Week 04 HW: Protein Design Part I
‘Week 4 HW: Protein Design Part I’
Documentation
Homework: Protein Design I — DUE BY START OF MAR 3 LECTURE
Part A. Conceptual Questions
Answer any NINE of the following questions from Shuguang Zhang: (i.e. you can select two to skip):
☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆☆
1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
(500gの肉に含まれるアミノ酸分子の数は?)
First, assuming this meat is beef, we calculate the protein content in 500 g of beef
(まず、この肉を牛肉と仮定して、牛肉500gに含まれるタンパク質の量を計算する)
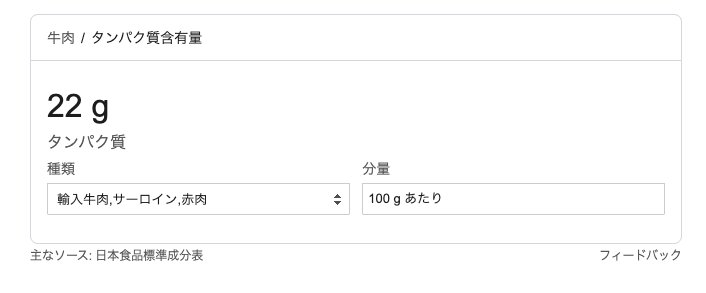



100gあたり約22gのタンパク質とすると、
There are about 22 g of protein per 100 g, so for 500 g, that becomes 110 g of protein
500g × 0.22 = 110g
平均アミノ酸 1分子 ≈ 100 Dalton
Although some references suggest an average amino acid mass of ~110 Daltons, I follow the problem statement (~100 Daltons) for this calculation.
1 Dalton ≈ 1 g/mol
100 Dalton ≈ 100 g/mol
110g / 100g/mol = 1.1 mol
アボガドロ定数:物質1モル(mol)中に含まれる原子や分子などの粒子数で、その値は正確に 6.02214076 *10²³ molecules/mol である。
Avogadro constant: The number of particles, such as atoms or molecules, contained in one mole (mol) of a substance. Its value is exactly
particles/mol.
1.1 mol × 6.02 × 10²³ molecules/mol ≈ 6.6 × 10²³ molecules
つまり、 6.6 × 10²³ molecules 個のアミノ酸分子

2. Why do humans eat beef but do not become a cow, eat fish but do not become fish?
(なぜヒトは牛肉を食べても牛にはならず、魚を食べても魚にはならないのか)
https://bio-sta.jp/biokids/chapter1/section7/
Quora: “Why don’t I turn into a fish when I eat fish or a cow when I eat beef? There are chemicals in our bodies that prevent foreign genetic material in a cell from being read and translated into mRNA. What are these chemical processes and how do they work?”
Digestion (消化)
牛のタンパク質は完全に分解され、アミノ酸になり、もとの構造情報(配列情報)は失われる
Beef proteins are completely broken down into amino acids; their original structural information is lost.
ペプシン・トリプシン・キモトリプシンで分解 個々のアミノ酸・小ペプチド
Broken down by proteases such as pepsin, trypsin, and chymotrypsin Individual amino acids and small peptides
↓
Absorption(吸収)
アミノ酸は「設計図」ではなく、分子レベルの「材料」として吸収される(小腸)
Amino acids are absorbed as molecular “building blocks,” not as blueprints.(absorbed in the small intestine)
↓
Synthesis(合成)
ゲノム(DNA)が設計図として機能し、ヒトのタンパク質として合成を指示する
DNA → mRNA → ribosomes
Resynthesized as human proteins
牛肉を摂取すると、牛のタンパク質は胃や消化酵素によって分解され、アミノ酸になる。この過程で、もとのアミノ酸配列という構造情報は失われる。 その後、アミノ酸は小腸で吸収され、血液中へと取り込まれる。 吸収されるのは「牛のタンパク質」ではなく、あくまでタンパク質を構成する材料。そして最終的に、ヒトのDNAが設計図として働き、これらのアミノ酸を用いてヒトのタンパク質が新たに合成される。よって、牛を食べてもヒトは牛にはならない。
When we consume beef, the cow’s proteins are broken down by gastric acid and digestive enzymes into individual amino acids. During this process, the original structural information—the amino acid sequence—is lost.
