Week 6 HW: Genetic Circuits Part I: Assembly Technologies
Assignment: DNA Assembly
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?

The Phusion High-Fidelity PCR Master Mix contains all the core reagents necessary for accurate and efficient DNA amplification:
- Phusion DNA Polymerase: A high-fidelity enzyme with 3′→5′ exonuclease proofreading activity that minimizes errors during DNA synthesis, especially important for mutation-based cloning (NEB, 2023).
- dNTPs (deoxynucleotide triphosphates): Provide the nucleotide building blocks (A, T, G, C) for DNA strand elongation.
- Reaction Buffer (with Mg²⁺): Maintains the ionic strength and conditions needed for optimal enzyme activity and DNA strand stability.
- Stabilizers & enhancers: Help maintain enzyme performance across temperature ranges and buffer pH changes during thermocycling.
(New England Biolabs (NEB), 2023).
2. What are some factors that determine primer annealing temperature during PCR?
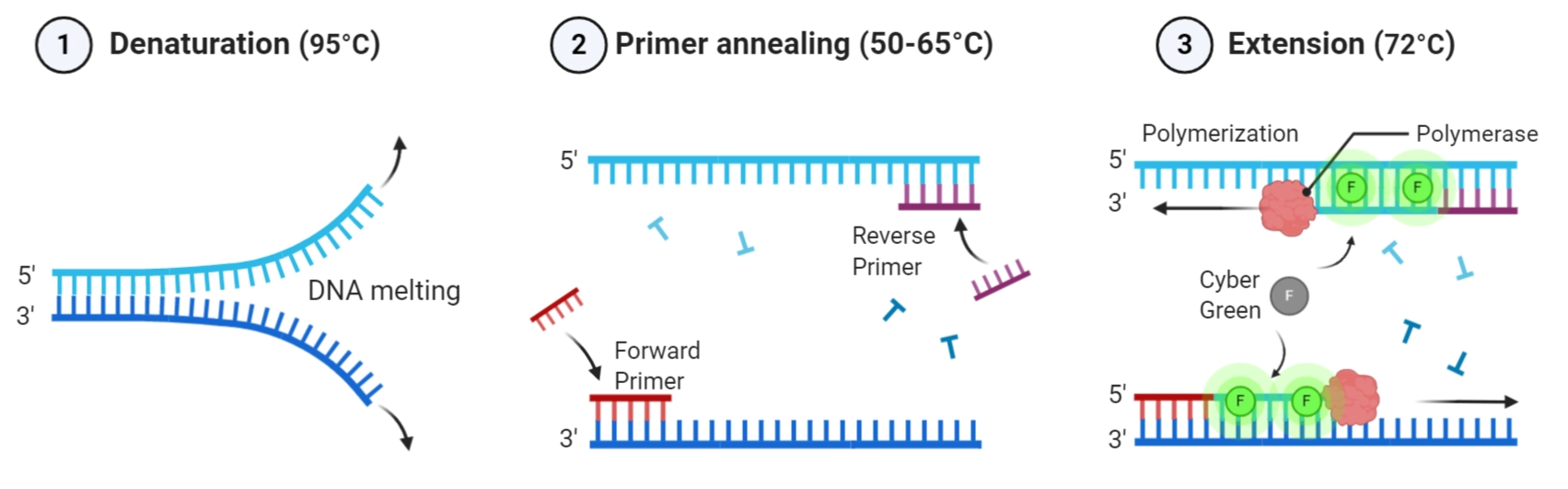

The annealing temperature (Ta) determines primer binding specificity. It depends mainly on:
- Primer length: Longer primers (18–22 bp typical) increase Tm (melting temperature).
- GC content: G–C pairs form three hydrogen bonds (vs. the two hydrogen bonds in A–T pairs), making high GC content primers bind more tightly (raising Tm).
- Salt concentration (especially Mg2+): Stabilizes primer-template binding.
- Secondary structures: Hairpins or dimers can lower effective primer availability, thus altering temperature behavior.
- Complementarity between primers and template: Mismatches like the ones introduced for mutagenesis in this particular lab can reduce effective binding and could require lower Tas.
(Addgene Primer Design Guide; Primer 3 Manual, 2022.)
