the vesicle. Upon detection, GFP is produced and can be measured using fluorescence spectroscopy or flow cytometry. This output provides a quantitative indication of arsenic concentration.

PART 3 — Freeze-Dried CFPS Application (Textiles)
11. What is the proposed application of cell-free systems in textiles?
The proposed concept involves smart textiles embedded with freeze-dried cell-free systems capable of detecting air pollutants. Upon exposure to environmental moisture, the system becomes activated and responds to pollutants such as nitrogen dioxide or heavy metals by producing a visible color change. This enables real-time monitoring of air quality through wearable materials.
12. How can the limitations of cell-free systems be addressed in this application?
Key limitations of cell-free systems include stability, activation requirements, and single-use constraints. These challenges can be addressed through:
• Enhancing stability via freeze-drying with protective agents such as trehalose
• Enabling activation through environmental moisture
• Designing modular and replaceable patches to mitigate single-use limitations
PART 4 — Genes in Space Proposal
13. What is the background and objective of this study?
Cosmic radiation in space poses a significant threat to DNA integrity and astronaut health, particularly through the induction of double-strand breaks. Understanding DNA repair mechanisms under microgravity conditions is essential for long-term space exploration and biotechnological advancements. This study aims to investigate the activity of the RecA protein in DNA repair using a cell-free system.
14. What is the molecular target and hypothesis?
The molecular target of this study is the recA gene and radiation-damaged DNA. The hypothesis is that microgravity and radiation conditions reduce the efficiency of RecA-mediated DNA repair. This effect is expected to be reflected in decreased fluorescence signals from a DNA repair reporter system.
15. What is the experimental plan?
The experiment will utilize the BioBits® cell-free protein expression system to produce RecA protein in vitro. Samples will include undamaged DNA as a control and DNA damaged by UV or radiation. DNA repair activity will be measured using a fluorescence reporter, with instruments such as the miniPCR thermal cycler and P51 Molecular Fluorescence Viewer. The resulting data will provide insights into DNA repair efficiency under space conditions.