Week 03 HW: Lab Automation
Assignment: Python Script for Opentrons Artwork — DUE BY YOUR LAB TIME!
Your task this week is to Create a Python file to run on an Opentrons liquid handling robot.
0. Review this week’s recitation and this week’s lab for details on the Opentrons and programming it.
1. Generate an artistic design using the GUI at opentrons-art.rcdonovan.com.
2. Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.
・You may use AI assistance for this coding — Google Gemini is integrated into Colab (see the stylized star bottom center);
it will do a good job writing functional Python, while you probably need to take charge of the art concept.
・If you’re a proficient programmer and you’d rather code something mathematical or algorithmic instead of using your GUI coordinates,
you may do that instead.
3. If the Python component is proving too problematic even with AI and human assistance, download the full Python script from the GUI website and submit that:
4. If you use AI to help complete this homework or lab, document how you used AI and which models made contributions.
5. Sign up for a robot time slot if you are at MIT/Harvard/Wellesley or at a Node offering Opentrons automation.
The Python script you created will be run on the robot to produce your work of art!
・At MIT/Harvard? Lab times are on Thursday Feb.19 between 10AM and 6PM.
・At other Nodes? Please coordinate with your Node.
6. Submit your Python file via this form.
Colab Link HTGAA26 Opentrons Colab _ShimadaSayaka
[https://colab.research.google.com/drive/13-Wv0oNY5XePGTKNubb3rZWO9Zlw5PAz?usp=sharing]
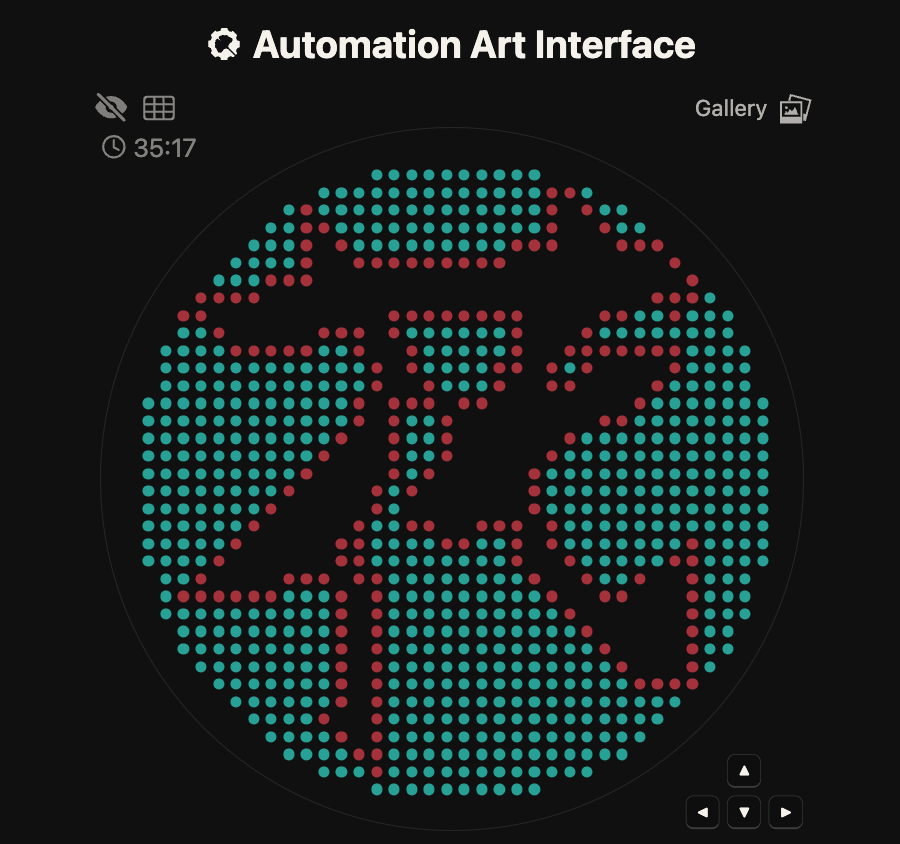
This is the character for “flower,” written by my late grandmother. She did not have access to education, so she taught herself to read and write after she was over 80 years old. These “flower” characters were practiced during that time. After she passed away, these practice writings were discovered. I want to bring my grandmother’s handwriting back to life. Last year when I participated in HTGAA, I had the idea but was unable to realize it, so this year I would like to make it happen.