Primers in this lab are designed for melting temperatures between 52–58 °C, within 5 °C of each other for optimal pairing.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
In this lab, PCR is used to create fragments with intentional mismatches and overlapping ends for Gibson Assembly, which restriction digestion alone cannot provide.PCR uses a DNA copying machine to make many copies of a specific DNA piece, with the start and end points controlled and chosen by the user. This method is very flexible because it allows adding changes to the DNA sequence, like mutations or tags, and works even when there are no natural cutting points present. In contrast, restriction enzyme digestion uses proteins that cut DNA only at specific, short sequences. This method is very precise and predictable at those cut points, but it is limited to locations where these natural cutting sequences exist. PCR is the go-to choice when you need to customize the DNA or when natural cutting sites aren’t available, but digestion is preferred for routine cutting and pasting of DNA fragments when convenient cut sites exist (Green & Sambrook, 2023).
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To ensure PCR and digested DNA fragments are suitable for Gibson Assembly:
- Verify that each fragment has 20–40 bp homologous overhangs complementary to adjacent fragments (as introduced by designed primers).
- Confirm that fragments are designed in the correct 5′→3′ orientation for annealing and ligation.
- Check purity and concentration: Check the concentration of the plasmid DNA and the purity of the fragments. DpnI digestion is useful to remove template plasmid DNA and Zymo cleanup to purify fragments. Nanodrop or Qubit can be useful to measure concentration.
- Ensure the fragment size matches the predicted outcome from Benchling simulation before assembly.
- Ensure there are no sequence errors by using a high-fidelity polymerase, and verify that overlapping regions match perfectly for efficient Gibson reaction.
(Gibson et al., 2009; NEBuilder HiFi DNA Assembly Guide, 2023).
5. How does the plasmid DNA enter the E. coli cells during transformation?
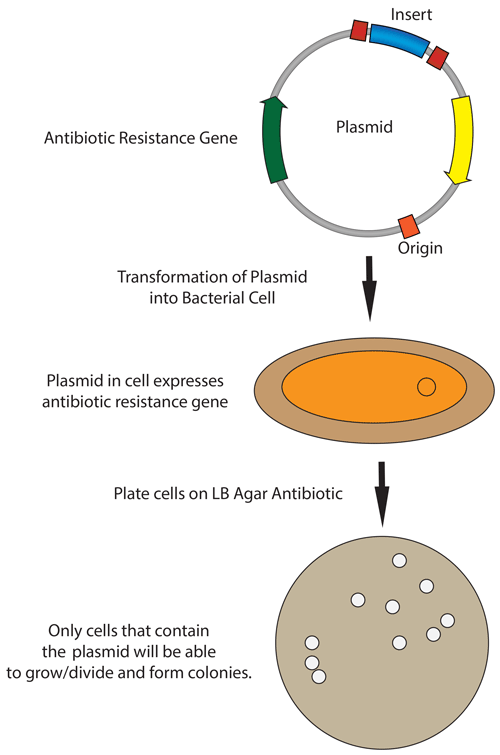

In the protocol from the lab, during heat-shock transformation the plasmid DNA enters the E.coli cells in the following steps:
- Competent E. coli cells are chilled to stabilize the membrane.
- A 42 °C heat shock creates temporary pores in the membrane by rapidly increasing membrane fluidity.
- The plasmid DNA enters the cell through diffusion across the pores.
- Cells are then returned to ice or the nutrient-rich SOC medium for recovery. The membrane then reseals and the plasmid begins replication.
- During plating on chloramphenicol LB agar, only transformed cells containing the plasmid survive due to antibiotic selection.
(NEB Transformation Protocol, 2023; Hanahan, 1983).
6. Describe another assembly method in detail (such as Golden Gate Assembly)
Golden Gate Assembly is a molecular cloning method that uses Type IIS restriction enzymes (for example BsaI) and DNA ligase in a single tube reaction to assemble multiple DNA fragments in one step and with high precision.
This is the step by step mechanism:
- Type IIS enzymes cut DNA outside their recognition sites, creating 4-base overhangs that can be custom-designed for assembly.
- The digestion and ligation occur in a thermocycling reaction that alternates between 37 °C (cutting) and 16 °C (ligation) steps.
- Because recognition sites are removed in the process, the resulting construct lacks unwanted “scar” sequences. This method has advantages. First, it can enable efficient one-pot assembly of multiple fragments, and it’s speedy and has high accuracy (so no need for overlapping sequences as in Gibson assembly).