The amino acids are then absorbed in the small intestine and transported into the bloodstream. What is absorbed is not “cow protein” itself, but merely the molecular building blocks that make up proteins. Finally, human DNA functions as the blueprint, directing the synthesis of new human proteins from these amino acids. Therefore, eating beef does not turn a human into a cow.
3. Why are there only 20 natural amino acids?
(なぜ天然のアミノ酸は20種類しかないのか?)
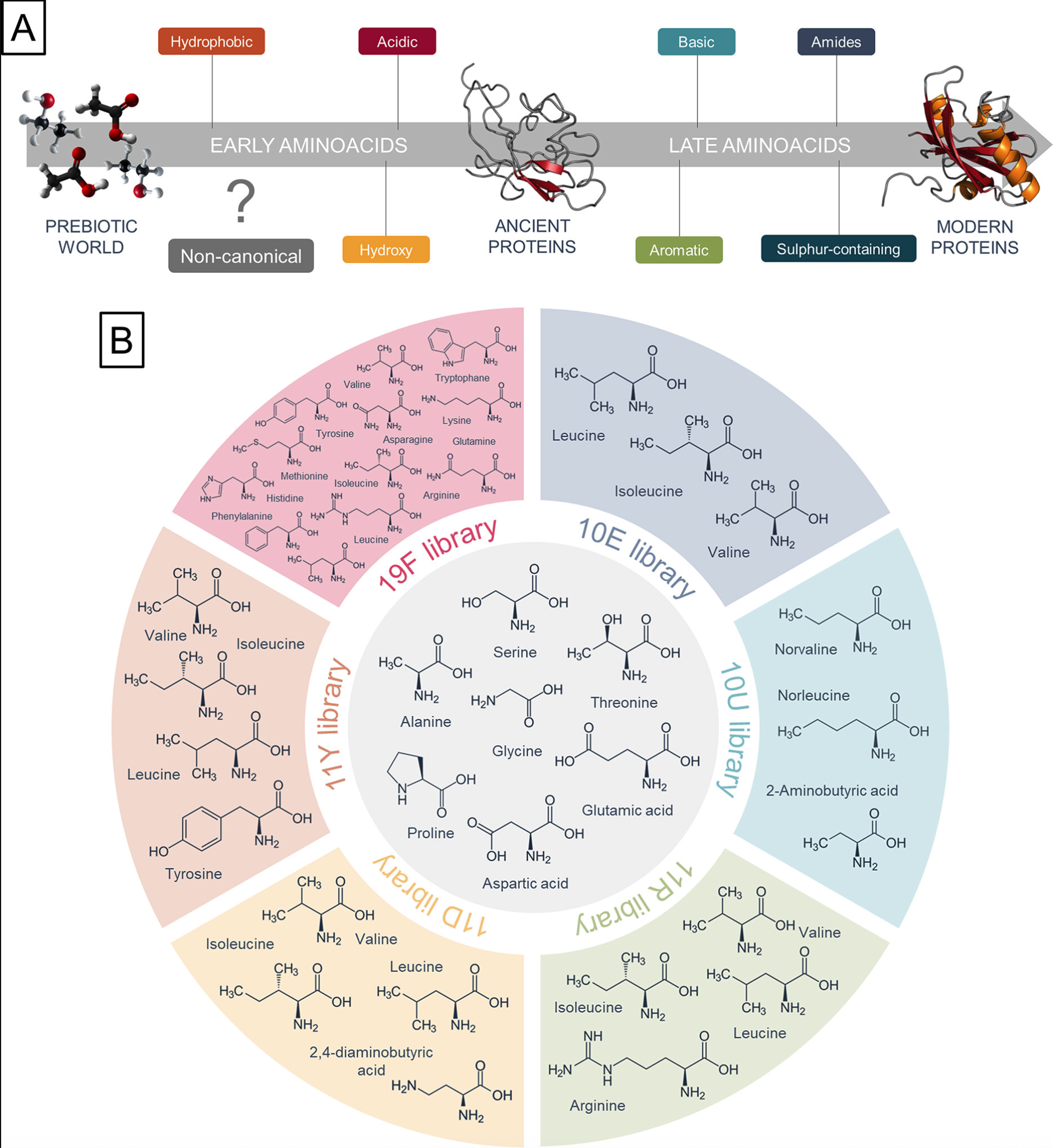
(A) Assumed early and late stages of amino acid alphabet incorporation during protein evolution and (B) design of peptide libraries based on this order(from Journal of the American Chemical Society Cite this: J. Am. Chem. Soc. 2023, 145, 9, 5320–5329)
From the above information, we can conclude:
01 Early Amino Acids (10) vs. Late Amino Acids (10)
原始地球の環境では、グリシン(Gly)やアラニン(Ala)などの「初期アミノ酸」は構造が単純で生成しやすく、自然界に豊富に存在していたと考えられている。
その後の進化の過程で、フェニルアラニン(Phe)やトリプトファン(Trp)のような、より複雑な「後期アミノ酸」が生合成経路を通じて取り込まれ、
最終的に約20種類の標準的なアミノ酸の集合へと収束した。
In the primordial environment, certain “early amino acids” (such as Gly and Ala), which were simpler and easier to form, were naturally abundant. Later in evolution, more complex “late amino acids” (e.g., Phe, Trp) were incorporated via biosynthetic pathways, eventually converging on a standard set of about 20 amino acids.
02 Optimization for Protein Folding
ジョンズ・ホプキンス大学の Steven Fried らによる研究では、機能的な球状タンパク質へと折りたたまれる能力(foldability)が、数多くの非タンパク質性アミノ酸の中から現在 の20種類が選ばれた重要な要因であった可能性が示唆されている。適切なアミノ酸の組み合わせは、より優れたタンパク質の折りたたみと機能をもたらした。
