Individual Final Project
In Silico Design of a Humanized Chimeric Arginine Deiminase
![cover image]()
![cover image]()
ABSTRACT
Arginine deiminase (ADI) is a bacterial enzyme with promising therapeutic potential for the treatment of arginine-auxotrophic cancers such as hepatocellular carcinoma, melanoma, and mesothelioma. ADI exerts its antitumor effect by depleting extracellular arginine, an amino acid essential for the survival of certain tumor cells lacking argininosuccinate synthetase expression. Despite its therapeutic promise, the bacterial origin of ADI leads to high immunogenicity, which may trigger immune responses, reduce enzyme efficacy, and limit repeated administration. This project aims to design a humanized chimeric ADI in silico by identifying immunogenic epitopes in bacterial ADI and substituting them with structurally analogous regions from the human arginine-metabolizing enzyme while preserving catalytic function. Structural modeling, immunogenic epitope prediction, molecular docking, and protein validation analyses will be performed to evaluate the stability and substrate-binding ability of the engineered chimera. The resulting humanized ADI is expected to exhibit reduced immunogenicity while maintaining therapeutic activity, providing a potential strategy for improving enzyme-based cancer therapies.
PROJECT AIMS
To identify immunogenic epitopes in bacterial arginine deiminase using in silico epitope prediction tools.
To identify structurally compatible regions in the human arginine-metabolizing enzyme for epitope substitution.
To construct a chimeric ADI model by replacing immunogenic bacterial epitopes with human peptide fragments.
BACKGROUND
Cancer cells often undergo metabolic reprogramming to sustain rapid proliferation. Some tumors, including melanoma, hepatocellular carcinoma, and mesothelioma, are unable to synthesize sufficient arginine due to low expression of argininosuccinate synthetase (ASS1). As a result, these cancers become dependent on extracellular arginine for growth and survival. This metabolic vulnerability makes arginine deprivation an attractive therapeutic strategy.
Arginine deiminase (ADI) is a microbial enzyme that catalyzes the hydrolysis of arginine into citrulline and ammonia, thereby depleting extracellular arginine and suppressing tumor growth. ADI derived from bacterial sources has shown significant antitumor activity, making it a promising candidate for enzyme therapy. However, as a foreign bacterial protein, ADI can induce immune responses in patients, resulting in antibody production, rapid clearance, hypersensitivity, and decreased therapeutic efficacy.
One promising approach to overcome this limitation is protein humanization, in which immunogenic regions of a bacterial therapeutic enzyme are replaced with structurally similar fragments from human proteins. This strategy has been successfully demonstrated in previous in silico studies on bacterial therapeutic enzymes to reduce antigenicity while maintaining enzymatic activity.
Therefore, this project proposes the in silico design of a humanized chimeric arginine deiminase, where immunogenic epitopes of bacterial ADI are replaced with homologous regions from a human arginine-metabolizing protein. This design aims to reduce immunogenicity while preserving the enzyme’s ability to bind arginine, thereby improving its therapeutic potential for cancer treatment.
