Week 1 HW: Principles and Practices

First, describe a biological engineering application or tool you want to develop and why. This could be inspired by an idea for your HTGAA class project and/or something for which you are already doing in your research, or something you are just curious about.
As a Biologist, I have a macro-scale perspective on life, from organisms to ecosystems to planetary systems, and have always been drawn to technological innovations. However, I am now curious about the fundamental question of what constitutes life at a micro-scale, and what does engineering its core principles entail. Still interested in biocomputational methods, I want to learn more about the intersection of bio-artificial intelligence and synthetic biology.
My initial research led me to concepts such as distributed computing, logic gates and perceptron-based learning algorithms. Then, I first encountered the term “biocomputer”, which I understand is analysing how living systems perform computation functions, and in some cases the living systems are used to perform those functions as well. In the research paper by Sarkar et al. (2021) titled “Engineered Bacteria Computationally Solve Chemically Generated 2X2 Maze Problems”, the authors programmed E.coli with genetic circuits to solve maze problems within a chemical mixture introduced inside the tubes where the bacterias were incubated in (Siobhan Roberts, 2021). They observed that the bacteria were able to solve the maze problems by analysing different maze configurations.
Inspired by this and other similar research, I would like to further explore the problem-solving capacities of other microorganisms. I am curious to see if similar genetic programming can be applied to other microbial species to solve maze problems and hopefully translate these results in a way that helps us understand new ways to optimize human-made machines.
I am excited to learn more about this in my HTGAA journey, especially knowing that Neuromorphic circuits/computing is part of the course’s curriculum. If I find new topics that spark my interest, I will add them to the list below: Biocomputers, logic gates, learning algorithms
Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals.
For an “ethical” future in relation to biocomputer and bio-artificial intelligence research, I propose three main principles:
Non-malfeasance ✮
Safety ⚘
Respect ☀
Transparency ✿
I propose the following goals, encompassed within one or more of the main principles mentioned previously:
A. Prevent creation/release of harmful organisms ⚘: When collaborating or working with living microorganisms, researchers should always avoid creating and/or releasing pathogenic organisms. This involves a thorough previous investigation on the particular species’ characteristics and potential risks of it being engineered and exposed to different lab procedures.
B. Minimize harm and resource use in experimentation ✮ ☀: Firstly, researchers should aspire to always minimize harm to all living organisms when working with them inside and out of the lab. Additionally, they should also avoid using more resources than they need, this requires a well thought out initial plan and constant readjustments of materials, time and procedures throughout the experimental portion of the research.
C. Ensure accurate public and scientific understanding ✿: Science has to be more democratized, especially when it is cutting-edge innovations like synthetic biology. I believe a way of doing so is by open communication with the general public using accessible friendly language.
D. Promote constructive applications of the technology ✮ ✿: True innovation should inspire applications that are ethical, fair, and beneficial for both human and more-than-human life. Achieving this requires active collaboration among diverse groups and expertise. By integrating diverse perspectives, we can better study expectations and needs, hopefully creating shared, mutualistic goals for our collective future.
Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”). Try to outline a mix of actions (e.g. a new requirement/rule, incentive, or technical strategy) pursued by different “actors” (e.g. academic researchers, companies, federal regulators, law enforcement, etc). Draw upon your existing knowledge and a little additional digging, and feel free to use analogies to other domains (e.g. 3D printing, drones, financial systems, etc.). Purpose: What is done now and what changes are you proposing? Design: What is needed to make it “work”? (including the actor(s) involved - who must opt-in, fund, approve, or implement, etc) Assumptions: What could you have wrong (incorrect assumptions, uncertainties)? Risks of Failure & “Success”: How might this fail, including any unintended consequences of the “success” of your proposed actions?
Previous risk analysis: the projects should be reviewed and approved by an Institutional Biosafety Committee (IBC) (Institutional Biosafety Committee, n.d.) and/or an established Ethics Review Board after presenting a thorough risk assessment of the chosen organism and genetic programming for biocomputational research.
Establishing welfare margins: there could be an international guideline created by a wide community of academics from science to ethics where there is an established welfare margin for microbial stress in experimental designs to minimize demonstrable harm without scientific necessity. These guidelines would be based on known and measurable physiological indicators, and would help promote a duty of care for all living systems, including microorganisms.