(Engler et al., 2008; NEB Golden Gate Assembly Technical Resource).
A. Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Here’s a diagram made by Mary Gearing that I think illustrates Golden Gate Assembly pretty clearly:
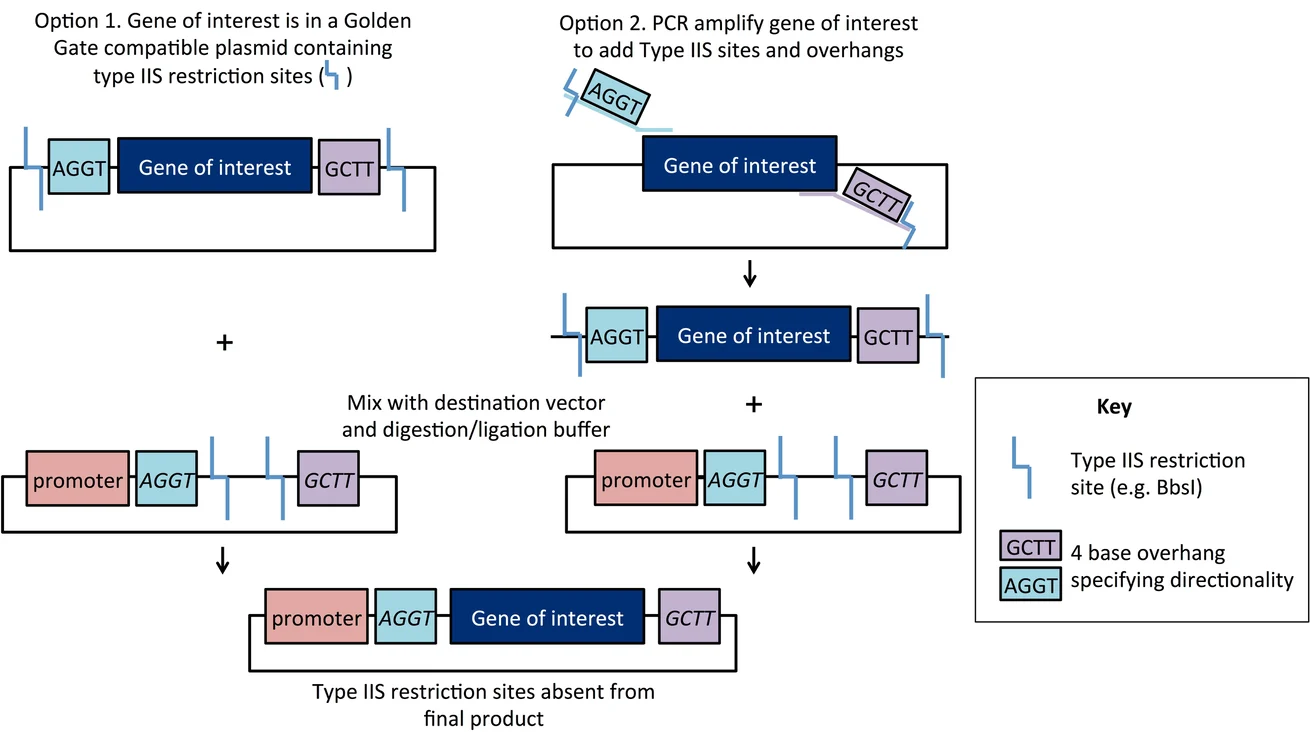

B. Model this assembly method with Benchling or Asimov Kernel!
To model it in Benchling, I decided to use pUC19 because it is a common, well-studied cloning vector. It has a high copy number, an ampicillin resistance gene for selection, and a multiple cloning site (MCS) where I can insert new DNA. Golden Gate Assembly uses Type IIS enzymes like BsaI. So after importing the pUC19 sequence, I located the multiple cloning site (MCS) in pUC19 and added GGTCTC before it and GAGACC after it (see pink annotations on the image below - these are the inserted BsaI cut sites). These are the recognition sites where BsaI will cut.