STEP-BY-STEP METHODS (IN SILICO WORKFLOW)
1. Retrieval of Protein Structures and Sequences
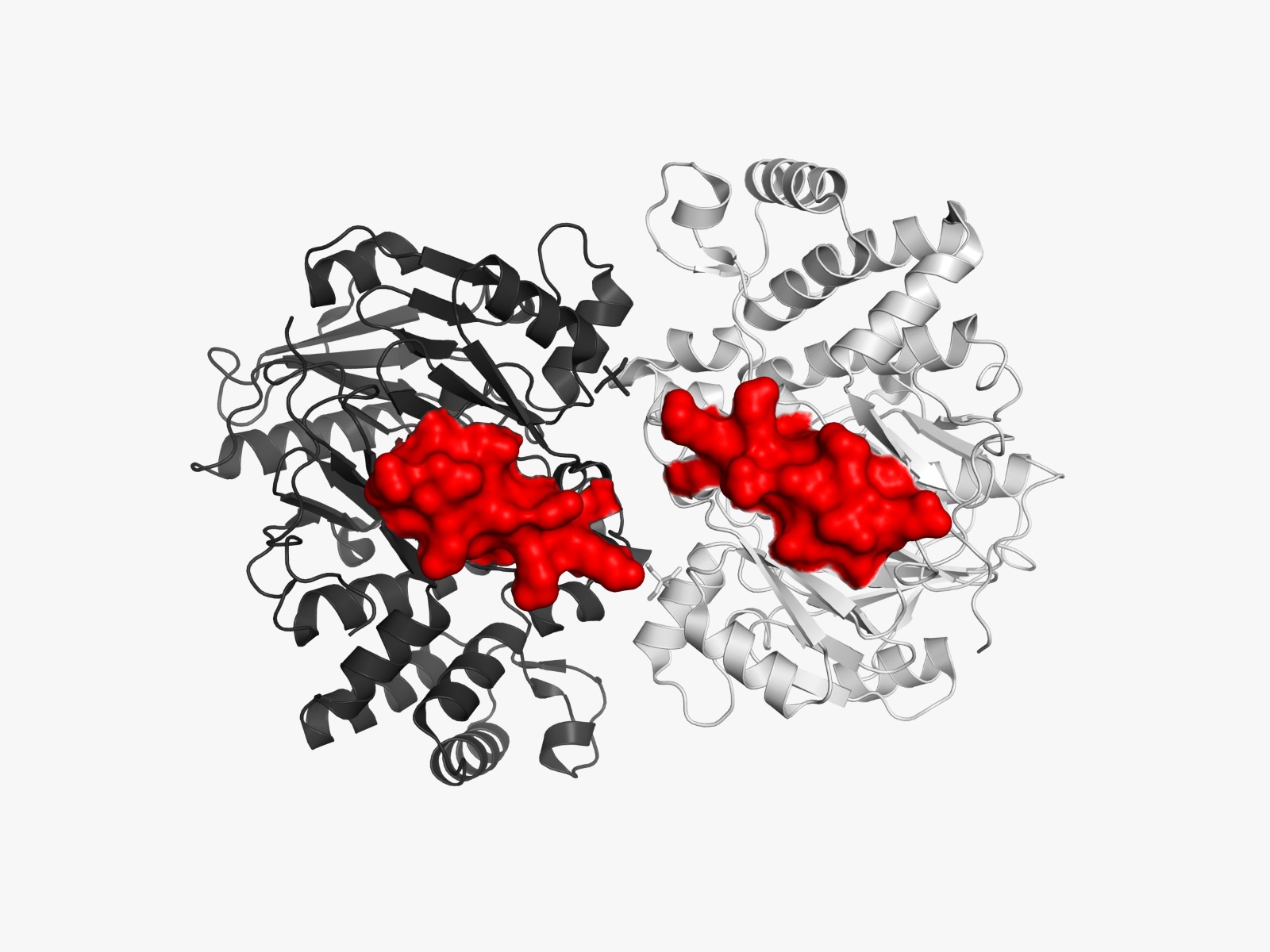
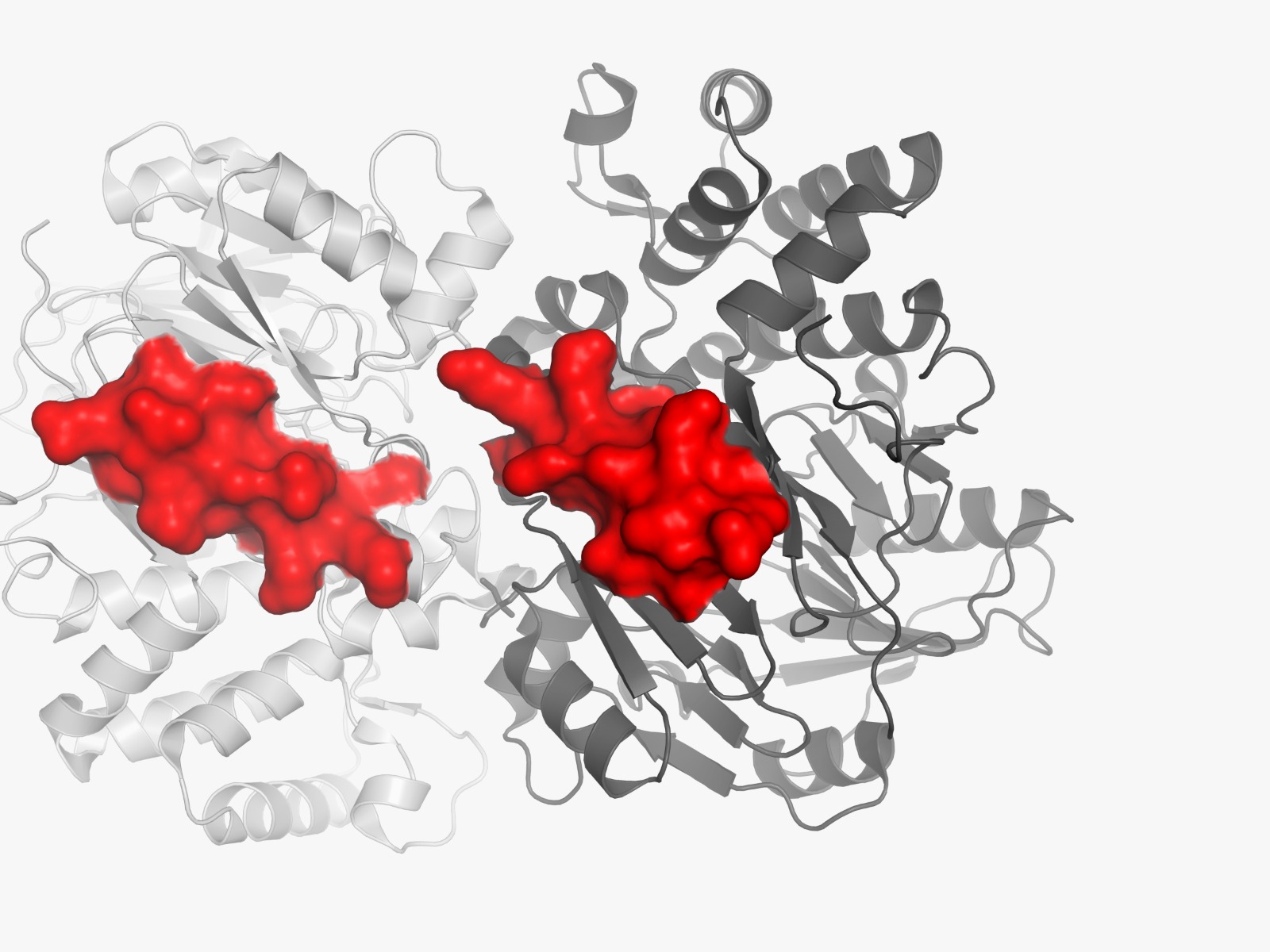
The amino acid sequences and three-dimensional structures of arginine deiminase (ADI) from Mycoplasma arginini (PDB ID: 1LXY) and human arginase I (PDB ID: 2AEB) will be retrieved from the Protein Data Bank (PDB).
2. Prediction of Immunogenic T-cell Epitopes in Bacterial ADI
To predict immunogenic T-cell epitopes in bacterial arginine deiminase (ADI; PDB ID: 1LXY), the protein’s amino acid sequence will be submitted to the NetMHCIIpan server for MHC class II binding prediction, focusing on regions likely to elicit T-cell immune responses. The sequence will be fragmented into overlapping peptides of 9–15 amino acids, with binding affinities evaluated against common HLA-DR alleles—HLA-DRB101:01, HLA-DRB103:01, HLA-DRB104:01, HLA-DRB107:01, HLA-DRB108:01, HLA-DRB111:01, HLA-DRB113:01, and HLA-DRB115:01—which serve as reference alleles with broad global frequency in the literature. Epitope density will be determined for each protein, selecting peptides classified as strong binders (SB, IC50 ≤ 1%) or weak binders (WB, IC50 ≤ 5%) as candidate immunogenic epitopes for targeted substitution.