Research led by Steven Fried at Johns Hopkins University suggests that the ability to fold into functional globular proteins (i.e., “foldability”) was a critical factor in why these particular 20 amino acids were chosen over countless non-proteinogenic alternatives.
The right blend of amino acids provided superior folding and functional performance.
03 Evolutionary Selection and Elimination
生命が正式に誕生する以前であっても、タンパク質形成のレベルで原始的な「自然選択」が起こっていた可能性がある。
タンパク質の折りたたみや触媒機能に有利なアミノ酸は残り、あまり有効でないものは自然に淘汰されていったと考えられる。
Even before the formal emergence of life, primitive “natural selection” may have taken place at the level of protein formation.
Amino acids beneficial for folding and catalytic functions were retained, while less effective ones were naturally discarded.
04 From GNC Code → SNS Code → Universal Genetic Code
ある仮説では、最も初期の遺伝暗号はグリシンやアラニンなど、ごく少数のアミノ酸しか扱わなかったとされている。その後、性能を高めるアミノ酸が一つずつ追加され、現在の20種類 のアミノ酸の体系へと発展した。後から追加されたアミノ酸は、より複雑な側鎖を持ち、タンパク質の機能的多様性や安定性を高める役割を果たしている。
One hypothesis is that the earliest genetic code involved only a few amino acids (like Gly, Ala) .
Over time, performance-enhancing amino acids were added one by one, leading to today’s repertoire of 20.
Late-added amino acids have more complex side chains and contribute greater functional variety and stability to proteins.
05 Exceptions (21st, 22nd, etc.)
一部の生物では、セレノシステイン(21番目)やピロリジン(22番目)といった追加のアミノ酸が利用されている。
しかし、これらは特別な翻訳機構を必要とする例外的な存在であり、標準的な20種類のアミノ酸体系の外に位置づけられている。
A few organisms utilize selenocysteine (21st) or pyrrolysine (22nd), but these require specialized translational machinery
They remain exceptions outside the standard 20.
