Week 11 HW: Building genomes
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork

1. Contribution
Honestly, I did not contribute anything to the canvas, as I was already too late to participate in the artwork. However, that is okay. I feel that I still have a lot to learn about this kind of project, and I was genuinely excited to see how enthusiastic everyone was while working on it.
2. What I liked about the project
This project provided us with the freedom to create unique and creative artwork. I was also truly impressed by how beautiful artistic outcomes can emerge from science, which broadened my perspective that science is not only intellectually engaging but also aesthetically inspiring. The global collaboration allowed me to connect with people from diverse backgrounds, and I learned a lot from that experience.
3. What could be improved
I hope that access to editing cell-free reaction compositions could still be maintained, but with some modifications. For example, it would be better if participants could create and explore their own versions without altering the collaborative work of others. Overall, however, the project was conducted very well.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
A. Functional Roles of Each Component
- Lysate
E. coli Lysate: The E. coli lysate provides the essential cellular machinery required for protein synthesis, including ribosomes, tRNAs, aminoacyl-tRNA synthetases, and various transcription and translation factors.
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase): This lysate contains T7 RNA polymerase, enabling efficient transcription of DNA templates into mRNA under the control of a T7 promoter.
- Salts and Buffer
Potassium Glutamate: Maintains ionic strength and mimics intracellular conditions, supporting optimal enzyme activity and protein stability.
HEPES-KOH (pH 7.5): Acts as a buffering agent to maintain a stable pH environment during the reaction.
Magnesium Glutamate: Provides magnesium ions, which are essential cofactors for ribosomal function and enzymatic activities in transcription and translation.
Potassium Phosphate Monobasic and Dibasic: Together form a phosphate buffer system that helps stabilize pH and supports metabolic reactions.
- Energy / Nucleotide System
Ribose: Serves as a precursor for nucleotide biosynthesis, enabling in situ generation of nucleotides.
Glucose: Functions as an energy source to regenerate ATP through metabolic pathways present in the lysate.
AMP, CMP, GMP, UMP: These nucleoside monophosphates act as precursors that are enzymatically converted into nucleoside triphosphates (NTPs) required for RNA synthesis.
Guanine: Serves as a substrate in the nucleotide salvage pathway, allowing the synthesis of GMP and subsequently GTP.
- Translation Mix (Amino Acids)
17 Amino Acid Mix: Provides the majority of amino acids necessary for protein synthesis.
Tyrosine: Supplied separately due to its specific solubility and stability requirements.
Cysteine: Added individually because it is prone to oxidation and requires controlled conditions.
- Additives
Nicotinamide: Acts as a precursor for NAD⁺, supporting redox reactions and cellular metabolism within the lysate.
- Backfill
Nuclease-Free Water: Used to adjust the final reaction volume without introducing nucleases that could degrade DNA or RNA.
B. Differences Between the 1-Hour and 20-Hour Systems
The 1-hour PEP-NTP system relies on the direct addition of high-energy molecules such as phosphoenolpyruvate (PEP) and nucleoside triphosphates (NTPs), enabling rapid protein synthesis but limiting reaction duration. In contrast, the 20-hour NMP-Ribose-Glucose system utilizes metabolic precursors like ribose and glucose to regenerate nucleotides and energy over time, resulting in a more sustainable and cost-effective system. This allows for prolonged protein production and improved reaction longevity.
C. Bonus Question
Transcription can still occur because guanine can be converted into GMP via the nucleotide salvage pathway. The GMP is then phosphorylated into GTP, which is required as a substrate for RNA synthesis during transcription.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
A. Biophysical and Functional Properties
- sfGFP (superfolder GFP)
sfGFP exhibits highly efficient folding and structural stability, allowing it to maintain strong fluorescence even under suboptimal conditions in cell-free systems.
- mRFP1
mRFP1 has relatively slow chromophore maturation, which can delay fluorescence development during shorter or suboptimal incubation periods.
- mKO2
mKO2 is sensitive to pH changes, and variations in pH can significantly affect its fluorescence intensity and stability.
- mTurquoise2
mTurquoise2 has a high quantum yield but requires proper folding conditions to achieve optimal fluorescence output.
- mScarlet-I
mScarlet-I displays rapid maturation and high brightness, making it well-suited for sustained fluorescence over long incubation periods.
- Electra2
Electra2 likely depends on sufficient oxygen availability for chromophore formation, which can influence fluorescence efficiency in cell-free reactions.
B. Hypothesis for Master Mix Optimization
To improve fluorescence of mRFP1, increasing magnesium ion concentration and optimizing energy sources such as glucose in the master mix may enhance protein synthesis and folding efficiency. This is expected to accelerate chromophore maturation and increase overall fluorescence intensity over a 36-hour incubation.