3. Allergenicity Prediction of Candidate Epitopes
Candidate immunogenic epitopes identified from bacterial ADI will undergo allergenicity prediction using AllerTOP v.2.0 to assess their potential to provoke allergic responses. Epitopes classified as probable allergens will be prioritized for substitution, ensuring targeted modification of regions most likely to elicit adverse immune reactions
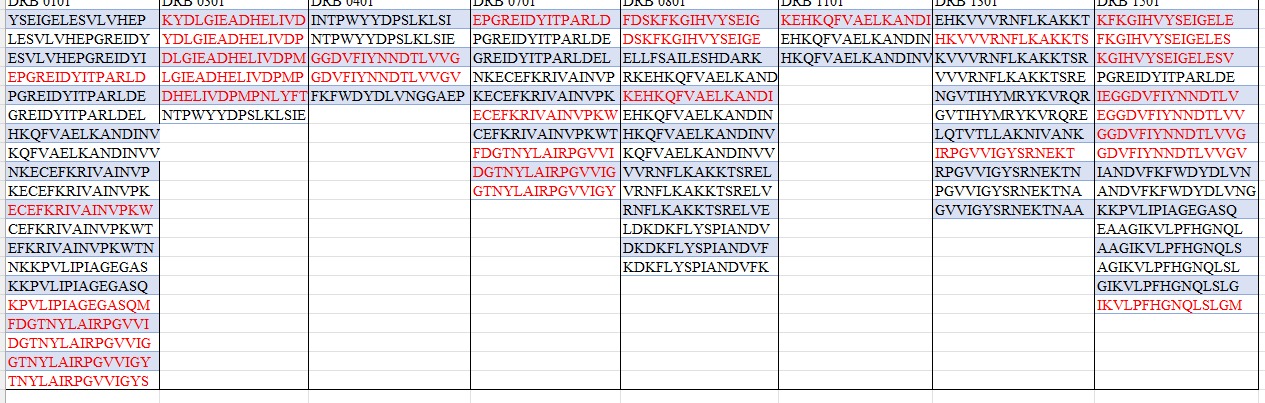

4. Epitope Alignment and Mapping
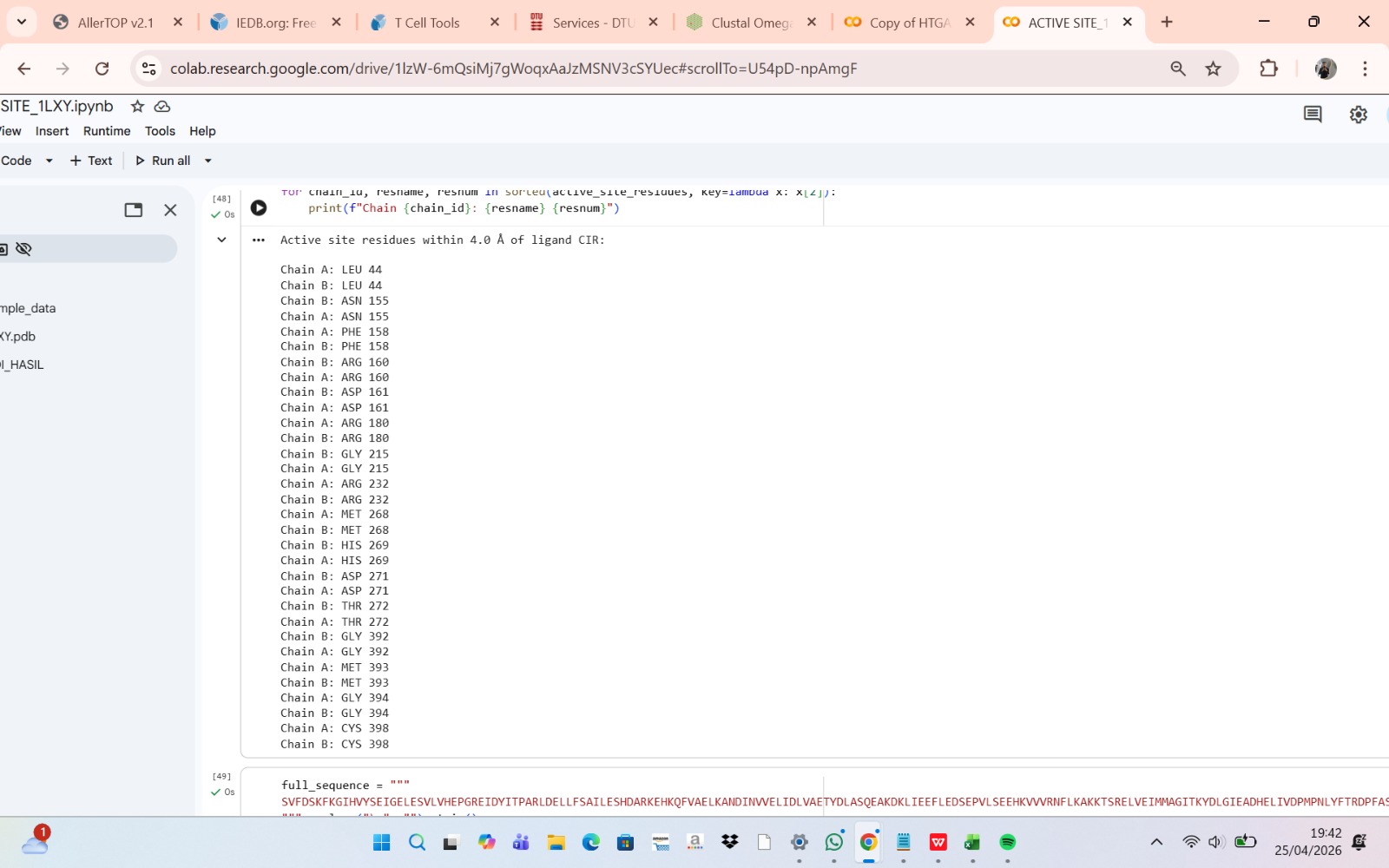
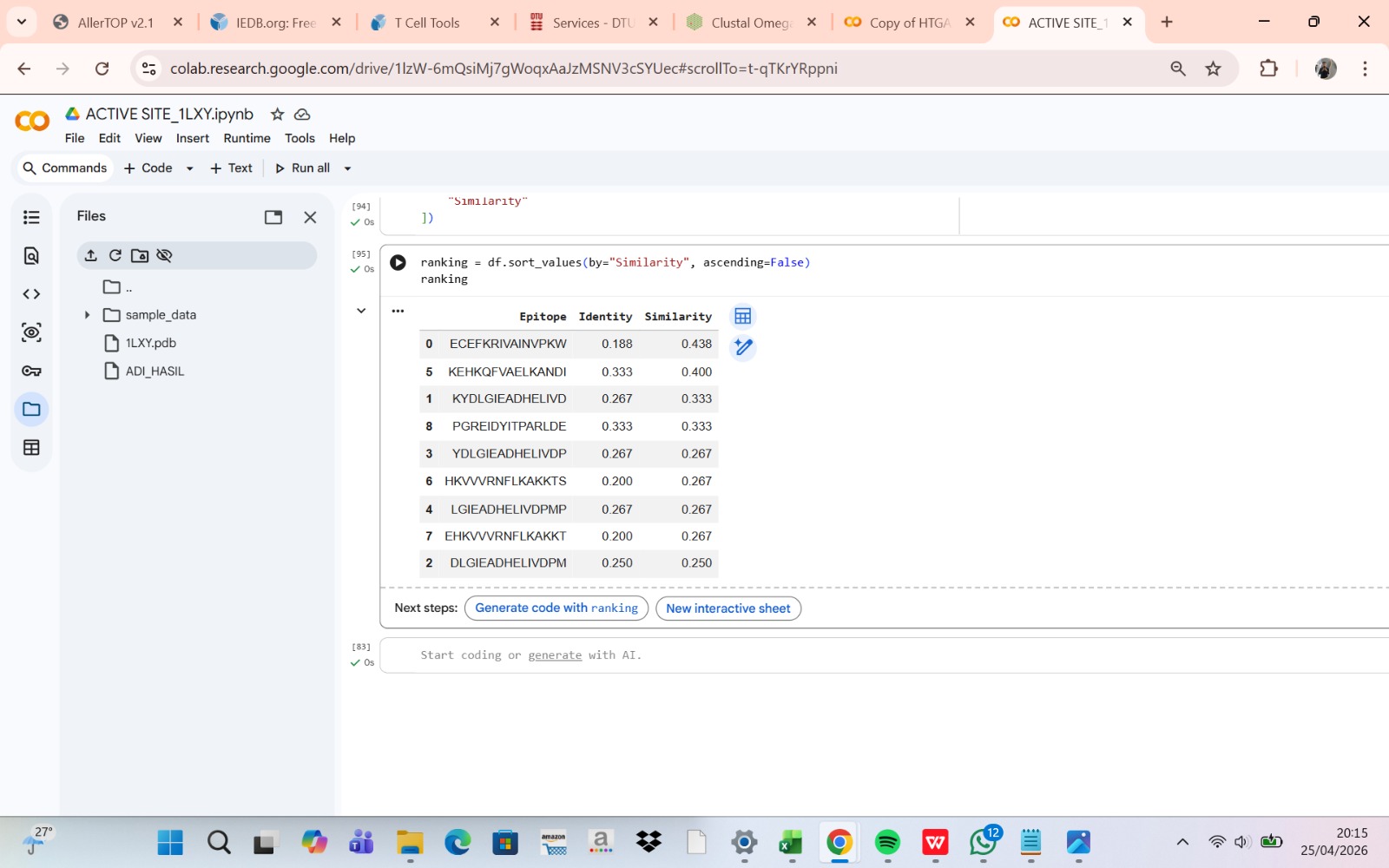
Not all predicted allergenic epitopes will be substituted; selection will prioritize those that are surface-exposed (solvent-accessible), readily recognizable by the immune system, located outside the active site, and non-essential for structural stability. This focuses substitution on the “most exposed” residues—defined as surface-accessible epitopes—rather than solely those with the highest immunogenicity scores. Active site residues were identified by detecting amino acid residues located within 4 Å of the native ligand in the crystal structure using BioPython. These residues were excluded from epitope substitution to preserve catalytic activity.

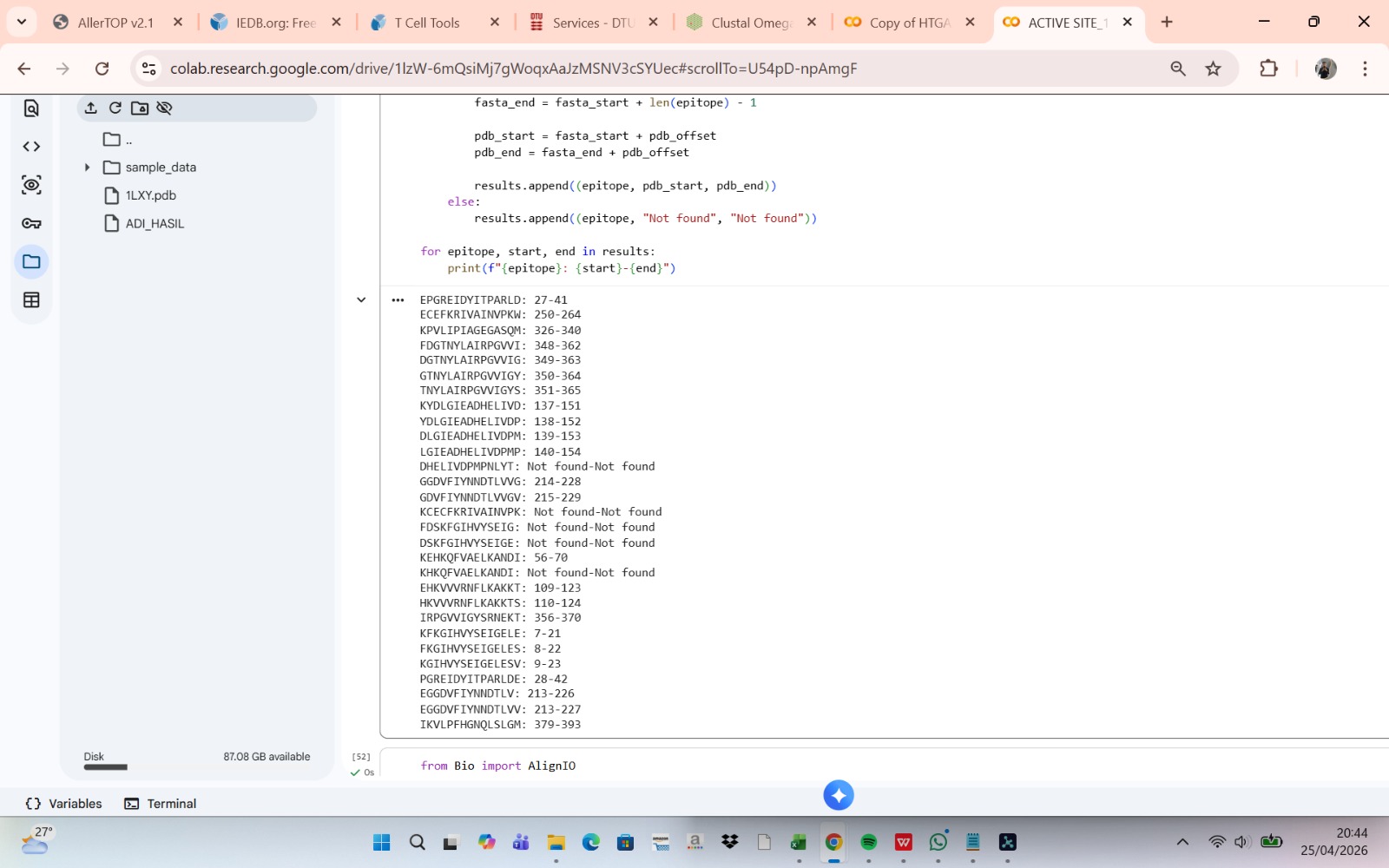
The linear positions of each epitope within the target protein sequence are mapped using Biopython (SeqIO