06 Why Non-Proteinogenic Amino Acids Were Not Adopted
分岐していないアルキル鎖を持つアミノ酸など、一部の非タンパク質性アミノ酸は、前生物的環境では豊富に存在していた可能性がある。
しかし、タンパク質の折りたたみや分子間相互作用にとって不利であったため、生命のタンパク質構成要素としては適さなかったと考えられる。
その結果、現在の20種類のアミノ酸の集合は、生命にとって非常に効率的でバランスの取れた構成であると見なされている。
Some non-proteinogenic amino acids—such as those with unbranched alkyl chains—might have been abundantly available prebiotically, but proved detrimental for folding or interaction, making them poor choices.
The set of 20 is viewed as a highly effective balance for life’s needs.
初期地球では、タンパク質の折りたたみ能力に優れたアミノ酸を選び出すような、自然選択と化学的な選別が起こっていたと考えられる。 生命が誕生し進化する過程で、いくつかの「高性能」なアミノ酸が追加され、最終的に20種類の標準的アミノ酸(加えて例外的な21番目・22番目)に至った。 この最終的なセットは、生物が機能するために必要な性質のバランスを最もよく満たす組み合わせであった可能性が高い。
In summary, natural selection and chemical filtering took place on the early Earth, favoring amino acids that excelled at protein folding. Once life arose and evolved, several “high-performance” amino acids were added, culminating in the total of 20 standard amino acids (plus the exceptional 21st and 22nd). This final set likely struck the optimal balance of functionality for living organisms.
4. Can you make other non-natural amino acids? Design some new amino acids.
(自然界に存在しない非天然アミノ酸を作ることは可能でしょうか。また、いくつか新しいアミノ酸を設計してみてください。)
[https://www.nature.com/articles/nature24031]
[https://www.nature.com/articles/nchembio.203]
自然界に存在しない非天然アミノ酸は作れる。アミノ酸は共通の骨格(NH2-CH-COOH)を持ち、側鎖(R基)を変えることで新しいアミノ酸が作れるからである。そのため、蛍光基、光反応基、金属結合基などを持つ新しいアミノ酸を人工的に設計できる。
Yes, it is possible to design non-natural amino acids.All amino acids share the same core structure (NH₂–CH–COOH) but differ in their side chain (R group). By modifying the side chain, scientists can create new amino acids with novel chemical properties. For example, one could design amino acids with fluorescent groups, photo-reactive groups, or metal-binding groups, expanding the functional diversity of proteins.
1 Fluorescent Amino Acid
通常のアミノ酸骨格にR基に蛍光分子をつける。→タンパク質を光らせる
R-group: a fluorescent moiety
Function: This amino acid would allow proteins to emit light, enabling visualization of protein location and dynamics in living cells.
2 Magnetic metal-binding amino acid (磁性金属結合アミノ酸): 金属結合させて磁性を形成するタンパク質
鉄(Fe)やコバルト(Co)イオンと結合できる配位子(リガンド)を持つ側鎖 →このアミノ酸は金属イオンを配位して結合することができ、それにより、タンパク質が磁性中心や触媒中心を形成できるようになる可能性がある
binds “Fe” or “Co” ions
R-group: a ligand capable of binding iron or cobalt ions
Function: This amino acid could coordinate metal ions, enabling proteins to form magnetic or catalytic centers.
5. Where did amino acids come from before enzymes that make them, and before life started?
(生命やアミノ酸合成酵素が存在する以前、アミノ酸はどのようにして生まれたのか?)
生命の起源となったアミノ酸は、宇宙から隕石に乗ってやってきたのか?
