Group Final Project
Hypothesis: Substitution of a bacteriophage’s replisome with an orthogonal T7 replisome for continuous hypermutation directed towards stability
Group Members:
Alan Bravo https://pages.htgaa.org/2026a-alan-bravo
Samudera Mukhalid https://pages.htgaa.org/2026a-samudera-mukhalid
Bacteriophage Engineering
Challange Choosen: Stability
Proposal
This proposal is inspired by the article by Diercks et al. (2024), which harnesses a highly error-prone replisome to drive extreme antibiotic resistance evolution. Primary Goal #1: Enhance bacteriophage resilience to varying environmental conditions, such as temperature and pH. Primary Goal #2: Achieve this stability without impairing the bacteriophage’s infectivity.
The proposal centers on engineering a bacteriophage based on the T7 replisome, as T7 encodes its own low-fidelity replication machinery. This enables the generation of a mutation-prone phage population, selectively evolved for excellence in the target trait: stability. The population is exposed to targeted stability-related stresses, followed by isolation of infectious survivors via plaque assays. Independent survivor clones are then sequenced, and their encoded proteins analyzed using Clustal Omega. This yields a consensus view of conserved residues versus sites recurrently altered under selection, enabling the design of an optimally stable phage construct.
Two key potential pitfalls:
• Clustal Omega results may be overly divergent, complicating the identification of conserved residues.
• No stable phages may emerge due to the replisome’s excessive faultiness.
Schematic
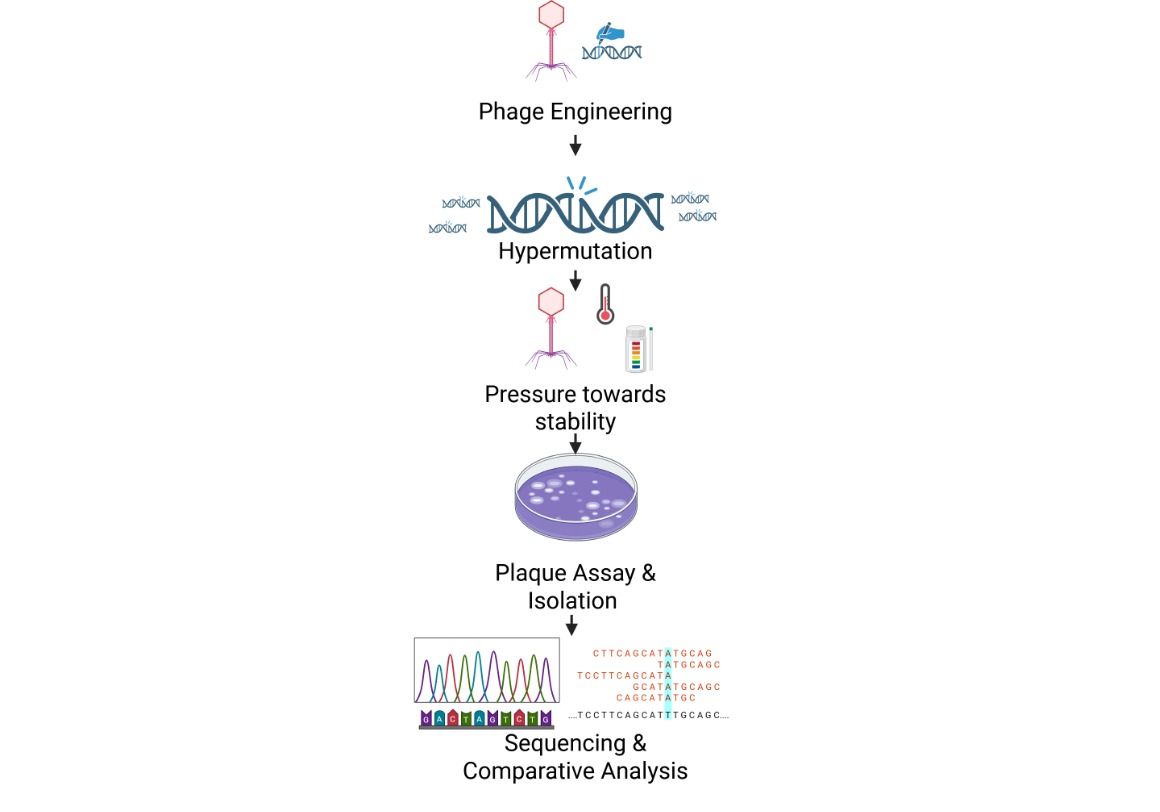

MSA result
Below are the results of the multiple sequence alignment we performed on a set of protein sequences. The rationale for selecting these sequences was to identify which residues in the bacteriophage endolysin are truly conserved, and which ones exhibit sufficient flexibility for engineering enhanced stability without compromising function. To achieve this, we avoided sequences that were either too identical or excessively divergent. Thus, the chosen sequences were deliberately selected within an intermediate homology range:
• Similar enough → Retain functional homology
• Different enough → Reveal clear patterns of conservation and variation"