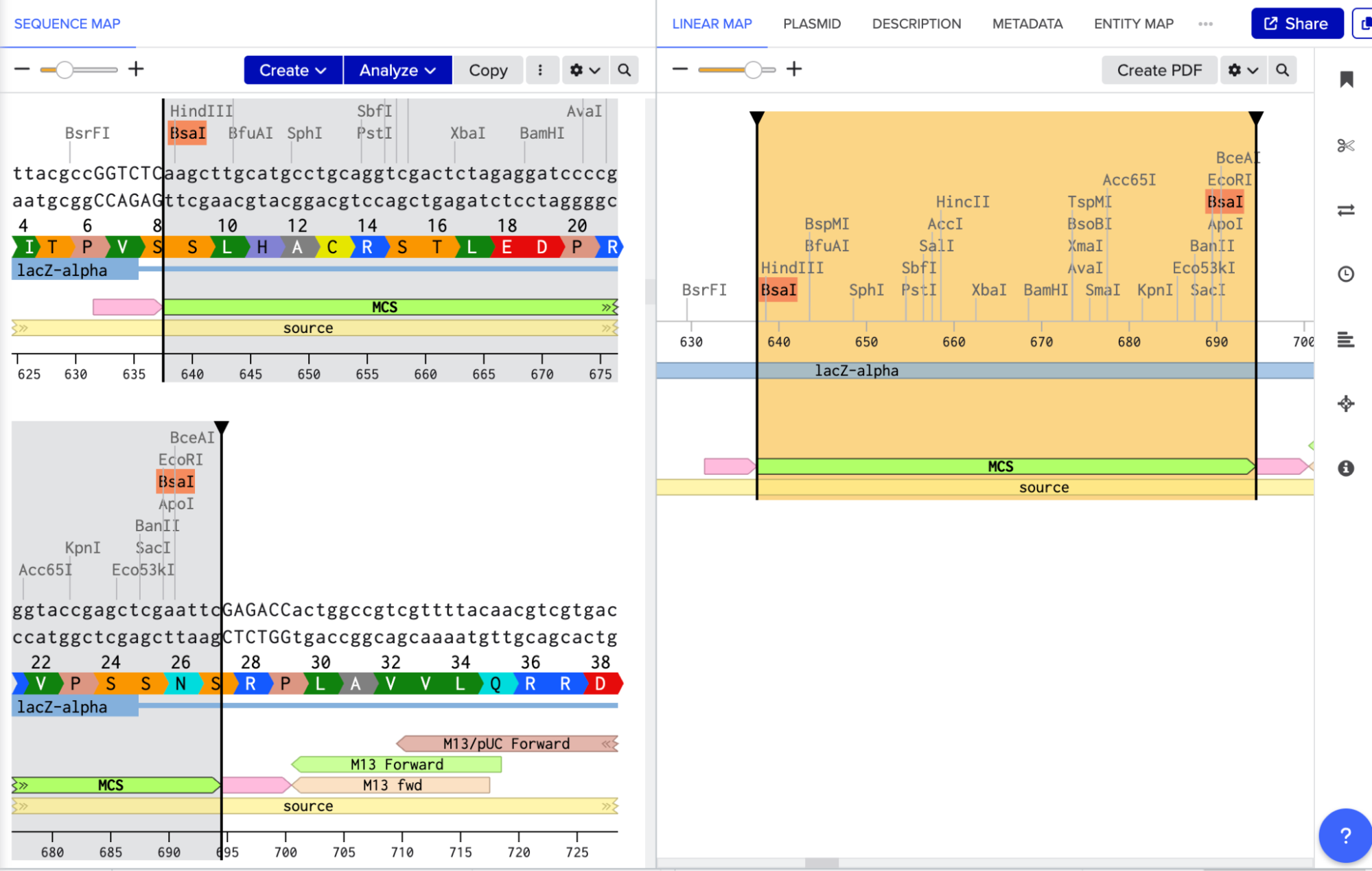

I needed a gene to insert, so I chose RFP (red fluorescent protein) as my insert. I added GGTCTC at the start and GAGACC at the end so it would be compatible with my backbone.
I opened Benchling’s Golden Gate Assembly tool. I selected my backbone fragment and my insert fragment, set the enzyme to BsaI, and clicked “Assemble.”