(I recognize this can be considered unnecessary as it is ambitious and could involve an almost philosophic discussion on the care for microorganisms in scientific research. However, I feel that as researchers we should prioritize not generating stress and/or pain to any living organism.)
Bioethics compliance: as a condition for publication, scientific journals should require a statement/certificate of ethical review by the researcher team and an established Ethics Review Board. This certificate states that the research methods are compliant with international bioethical laws and guidelines (such as the Universal Declaration on Bioethics and Human Rights or Oviedo Convention in Europe) (Fondation Brocher, 2023). Peer reviewers are also encouraged to revise and comment on the bioethical approaches of the experimental procedures.
Research efficiency and sustainability standards: Synthetic biology labs (and all research institutions in general) should focus on research efficiency and establishing sustainability standards. I propose a series of documents that would provide a skeleton for periodic resource efficiency check-ins during lab meetings. To motivate research teams to adhere to this strategy, institutions could create an annual recognition for research teams that demonstrate a responsible use of resources and waste while maintaining rigorous science. Also, being awarded previously could increase the chances of acquiring further funding for the research.
Public engagement and education: A portion of research funding must be used for the researchers to actively engage with the public using (or teaming up with) scientific communication initiatives (public forums, workshops, interactive talks, etc.), explaining the key takeaways from their research and the limits of biocomputation to avoid sensationalism or misinterpretation.
Key actors summary:
- Research team
- Institutional Biosafety Committee (IBC)
- Ethics Review Board
- Scientific journal
- Peer reviewers
- Funding agencies
- The general public
- Scientific communicators
- Institutions
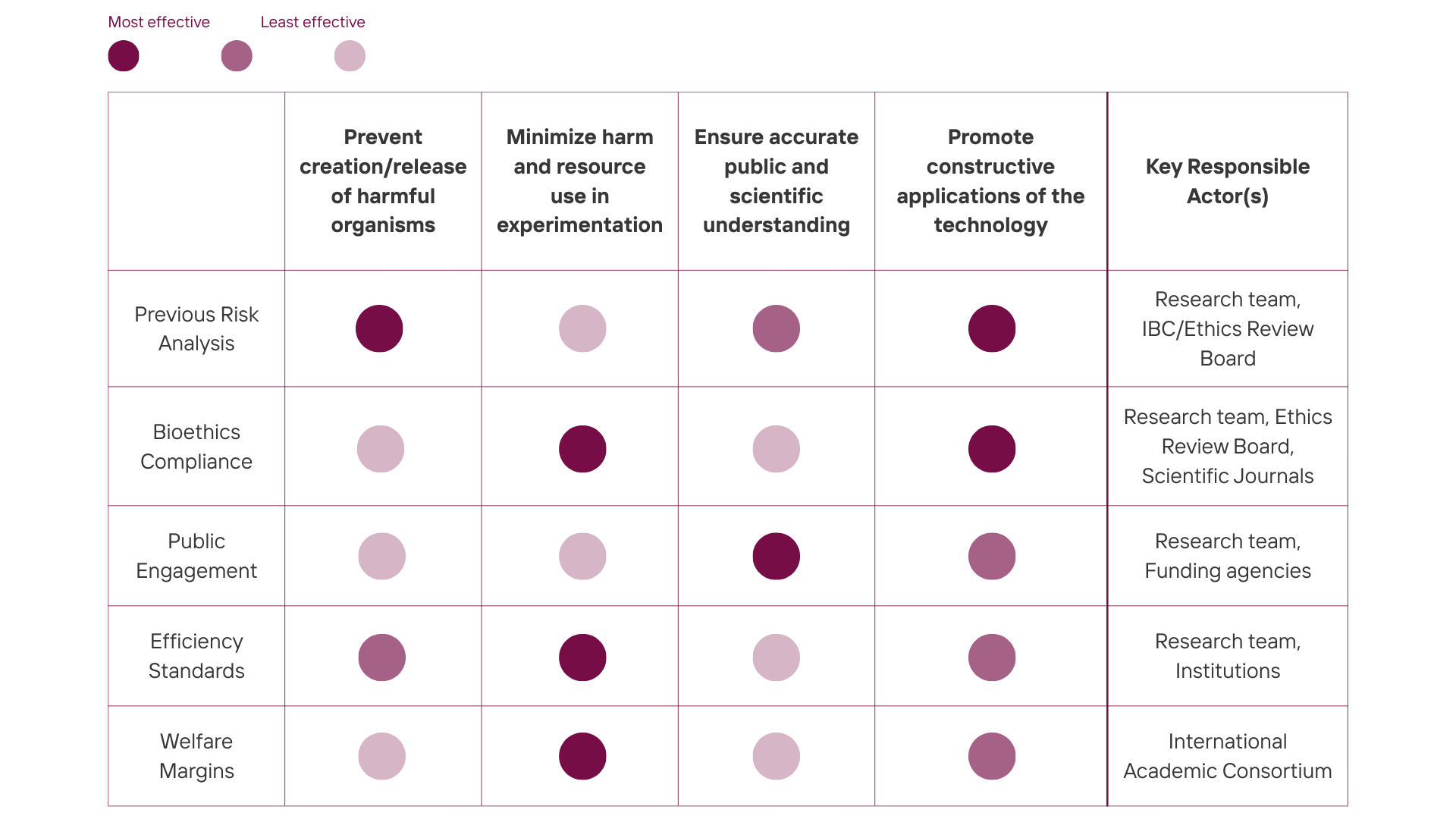
Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals. The following is one framework but feel free to make your own:
Governance actions are scored 1 (least effective) to 3 (most effective).
| Governance Action | Prevent creation/release of harmful organisms ⚘ | Minimize harm and resource use in experimentation ✮ ☀ | Ensure accurate public and scientific understanding ✿ | Promote constructive applications of the technology ✮ ✿ |