[https://wired.jp/article/did-the-seeds-of-life-ride-to-earth-inside-an-asteroid/]
生命のもとは、暗闇ででたらめに生まれた(ユーリ・ミラー)
[https://lne.st/2016/01/14/the-origin-of-life/]
隕石衝突でアミノ酸生成 生命誕生に手掛かり 東北大など実証
[https://www.sankei.com/article/20200614-CZWXTGRD3BN57HKSS7KAQTC4AY/]
Based on the references above, there are several possibilities regarding where amino acids came from before life and its enzymes existed:
Meteorite/Interstellar Cloud Origin
It is widely known that meteorites contain various amino acids. During the early formation of the solar system, small bodies known as “asteroids” could have delivered amino acids to Earth upon impact. In interstellar clouds, ice and gas may have generated amino acids when exposed to ultraviolet or cosmic rays, which were then brought to Earth via these “cosmic time capsules” (i.e., meteorites). In the near future, uncontaminated samples returned from asteroids Bennu and Ryugu are expected to shed more light on this scenario.
Natural Synthesis on Earth
Experiments like the Miller-Urey experiment and scenarios involving deep-sea hydrothermal vents point to the possibility that amino acids could have been produced right here on Earth. Factors such as lightning, volcanic activity, temperature, and pressure may have triggered organic molecules to form and accumulate into the amino acids that eventually led to proteins.
A Combined, Multistep Process
It is also possible that amino acids came from both meteorites and Earth-based synthesis, mixing to form a “primordial soup” . Multiple routes—asteroid impacts, hydrothermal vents, and interstellar ices—could have simultaneously supplied amino acids to Earth, thereby facilitating the emergence of life.
生命が誕生する前のアミノ酸は
・原始地球の化学反応
・宇宙由来の有機分子
・深海の化学反応
などによって作られたと考えられる。
Before life began, amino acids are thought to have been formed through several processes, including chemical reactions on the early Earth, organic molecules delivered from space, and chemical reactions occurring in deep-sea environments such as hydrothermal vents.
6. If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
(D型アミノ酸でαヘリックスを作った場合、そのらせんは右巻きと左巻きのどちらになると予想されるか?)
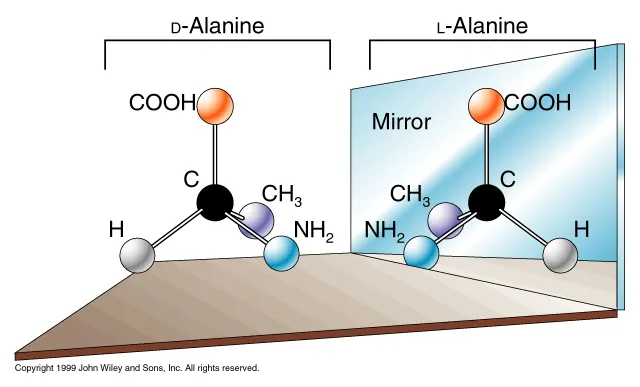
from “Biological Homochirality: One of Life’s Greatest Mysteries”
Alpha helix αヘリックス
[https://en.wikipedia.org/wiki/Alpha_helix]
[https://ja.wikipedia.org/wiki/%CE%91%E3%83%98%E3%83%AA%E3%83%83%E3%82%AF%E3%82%B9]
[https://www.nature.com/scitable/topicpage/protein-structure-14122136/]
“The amino acids in an α-helix are arranged in a right-handed helical structure.”
Science Direct “Right-Handed Alpha-Helix”
[https://www.sciencedirect.com/topics/chemistry/right-handed-alpha-helix]
L-amino acids can form only right-handed α-helices in protein structures.
National Library of Medicine “A mixed chirality α-helix in a stapled bicyclic and a linear antimicrobial peptide revealed by X-ray crystallography”
[https://pmc.ncbi.nlm.nih.gov/articles/PMC8637766/]
L-chiral amino acids form right-handed helices,
D-chiral amino acids form left-handed helices.
Natural proteins are composed almost entirely of L-amino acids, and these amino acids preferentially form right-handed α-helices due to their stereochemistry.
7. Can you discover additional helices in proteins?
(タンパク質の中に、さらに別のヘリックス構造を発見することは可能か?)