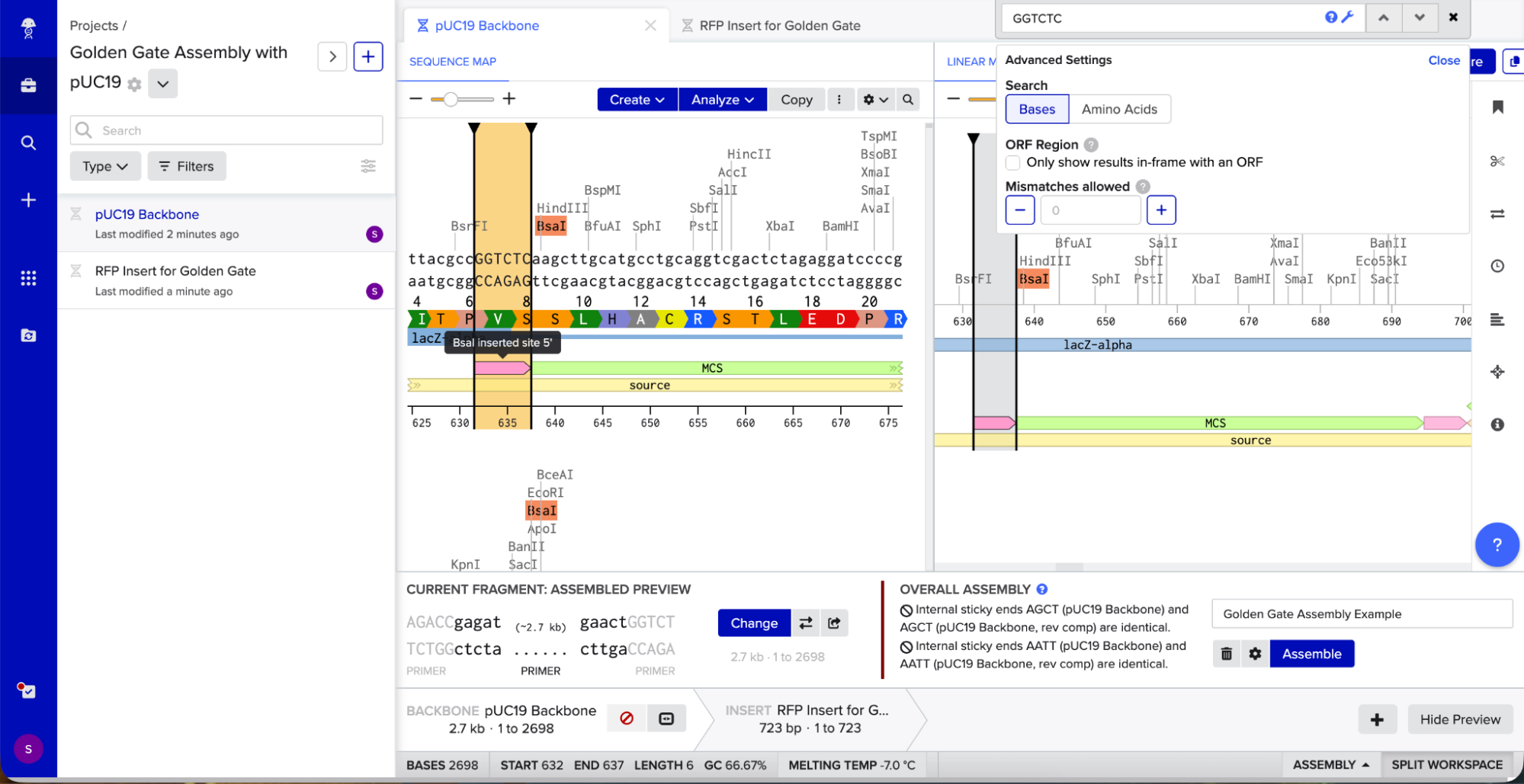

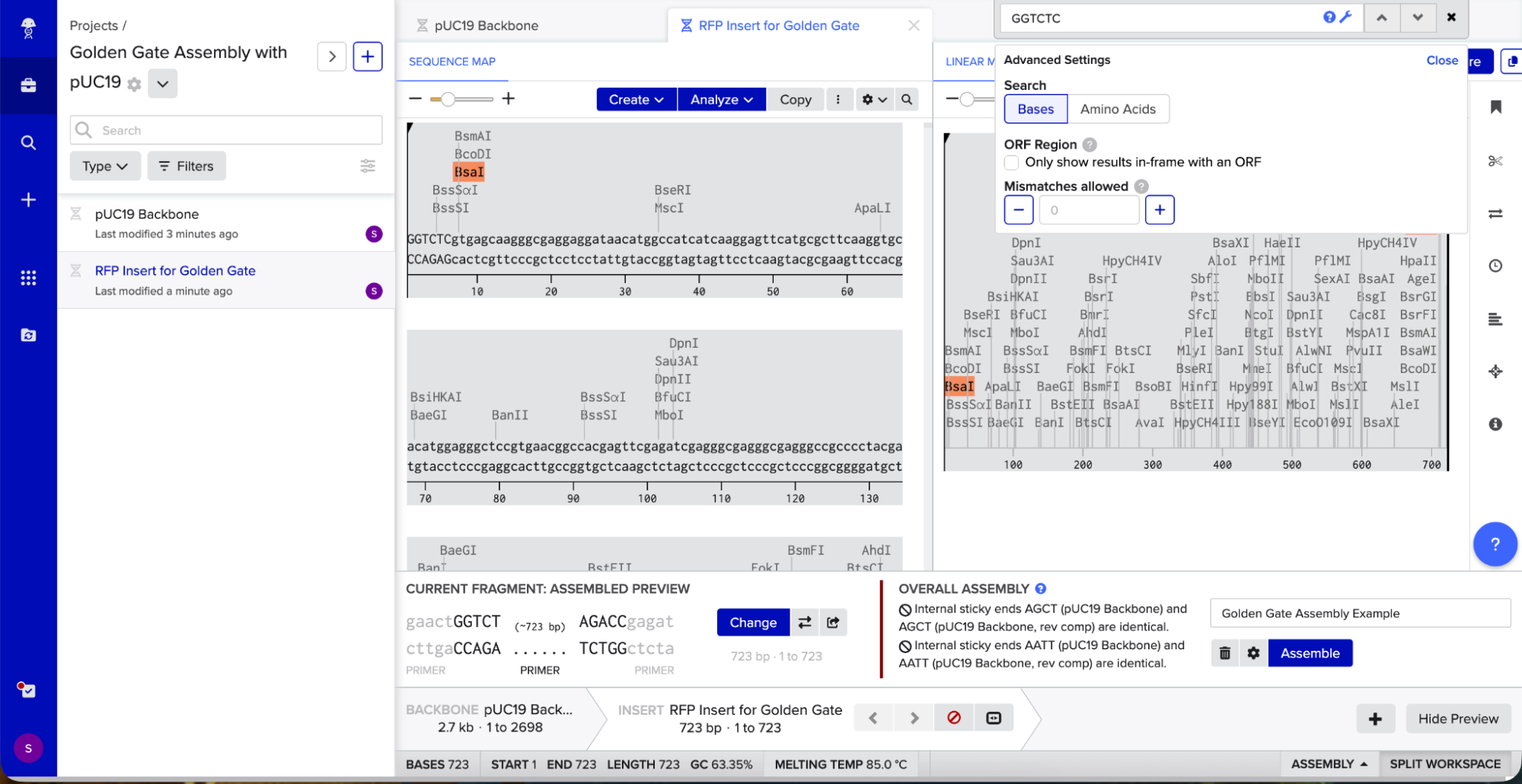

Benchling created a new circular plasmid with my RFP insert now inside the pUC19 backbone. I also checked where the backbone meets the insert, and the BsaI sites (GGTCTC and GAGACC) were gone. This is the proof that Golden Gate Assembly worked because the recognition sites are removed during assembly, leaving no “scar” sequence.