|---|---|---|---|---|
| Previous risk analysis | 3 | 1 | 2 | 3 |
| Establishing welfare margins | 1 | 3 | 1 | 2 |
| Bioethics compliance | 1 | 3 | 1 | 3 |
| Research efficiency and sustainability standards | 2 | 3 | 1 | 2 |
| Public engagement and education | 1 | 1 | 3 | 3 |
Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties. For this, you can choose one or more relevant audiences for your recommendation, which could range from the very local (e.g. to MIT leadership or Cambridge Mayoral Office) to the national (e.g. to President Biden or the head of a Federal Agency) to the international (e.g. to the United Nations Office of the Secretary-General, or the leadership of a multinational firm or industry consortia). These could also be one of the “actor” groups in your matrix.
Based on the scoring matrix, my top priorities are: the previous risk analysis, the bioethics compliance and the public engagement and education. I believe these three address the most critical breaking points. Risk analysis is non-negotiable because it prevents harmful microorganisms from spreading and endangering other living forms; bioethics compliance legitimizes research and promotes duty of care for all living organisms; and public engagement and education helps build public trust and accurate understanding necessary for the field’s long-term survival. The other governance options, on the other hand, while important, are not critical. They should be encouraged as best practices as they address less immediate risks.


Reflecting on what you learned and did in class this week, outline any ethical concerns that arose, especially any that were new to you. Then propose any governance actions you think might be appropriate to address those issues. This should be included on your class page for this week.
This week’s discussion on a collaborative bio-future and the role of trust was interesting. I agree trust is essential for ethical progress, but it raised a practical concern for me: I realized I don’t fully understand the current, specific mechanisms and laws for it. I wonder what specific laws, committees, and step-by-step procedures actually check research ethics today? To address this knowledge gap I think there should be more scientific communication around this. It would be a road to strengthen trust and general understanding of ethics as a key priority for scientific research.