Nature communications “Exo-chirality of the α-helix”
[https://www.nature.com/articles/s41467-024-51072-8]
最近の研究(例えば「αヘリックスの外部ヘリカル対称性(Exo-helical symmetries of the α-helix)」に関する研究)では、これまでαヘリックスの「主鎖ヘリックス(main-chain helix)」として一括りにされてきた構造に、周期的な側鎖パターンによって形成される 「外部ヘリックス(exo-helix)」 が伴っている可能性が示されている。
この発見は、以下のような理由から、タンパク質の中にはまだ 「未発見のヘリックス構造」 が存在している可能性を示唆しています。
Recent studies (such as the one on “Exo-helical symmetries of the α-helix”) indicate that what was previously grouped together as the “main-chain helix” of an α-helix may, in fact, be accompanied by an “external helix (exo-helix)” formed through periodic side-chain patterns This discovery suggests that there may still be “unidentified helices” hidden in proteins, for several reasons:
Helices beyond the α-helix main chain:
Traditionally, attention has been focused on the rotation angle and hydrogen bonding of the α-helix backbone
However, it has recently been shown that side chains, which follow periodic repeat patterns (i, i + x), can assume a different helical symmetry (exo-helical symmetry)
Theoretical and spectroscopic confirmation:
In the cited work, the authors integrated NBD (nitrobenzoxadiazole) chromophores into peptide side chains, then employed CD spectroscopy and molecular dynamics simulations to demonstrate the presence of an exo-helix
In other words, even though the backbone forms the same α-helix, periodic side chains can produce a separate, outward-facing helix that may be right-handed or left-handed
A framework for finding “additional helices”:
This exo-helix concept implies that, even in existing PDB structures, certain local or partial helical features might have gone unnoticed, depending on computational/experimental conditions
In effect, alongside the “standard secondary structure,” there may be a “hidden external helix”—an entirely new perspective that strongly suggests there is still ample room to discover unidentified helical motifs in proteins
Potential:
The exo-helix concept could be valuable for protein design and the creation of functional synthetic peptides.
It points to a new direction in which “additional helices” could significantly affect molecular interactions and structural conformations
8. Why are most molecular helices right-handed?
(なぜ多くの分子ヘリックス(らせん構造)は右巻きなのか?)
・“Why are α-helices in proteins mostly right handed?
[https://www.ch.ic.ac.uk/rzepa/blog/?p=3802]
・Alpha Helix “Bioinformatics: Concepts, Methods, and Data”
[https://www.sciencedirect.com/topics/medicine-and-dentistry/alpha-helix]
- YouTube Erik Lindahl
“Lecture 05, concept 08: The alpha helix is right-handed due to L amino acids
"
[https://www.youtube.com/watch?v=rdkXOxLHDws]
ほとんどすべての生物のタンパク質は、L型アミノ酸(「左手型」の形)のみから構成されているため、αヘリックスは自然に右巻き構造をとる。
理論的には、もし**D型アミノ酸(L型アミノ酸の鏡像)**を使用すれば、左巻きのαヘリックスを作ることも可能である。
しかし、生物は主にL型アミノ酸を使用するように進化しており(そのキラリティが固定されているため)、この条件下で形成されるヘリックスは結果として右巻きになる。
Because virtually all biological proteins are built exclusively from L-amino acids (the “left-handed” form), α-helices naturally adopt a right-handed conformation.
Theoretically, if one were to use D-amino acids (the mirror image of L-amino acids), left-handed α-helices would be possible.
However, since living organisms predominantly chose L-amino acids (their chirality is fixed), the helices formed under these conditions end up being right-handed.
Ramachandran Plot Stability
ラマチャンドランプロットにおける安定性
L型アミノ酸は、ラマチャンドランプロットの中で右巻きαヘリックスの領域に安定して収まる。
そのため、この構造はエネルギー的にも立体的にも有利である。
L-amino acids fit securely into the Ramachandran plot’s region for right-handed α-helices, making this conformation energetically and sterically favorable.
