Individual Final Project


MycoBoard engineers the fungus Neurospora crassa to grow flat hyphal mats that function as biodegradable, breadboard-like electronic substrates. Overexpression of the metallothionein gene cmt drives silver ion capture and reduction into conductive nanoparticles along hyphal walls, making the fungus an active circuit architect of the object itself. Mats grown in molds guide the hyphal geometry via its thigmotropism, and conductive tracks are applied via stamp or screen print to emulate the typical Breadboard. The result decomposes in soil within weeks or months, unlike conventional FR4 fiberglass PCBs that contribute to the ongoing e-waste crisis.
For aims 2 and 3 I have broken down the steps to achieve them. Further below you’ll find the step-by-step procedure for aim 1.
Aim 1 - Experimental:
Design and computationally validate the cmt overexpression construct for N. crassa, simulating expected silver biosorption and resistance outcomes
Aim 2 - Development:
Translate the validated computational design into a physical prototype: grow engineered N. crassa mats inside molds and validate by working LED circuit powered through a biologically grown conductive track.
- Transform verified construct into N. crassa via spheroplast electroporation
- Grow flat mats inside laser-cut or 3D-printed breadboard-like shaped molds
- N.crassa hyphae follow channel walls via thigmotropism, forming predefined track layout
- Soak mat in AgNO₃ medium, heat-fix and dry
- Map resistance across mat with multimeter to confirm conductivity along tracks
- Mount a simple LED + resistor circuit with conductive adhesive and validate conductivity
Aim 3 - Visionary:
Replace physical molds with optogenetic control of cmt expression, using an interface to draw circuit layouts that the fungus then follows while growing.
- Engineer light-inducible promoter controlling cmt
- Design circuit layout and project onto growing mat. Illuminated zones activate cmt, silver deposits follow the projected pattern precisely
- Characterise mat decomposition in soil, revise chelation wash protocol if helpful
- Generalise platform to other metals (perhaps copper via CUP1) and fungal chassis (perhaps other species like Fusarium or Ganoderma)
- Expected result: custom biodegradable boards grown from spores, compostable in months
Aim 1 Process
I went to FungiDB, created an account, located the Neurospora crassa cmt gene (gene ID: NCU05561), and imported its coding sequence (CDS) into Benchling.
- The cmt (copper metallothionein) gene encodes a small, cysteine-rich metal-binding protein. In N. crassa, cmt is naturally induced by copper and has been shown to bind heavy metals (Romeyer et al., 1990). For MycoBoard, I want to repurpose cmt to capture and reduce silver ions (Ag⁺) into conductive nanoparticles along the walls of the fungi.
I created a new Benchling file. This file will contain the full, linear DNA construct for direct transformation into N. crassa.
- Neurospora crassa is efficiently transformed with linear DNA via spheroplast electroporation (Nargang & Nargang,1996; Singh et al., 2026; Ploessl, 2022). Unlike bacteria, fungi do not require a circular plasmid backbone for integration; linear fragments recombine into the genome via non-homologous end joining (NHEJ) or homologous recombination. Designing the construct as a linear cassette simplifies the final transformation protocol and avoids unnecessary vector sequences.
I searched FungiDB for the ccg-1 gene (NCU03753). Before downloading the FASTA sequence, I adjusted the settings to retrieve -1500 nucleotides upstream of the start codon (the “Unspliced Genomic Sequence” with 1500 bp of flanking region). This gave me the 1500 bp promoter region plus the ccg-1 gene itself. I extracted only the promoter portion (the sequence before the ATG start codon of ccg-1) and pasted it into my linear cassette.
- The ccg-1 (clock-controlled gene 1) promoter is a well-characterized, strong constitutive promoter in Neurospora crassa (Wang et al., 2023). It drives high levels of transcription without the need for induction, making it ideal for expressing cmt throughout mat growth. Choosing a 1500 bp upstream region ensures inclusion of important regulatory elements (Ohrnberger & Akins, 1995).
I pasted the full N. crassa cmt CDS (84 bp) immediately after the promoter sequence. Then, I removed the native stop codon (TAA) from the end of cmt. Immediately before where the stop codon had been, I inserted the HA tag sequence (TACCCATACGATGTTCCAGATTACGCT, 27 bp) and annotated it separately. Finally, I added a new stop codon (TGA) after the HA tag.
I searched for the pCSN43 vector (a standard fungal transformation plasmid) (Staben et al., 1989; Carroll, 1994) and downloaded its GenBank file from NovoPro. I opened the file in Benchling, identified the trpC promoter, hph (hygR) gene, and trpC terminator, and copied the entire region from the start of the promoter to the end of the terminator. I pasted this cassette at the end of my linear construct (after the cmt-HA stop codon) and added a final stop codon (TAA) at the very end of the hygR gene.