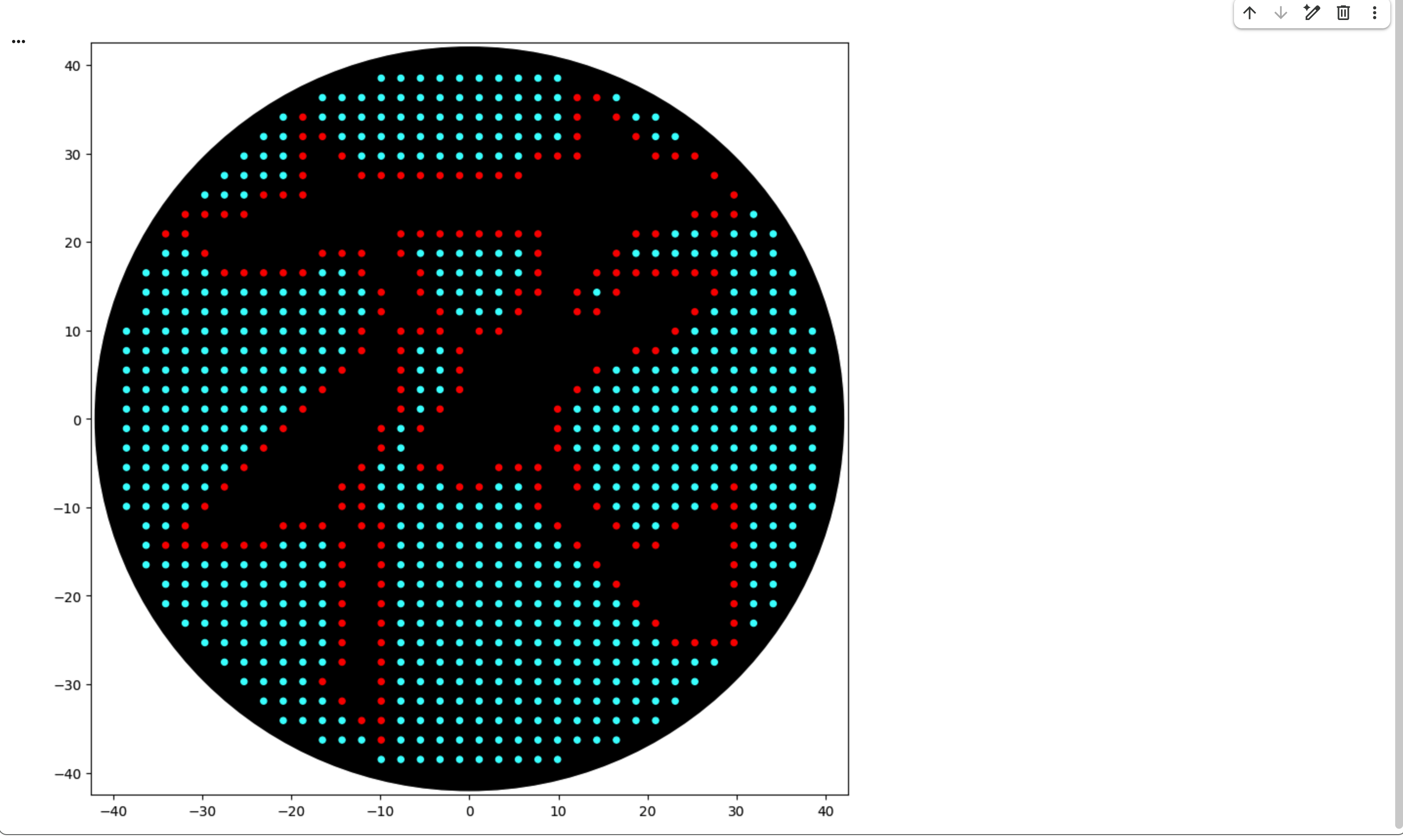
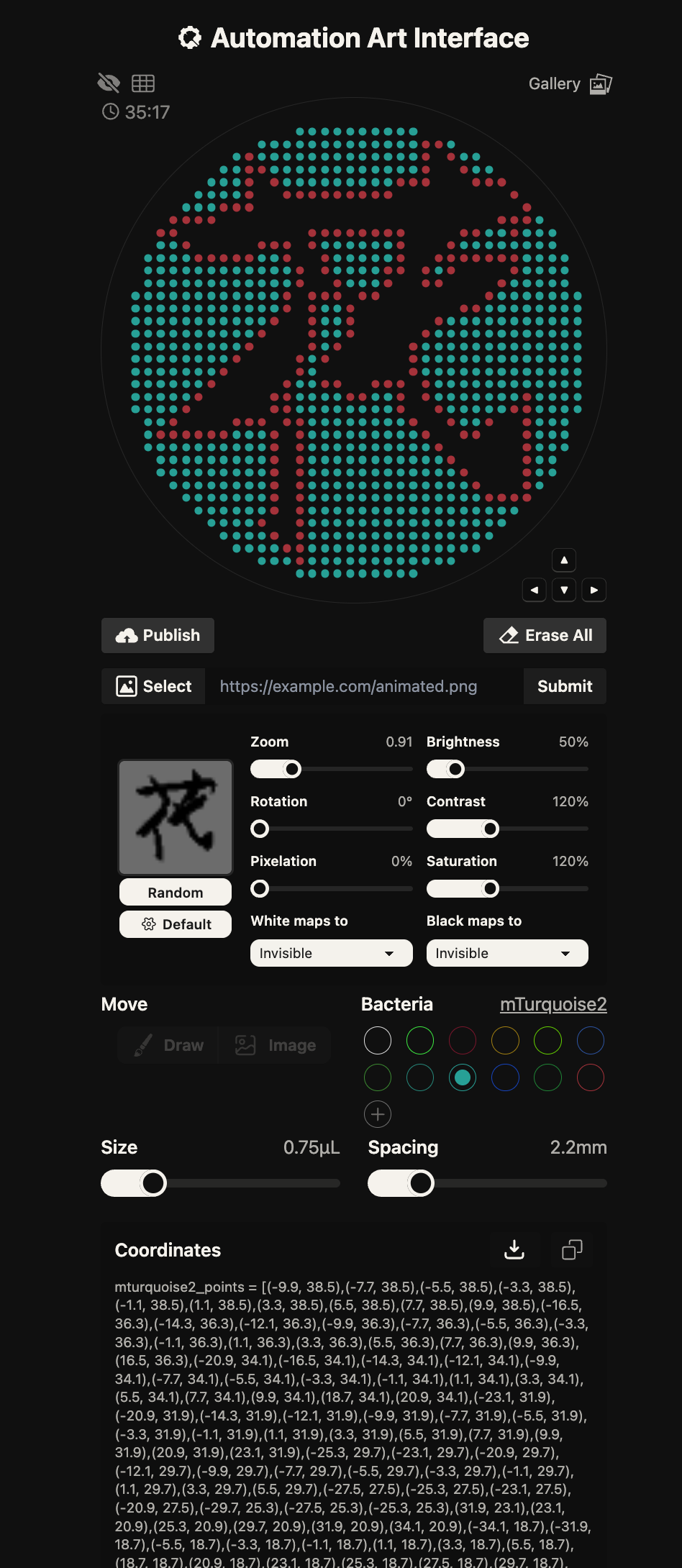
First, I created my design using the Automation Art Interface. However, even after publishing it to the gallery, clicking on my own work did not display it properly. Therefore, I copied the coordinates from the “Coordinates” section and used them in Colab.
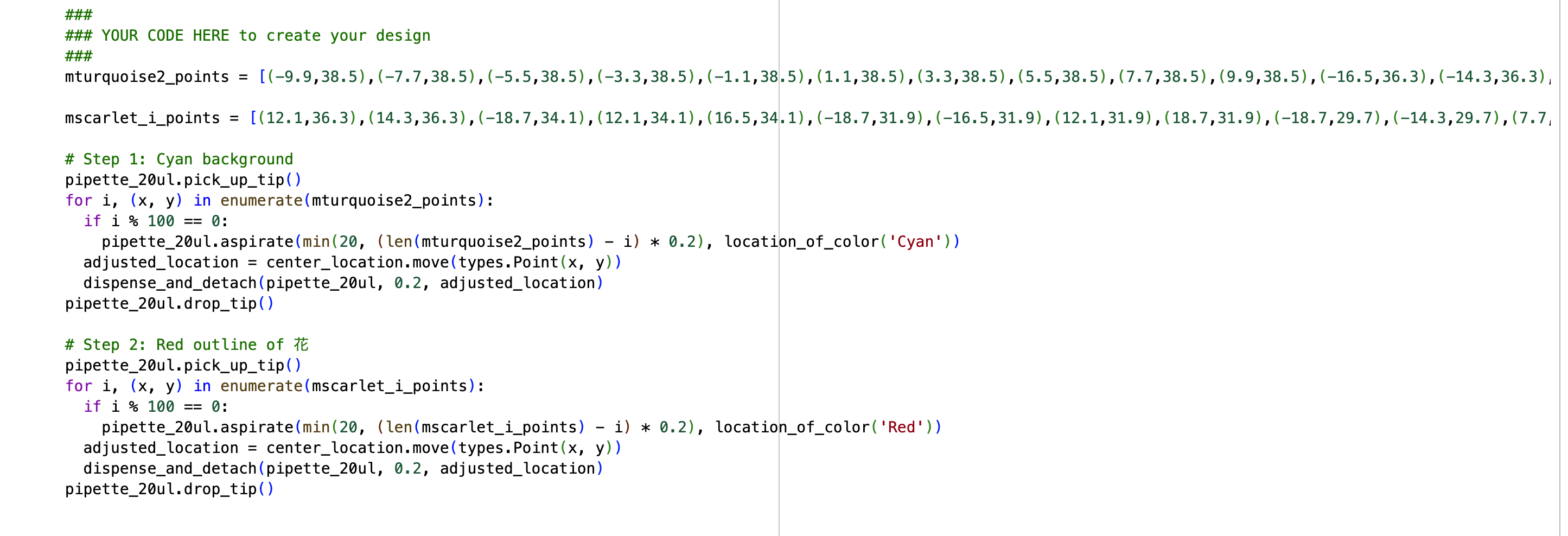
In the “your code” section, Cyan and Blue were not defined in the well_colors dictionary. However, since they were listed as available colors, I added them manually by defining D1 as Cyan and E1 as Blue.
[https://opentrons-art.rcdonovan.com/?id=6o741ir7g0gj0ri]

Post-Lab Questions
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely.
For this week, we’d like for you to do the following:
1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
- AssemblyTron: flexible automation of DNA assembly with Opentrons OT-2 lab robots
Author: John A Bryant Jr 1, Mason Kellinger 2, Cameron Longmire 3, Ryan Miller 4, R Clay Wright 5,
Published: 22 December 2022
[https://doi.org/10.1093/synbio/ysac032]
この論文は、j5というDNAアセンブリ設計ソフトウェアとOpentrons OT-2液体ハンドリングロボットを統合したオープンソースのPythonパッケージ「AssemblyTron」を紹介している。