Mirror-Image (Chirality) Influence 鏡像関係(キラリティ)の影響
物理法則そのものが、右巻きか左巻きかのどちらかを必ず選ばせるわけではない。
しかし、生命で使われているアミノ酸はすべてL型(特定のキラリティ)であるため、それらを組み合わせると結果として右巻きのヘリックスが形成される。
Physics itself does not mandate a preference for right- or left-handed forms. But because the amino acids in life are all L-type (with a specific chirality), assembling them leads to a right-handed helix.
Left-Handed Helices Are Rare in Practice
左巻きヘリックスは実際には非常にまれ
理論的には、D型アミノ酸を使えば左巻きαヘリックスが形成される可能性がある。しかし、このような構造は自然のタンパク質では非常にまれである。そのため、実際には「αヘリックス」と言うと、通常は右巻きのヘリックスを指す。
Although D-amino acids would theoretically yield left-handed α-helices, these are extremely uncommon in natural proteins
Hence, for all practical purposes, “α-helix” refers to the right-handed version.
9. Why do β-sheets tend to aggregate?
・What is the driving force for β-sheet aggregation?
(なぜβシートは凝集(集まる)しやすいのか? βシート凝集の駆動力は何か?)
“Natural β-sheet proteins use negative design to avoid edge-to-edge aggregation” [https://pmc.ncbi.nlm.nih.gov/articles/PMC122420/#:~:text=In%20hindsight%2C%20it%20seems%20obvious,other%20%CE%B2%20strands%20they%20encounter]
From the above cited paper, the following points can be understood:
Exposed “Edge” Strands
βシートでは、それぞれのストランドが主鎖の水素結合によって整列している。
しかし、端(エッジ)に位置するストランドは、近くにある別のβストランドと追加の水素結合を形成できる状態のまま残る。
そのため、これらのエッジが保護されていない場合、複数のβシートが積み重なったり結合したりして、大きな凝集体を形成する可能性がある。
In a β-sheet, each strand aligns via hydrogen bonds along its backbone.
However, the edge strands are still available to form additional hydrogen bonds with any nearby β-strand.
Consequently, if those edges remain unprotected, multiple β-sheets can stack or merge, leading to large aggregates.
Natural “Edge Protection” in Proteins
多くの天然のβシートタンパク質では、制御されない凝集を防ぐためにさまざまな設計上の工夫が存在する。
例:
・ループ構造がシートの端を覆う
・プロリン残基や「バルジ(bulge)」を挿入して、規則的なβ構造をわずかに歪ませる
・内側に向いた帯電残基を配置する
・βバレルのような閉じた構造を形成し、露出したエッジをなくす
一方で、このような保護機構を持たない短いβシート断片や、人工的に設計されたβタンパク質は、容易に凝集して不溶性の繊維(ファイバー)を形成しやすくなる。
・Many native β-sheet proteins avoid uncontrolled aggregation by employing various design strategies:
loops covering the edges, inserted prolines or “bulges” that distort regular β-structure, charged residues pointing inward, or forming closed barrels so that no true edges remain exposed.
・In contrast, short β-sheet fragments or artificially designed β-proteins without these protective features readily aggregate into insoluble fibers.
Driving Forces of Aggregation 凝集の駆動力
Extended Hydrogen-Bond Network 水素結合ネットワークの拡張:
When an edge strand meets another β-strand, a series of hydrogen bonds form, creating a significant energetic gain that propels aggregation βシートの端にあるストランドが別のβストランドと接触すると、一連の水素結合が形成される。このときエネルギー的に大きな安定化が得られるため、凝集が進みやすくなる。
Hydrophobic Interactions 疎水性相互作用:
Clustering β-sheets often bury hydrophobic side chains, lowering the system’s free energy and favoring aggregation
βシートが集まると、疎水性の側鎖が内部に埋もれるように配置される。これによって系の自由エネルギーが低下し、凝集がより起こりやすくなる。
Other Interactions その他の相互作用:
Electrostatic and van der Waals (dispersion) forces can further stabilize the growing assembly, making β-sheet clusters more likely to form
静電相互作用やファンデルワールス力(分散力)も、形成されつつある集合体をさらに安定化させる。その結果、βシートのクラスターは形成されやすくなる。
10. Why do many amyloid diseases form β-sheets?
・Can you use amyloid β-sheets as materials? (なぜ多くのアミロイド病ではβシート構造が形成されるのか? また、アミロイドβシートを材料として利用することはできるのか?)