As a summary, here’s the explanation for each part of the construct:
- Pccg-1 promoter: is the ON switch for the gene. It is always active in N.crassa in most experimental conditions, which means its constitutive, making the gene be ON always.
- cmt gene: is the actual gene that encodes the methallothionein protein that binds the silver ions for the fungal mat. It is codon optimized for N.crassa
- HA tag: to check if the protein is actually being made with a Western Blot.
- trpC promoter: drives the expression of the antibiotic resistant gene (HygR), also an active promoter in filamentous fungi.
- HygR: is the selection marker to see which fungal cells actually took up the construct. Hygromycin is an antibiotic that kills N.crassa cells. The cells that successfully incorporate the construct will express HygR, neutralizing the antibiotic and surviving. When growing the engineered cells, only the ones with the introduced DNA will grow, expressing the cmt gene.
- trpC terminator: is the end of the HygR transcription unit.
- In Benchling, I annotated all functional elements:
- Pccg-1 promoter (blue)
- cmt (green)
- HA tag (yellow)
- trpC promoter (purple)
- trpC terminator (pink)
- hygR (hph) (red)
- STOP codons (grey)
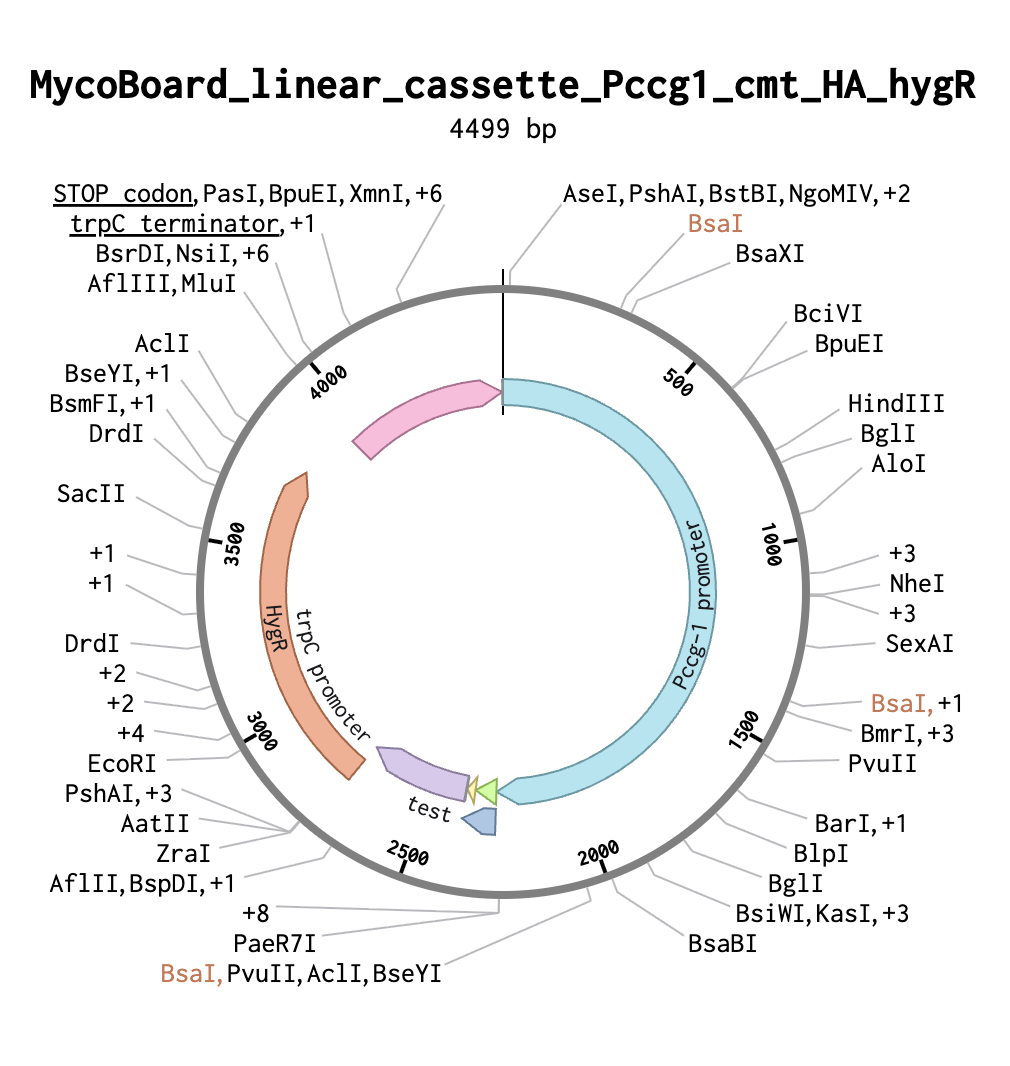

I then ran codon optimization on the cmt gene alone (using Neurospora crassa as the target organism) and replaced the original cmt sequence with the optimized version. Finally, I verified the reading frame by selecting the cmt and HA tag annotations together and confirming that the protein translation showed.