module). The full protein sequence is read from a FASTA file, and each epitope peptide is located via substring matching to determine the start and end residues. This positional information facilitates overlap analysis with active site residues and three-dimensional structural mapping.




BIBLIOGRAPHY REFERENCE
Pedroso, A., Herrera Belén, L., Beltrán, J. F., Castillo, R. L., Pessoa, A., Pedroso, E., & Farias, J. G. (2023). In silico design of a chimeric humanized L-asparaginase. International Journal of Molecular Sciences, 24(8), Article 7550. https://doi.org/10.3390/ijms24087550
Group Final Project
![cover image]()
![cover image]()
Hypothesis: Substitution of a bacteriophage’s replisome with an orthogonal T7 replisome for continuous hypermutation directed towards stability
Group Members:
Alan Bravo https://pages.htgaa.org/2026a-alan-bravo
Samudera Mukhalid https://pages.htgaa.org/2026a-samudera-mukhalid
Bacteriophage Engineering
Challange Choosen: Stability
Proposal
This proposal is inspired by the article by Diercks et al. (2024), which harnesses a highly error-prone replisome to drive extreme antibiotic resistance evolution. Primary Goal #1: Enhance bacteriophage resilience to varying environmental conditions, such as temperature and pH. Primary Goal #2: Achieve this stability without impairing the bacteriophage’s infectivity.