複数のDNA組み立て手法(PCR、Golden Gate assembly、相同性依存型IVAなど)を、自動化し、手動と同等の精度まで成功したことが実証された。
AssemblyTronは合成生物学における時間・コスト・廃棄物を削減でき、合成生物学をより広かれることで、生物学研究そのものを加速させる可能性があると言える。
This paper introduces “AssemblyTron,” an open-source Python package that integrates j5 DNA assembly design software with the Opentrons OT-2 liquid handling robot. It was demonstrated that multiple DNA assembly methods — including PCR, Golden Gate assembly, and homology-dependent IVA — were successfully automated, achieving accuracy comparable to manual methods.
AssemblyTron has the potential to reduce time, cost, and waste in synthetic biology, and by making synthetic biology more widely accessible, it could accelerate biological research itself.
2. Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
While your description/project idea doesn’t need to be set in stone, we would like to see core details of what you would automate. This is due at the start of lecture and does not need to be tested on the Opentrons yet.
Example 1: You are creating a custom fabric, and want to deposit art onto specific parts that need to be intertwined in odd ways. You can design a 3D printed holder to attach this fabric to it, and be able to deposit bio art on top. Check out the Opentrons 3D Printing Directory.
Example 2: You are using the cloud laboratory to screen an array of biosensor constructs that you design, synthesize, and express using cell-free protein synthesis.
(1). Echo transfer biosensor constructs and any required cofactors into specified wells.
(2). Bravo stamp in CPFS reagent master mix into all wells of a 96-well / 384-well plate.
(3). Multiflo dispense the CFPS lysate to all wells to start protein expression.
(4). PlateLoc seal the plate.
(5). Inheco incubate the plate at 37°C while the biosensor proteins are synthesized.
(6). XPeel remove the seal.
(7). PHERAstar measure fluorescence to compare biosensor responses.
Final Project Ideas
As explained in this week’s recitation, add 1-3 slides with 3 ideas you have for an Individual Final Project in the appropriate slide deck for MIT/Harvard/Wellesley students or for Commited Listeners. Be sure to put your name on your slide(s); for CLs, also put your city and country on your slide(s) and be sure you’re putting your slide(s) in your Node’s section of the deck.