References:
Fondation Brocher. (2023, March 16). Bioethics and laws - Brocher. https://fondation-brocher.ch/bioethics-and-laws/#weglot_switcher
Institutional Biosafety Committee. (n.d.). National Institute of Environmental Health Sciences. https://www.niehs.nih.gov/about/boards/ibc
Roberts, S. (2021, November 11). An E. coli biocomputer solves a maze by sharing the work. MIT Technology Review. https://www.technologyreview.com/2021/11/09/1039107/e-coli-maze-solving-biocomputer/
Sarkar, K., Bonnerjee, D., & Bagh, S. (2021). Engineered Bacteria Computationally Solve Chemically Generated 2X2 Maze Problems. Homi Bhabha National Institute (HBNI). https://doi.org/10.1101/2021.06.16.448778
Week 2 Lecture Prep
Homework Questions from Professor Jacobson:
- Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
According to Albertson & Preston (2006), the estimation for errors that error-prone DNA polymerase is once every 104–105 nucleotides polymerized, it can be lower for polymerases that have proofreading activity and can correct mistakes. For example, “twelve of the 15 known human DNA polymerases have no proofreading activity and are error-prone” (Albertson & Preston, 2006). Compared to the human genome, which is 3.2 billion base pairs long, an error-prone polymerase would make approximately 32,000 errors per cell division. However, there are ways to correct mistakes and significantly lower this statistic: error correcting polymerases, mismatch repair, recombination repair, or double-strand break repair (Dav University, n.d.).
- How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
In double-stranded DNA, there are six possible reading frames: three reading from the top strand, and three reading from the bottom strand. However, just one of the six frames is used to code for a protein, the rest of them do not work because a start codon is necessary to define the frame, and the ribosome binds specifically to the correct initiation site, determining that reading frame for the gene.
Homework Questions from Dr. LeProust:
- What’s the most commonly used method for oligo synthesis currently?
Phosphoramidite synthesis
- Why is it difficult to make oligos longer than 200nt via direct synthesis?
Chemical synthesis methods, including the phosphoramidite process, cannot reliably produce oligonucleotides longer than 200 nucleotides. This limitation is due to accumulating errors with each synthetic cycle (Hoose et al. 2023, cited in Yin et al., 2024).
- Why can’t you make a 2000bp gene via direct oligo synthesis?
This is because the length is superior to the 200nt that can be reliably created during phosphoramidite synthesis. So to achieve the 2000bp gene, you would have to do multiple rounds of smaller oligos and then stitch them together.
Homework Question from George Church: Choose ONE of the following three questions to answer; and please cite AI prompts or paper citations used, if any.
- What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
The main 10 aminoacids are: Arginine, Isoleucine, lysine, Methionine, Phenylalanine, Histidine, Leucine, Threonine, Tryptophan and Valine. Now, according to the Jurassic Park Wiki, the Lysine Contingency “is intended to prevent the spread of the animals in case they ever got off the island. Dr. Wu inserted a gene that creates a single faulty enzyme in protein metabolism. The animals can’t manufacture the amino acid lysine. Unless they’re continually supplied with lysine by us, they’ll slip into a coma and die”
It seems logical, because it is a second barrier of security in case the animals escape off the island. However, it is mostly a flawed hypothesis. Lysine is already an essential amino acid for all animals, meaning it must be obtained through diet, not synthesized internally. The dinosaurs would have needed to consume lysine-rich foods (meat, legumes, etc.) regardless of their engineering. So in the case of the dinosaurs escaping, other animals or plants would provide them with the necessary lysine, allowing them to survive. Although, maybe another hypothesis could be that the genetic modification may have created an exaggerated dependency on lysine, requiring amounts far greater than any natural diet could provide. In this scenario, Dr. Wu could have supplied a specially concentrated lysine supplement on the island to meet this particular need. If they escaped, even consuming lysine-rich foods in the wild would fail to meet their requirement, which would be a more clever (yet still very science-fiction oriented) option.
References:
Albertson, T. M., & Preston, B. D. (2006). DNA Replication Fidelity: proofreading in Trans. Current Biology, 16(6), R209–R211. https://doi.org/10.1016/j.cub.2006.02.031
Dav University. (n.d.). DNA proofreading and repair. DAV UNIVERSITY. https://davuniversity.org/images/files/study-material/DNA%20proofreading,EDU161.pdf
Hoose A. Vellacott R. Storch M. Freemont P. S. Ryadnov M. G. DNA synthesis technologies to close the gene writing gap. Nat. Rev. Chem. 2023;7:144–161. doi: 10.1038/s41570-022-00456-9. https://dx.doi.org/10.1038/s41570-022-00456-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
Yin Y, Arneson R, Yuan Y, Fang S. Long oligos: direct chemical synthesis of genes with up to 1728 nucleotides. Chem Sci. 2024 Dec 18;16(4):1966-1973. doi: 10.1039/d4sc06958g. PMID: 39759933; PMCID: PMC11694485.