After codon optimization and reading frame verification, I prepared the Twist order. Since the sequence is longer than 1.8 kb, I decided to do two similarly sized fragments.
- Fragment 1 (2,381 bp):
- Start: Beginning of Pccg-1 promoter
- End: Stop codon (TGA) after HA tag
- Fragment 2 (2,118 bp):
- Start: Beginning of PtrpC promoter
- End: End of TtrpC terminator
The TWIST order would come up to $359.92 for both fragments.
Here’s the full Benchling project: https://benchling.com/sarage/f_/IaAEWedyIi-mycoboard/
And here’s the Benchling to the linear cassette only: https://benchling.com/s/seq-pqP10qcYGTRgqqZgmW3H?m=slm-BGJ3tk6YS3hciLTI2bAD
References:
Nargang, C. E., & Nargang, F. E. (1996). Procedure for preparing and transforming spheroplasts of Neurospora crassa.
Singh, K., Sharma, S., Kalia, A., & Manchanda, P. (2026). Advancement in Mushroom Transformation: From Conventional Techniques to Modern Genetic Engineering. Journal of Basic Microbiology, 66(1), e70132.
Ploessl, D. (2022). Developing nuclear and mitochondrial DNA editing techniques for engineering yeasts as novel microbial factories and disease models (Doctoral dissertation, Iowa State University).
Wang, Z., Bartholomai, B. M., Loros, J. J., & Dunlap, J. C. (2023). Optimized fluorescent proteins for 4-color and photoconvertible live-cell imaging in Neurospora crassa. Fungal Genetics and Biology, 164, 103763.
Staben, C., Jensen, B., Singer, M., Pollock, J., Schechtman, M., Kinsey, J., & Selker, E. (1989). Use of a bacterial hygromycin B resistance gene as a dominant selectable marker in Neurospora crassa transformation. Fungal Genetics Reports, 36(1), 79.
Danninger, D., Pruckner, R., Holzinger, L., Koeppe, R., & Kaltenbrunner, M. (2022). MycelioTronics: Fungal mycelium skin for sustainable electronics. Science Advances, 8(45), eadd7118.
Riquelme, M., Aguirre, J., Bartnicki-García, S., Braus, G. H., Feldbrügge, M., Fleig, U., … & Fischer, R. (2018).
Fungal morphogenesis, from the polarized growth of hyphae to complex reproduction and infection structures. Microbiology and Molecular Biology Reviews, 82(2), 10-1128.
Rai, M., Bonde, S., Golinska, P., Trzcińska-Wencel, J., Gade, A., Abd-Elsalam, K. A., … & Ingle, A. P. (2021).
Fusarium as a novel fungus for the synthesis of nanoparticles: mechanism and applications. Journal of Fungi, 7(2), 139.
Ohrnberger, J., & Akins, R. A. (1995). Cloning of the copper-inducible metallothionein (cmt) promoter from Neurospora crassa. Fungal Genetics Reports, 42(1), 57-58.
Baldé, C. P., Kuehr, R., Yamamoto, T., McDonald, R., D’Angelo, E., Althaf, S., … & Wagner, M. (2024). Global e-waste monitor 2024.
Juan, W. A. N. G., Kunhui, S. H. U., & Youbin, S. I. (2017). Effects of silver nanoparticles on soil microbial communities and bacterial nitrification in suburban vegetable soils. Pedosphere, 27(3), 482-490.
Carroll, A. M., Sweigard, J. A., & Valent, B. (1994). Improved vectors for selecting resistance to hygromycin. Fungal Genet. Newsl, 41(22), 135-143.
Romeyer, F. M., Jacobs, F. A., & Brousseau, R. (1990). Expression of a Neurospora crassa metallothionein and its variants in Escherichia coli. Applied and environmental microbiology, 56(9), 2748-2754.
Ohrnberger, J., & Akins, R. A. (1995). Cloning of the copper-inducible metallothionein (cmt) promoter from Neurospora crassa. Fungal Genetics Reports, 42(1), 57-58.