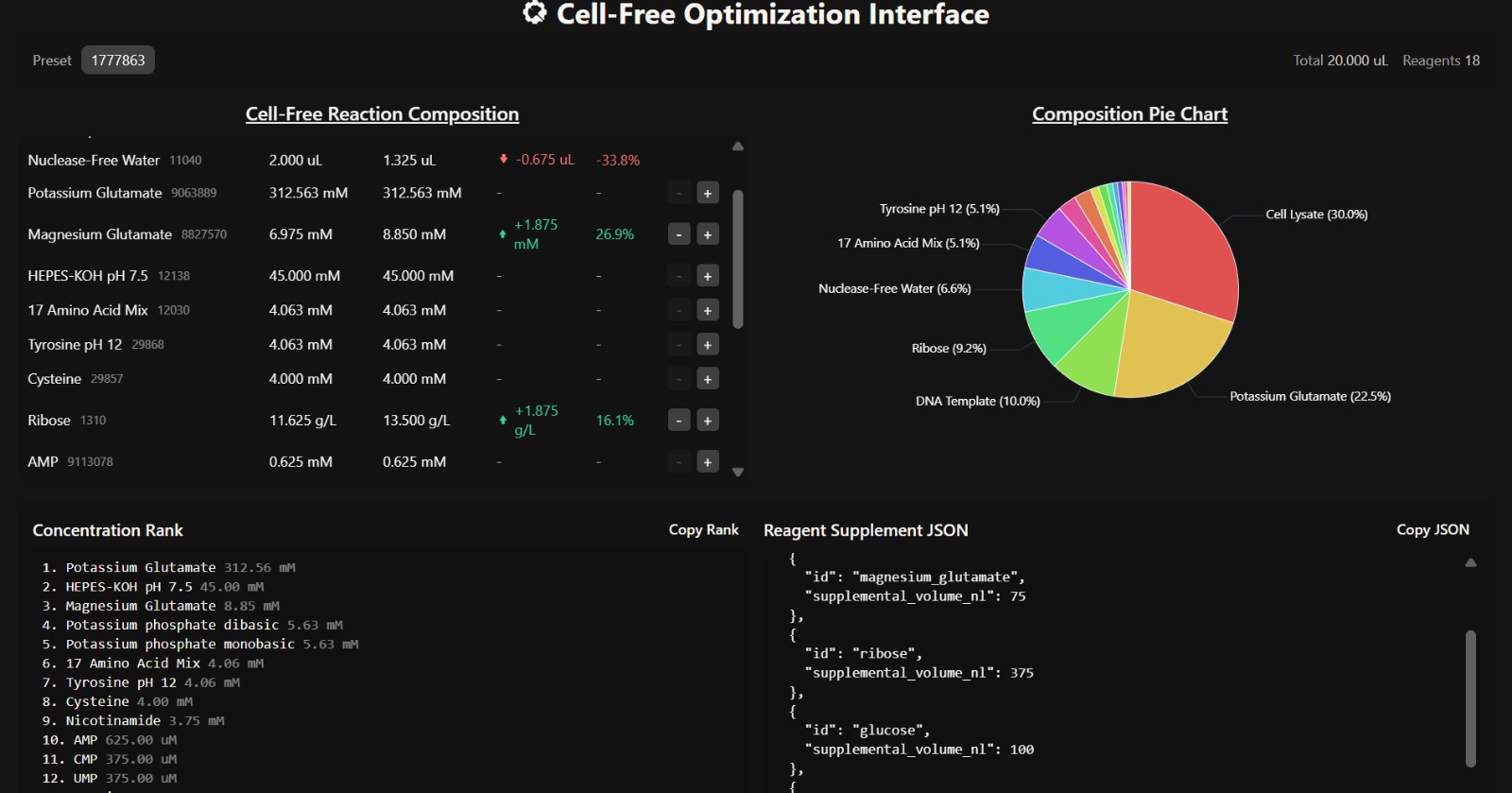

C. Interpretation of Cell-Free Reaction Composition
- Overall Composition Insight
The composition is dominated by cell lysate (30%) and potassium glutamate (22.5%), indicating that the system strongly mimics intracellular conditions and provides sufficient enzymatic machinery for transcription and translation. The presence of ribose (9.2%) and glucose supports a metabolically sustained system, enabling prolonged protein synthesis over a 36-hour incubation.
- Translation and Folding Efficiency
The increased magnesium glutamate concentration (8.85 mM) suggests enhanced ribosomal activity and improved translation efficiency. Magnesium ions are critical cofactors for ribosome stability and enzymatic reactions, meaning this adjustment likely promotes better protein folding and higher protein yield.
- Energy Sustainability
The elevated glucose (2.25 g/L) and ribose (13.5 g/L) concentrations indicate a system optimized for long-term energy regeneration and nucleotide synthesis. This supports continuous ATP production and RNA synthesis, which are essential for sustained protein expression.
- Metabolic Support
The presence of nicotinamide (3.75 mM) enhances metabolic stability by supporting NAD⁺ regeneration, which is essential for redox balance and energy metabolism within the lysate.
- Limitation in Supplement Strategy
Although key reagents were added, the supplemental volumes are relatively low, and a significant portion (1.325 µL) is still allocated to nuclease-free water. This may dilute the effect of critical reagents such as magnesium and glucose, potentially limiting the overall improvement in fluorescence.
- Expected Outcome
This composition is expected to produce moderate to high fluorescence output, particularly for proteins that benefit from sustained energy supply and improved folding conditions (e.g., mRFP1). However, the relatively small increase in key supplements may prevent the system from reaching maximum fluorescence potential.