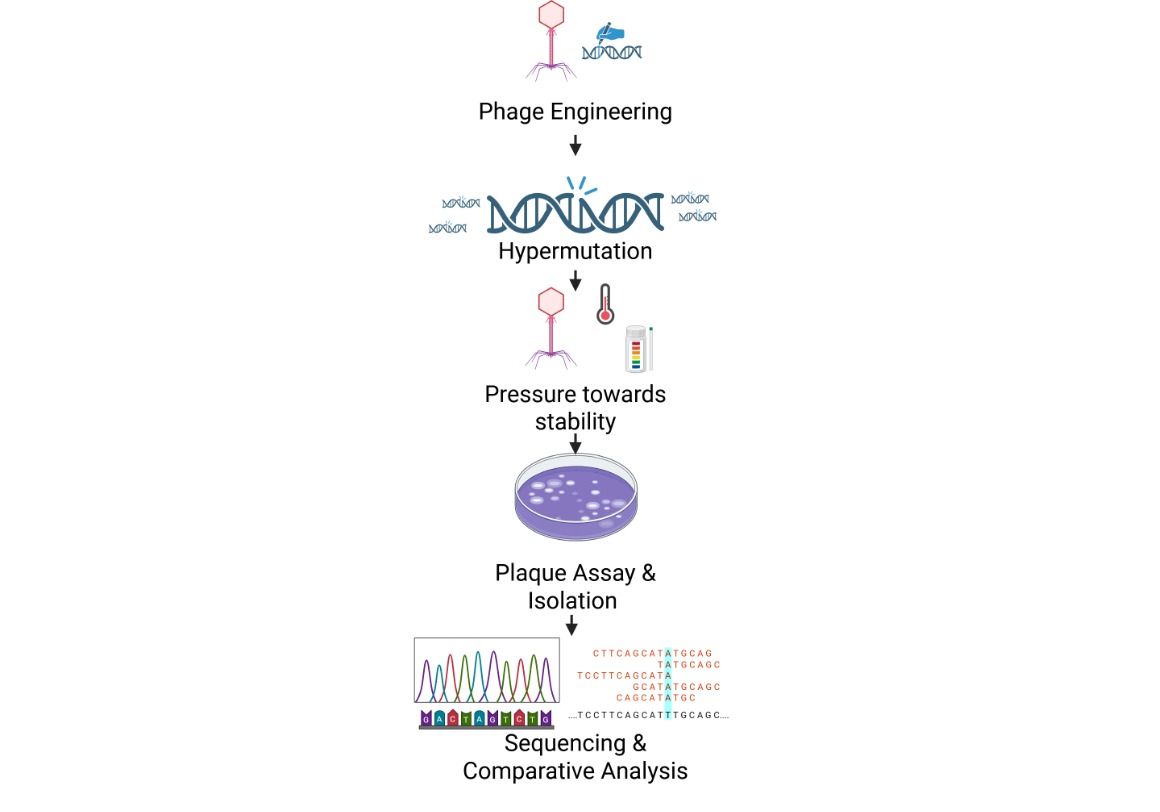
The proposal centers on engineering a bacteriophage based on the T7 replisome, as T7 encodes its own low-fidelity replication machinery. This enables the generation of a mutation-prone phage population, selectively evolved for excellence in the target trait: stability. The population is exposed to targeted stability-related stresses, followed by isolation of infectious survivors via plaque assays. Independent survivor clones are then sequenced, and their encoded proteins analyzed using Clustal Omega. This yields a consensus view of conserved residues versus sites recurrently altered under selection, enabling the design of an optimally stable phage construct.
Two key potential pitfalls:
• Clustal Omega results may be overly divergent, complicating the identification of conserved residues.
• No stable phages may emerge due to the replisome’s excessive faultiness.
Schematic

MSA result
Below are the results of the multiple sequence alignment we performed on a set of protein sequences. The rationale for selecting these sequences was to identify which residues in the bacteriophage endolysin are truly conserved, and which ones exhibit sufficient flexibility for engineering enhanced stability without compromising function. To achieve this, we avoided sequences that were either too identical or excessively divergent.
Thus, the chosen sequences were deliberately selected within an intermediate homology range:
• Similar enough → Retain functional homology
• Different enough → Reveal clear patterns of conservation and variation"