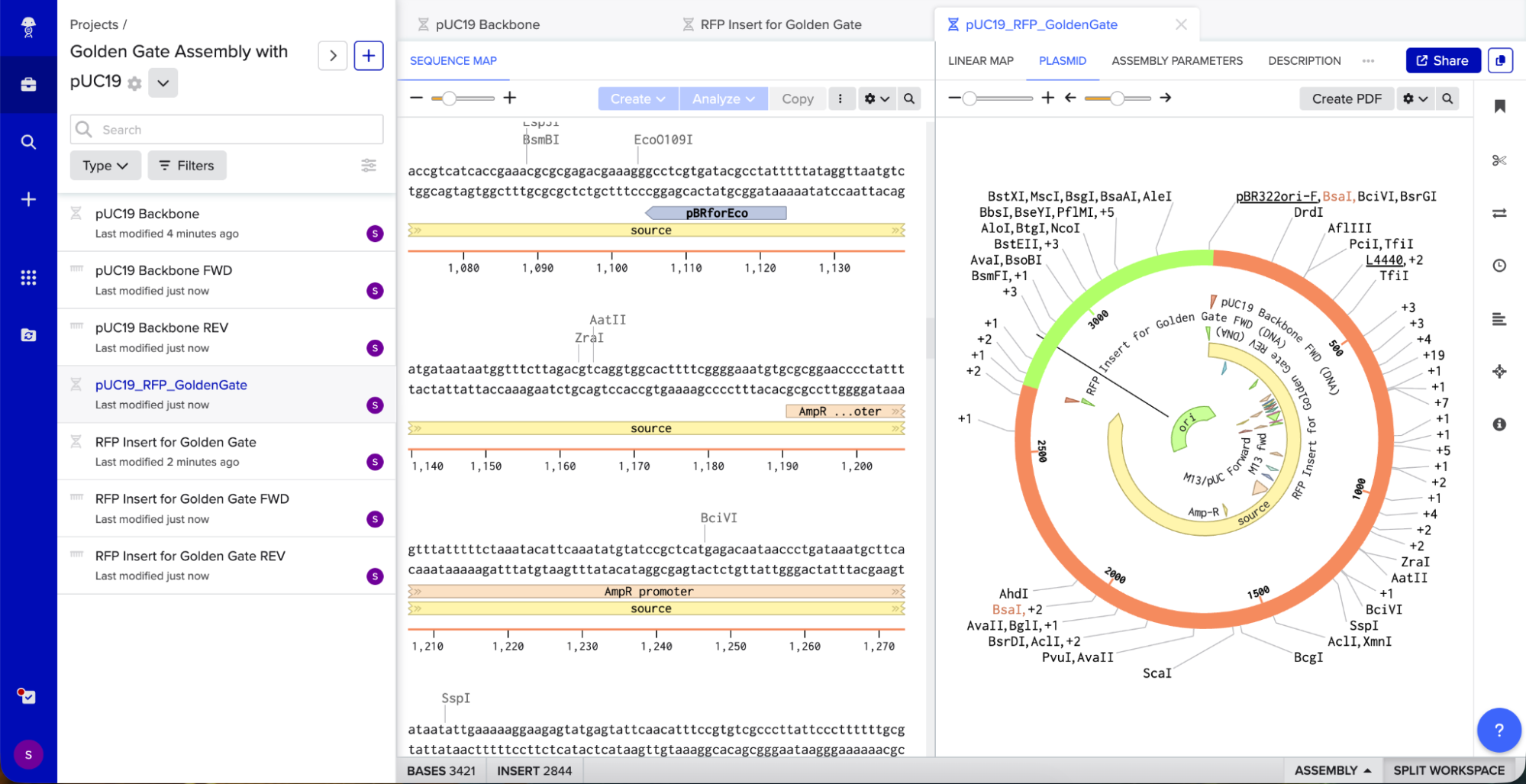

Here’s the link to the project in case you want to check it out: https://benchling.com/s/seq-0WBjIVZttA5D9Pd7fUMv?m=slm-VNUyspCpRxjNPsZrUfhB
References:
- New England Biolabs (NEB) Phusion® High-Fidelity DNA Polymerase—Product Manual, 2023
- Addgene Primer Design Guide; Primer 3 Manual, 2022.
- Green & Sambrook, Molecular Cloning: A Laboratory Manual, 4th ed.; NEB Cloning Guide (2023).
- Gibson et al., Nature Methods, 2009, 6(5):343–345.
- NEBuilder HiFi DNA Assembly Guide, NEB 2023.
- NEB Transformation Protocol, 2023.
- Hanahan, D. (1983). “Studies on transformation of Escherichia coli with plasmids.” J. Mol. Biol., 166(4):557–580.
- Engler, C., et al. (2008). “A one pot, one step, precision cloning method with high throughput capability.” PLoS ONE, 3(11): e3647.
- NEB Golden Gate Assembly Technical Resource.