アミロイドーシスに関する調査研究班
[http://amyloidosis-research-committee.jp/about/]
What Are Amyloid and Amyloidosis? アミロイドとアミロイドーシスとは何か
アミロイドとは、体内のタンパク質の形や性質が変化することで生じる物質であり、その結果タンパク質が集合して、水や血液に溶けにくい繊維状の沈着物を形成したものを指す。
このアミロイドが体内の組織に蓄積する病気は、総称して アミロイドーシス(amyloidosis) と呼ばれる。
Amyloid is a substance formed when the shape and properties of proteins in our bodies change, causing them to aggregate into fibrous deposits that are poorly soluble in water or blood. Diseases in which amyloid accumulates within the body’s tissues are collectively referred to as amyloidosis.
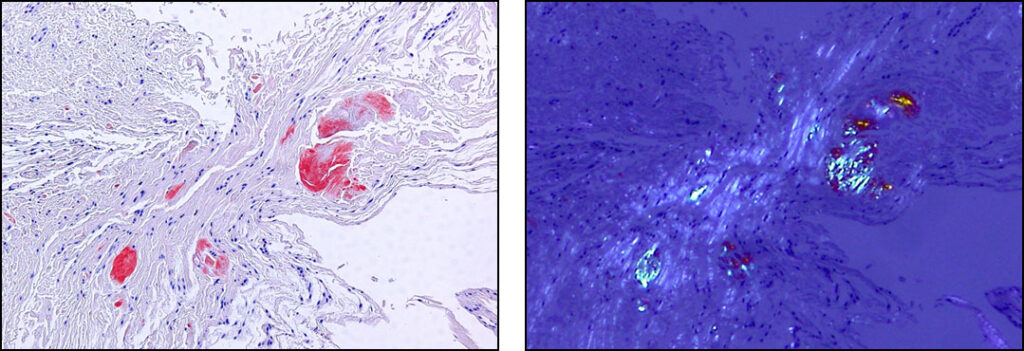
Microscopic Images of Amyloid Deposited in Tissue (Biopsy of the Carpal Ligament)
- In the left image, the red-stained area is amyloid (stained with Congo red)
- In the right image, when viewed under polarized light, the amyloid fluoresces with an apple-green color
Structure “Biology of Amyloid: Structure, Function, and Regulation”
[https://www.sciencedirect.com/science/article/pii/S0969212610003084]
Amyloid fibrils are fibrous structures composed of repeatedly aligned “cross-β sheet” arrangements, and they have long been linked to numerous diseases including Alzheimer’s disease (AD), Parkinson’s disease, and prion diseases.
More recently, researchers have discovered “functional amyloids” that carry out physiological roles despite having a similar structural basis.
The fundamental architecture of amyloid fibrils consists of protein chains extended and stacked in a regular cross-β configuration.
This fibrous arrangement is highly stable, so once protein misfolding initiates, it can recruit numerous identical molecules and rapidly grow into large aggregates.
Consequently, it is almost inevitable that the amyloid deposits observed in many neurodegenerative disorders (e.g., Alzheimer’s and Parkinson’s) take the form of β sheet–based fibrils.
Thus, in response to the question “Why do amyloid diseases form β sheets?”, one can argue that misfolded proteins adopt stable cross-β fibrils, leading to extensive aggregation characteristic of these conditions
アミロイド線維(amyloid fibrils)は、繰り返し整列した 「クロスβシート(cross-β sheet)」 配列から構成される繊維状構造であり、長い間、アルツハイマー病(AD)、パーキンソン病、プリオン病など多くの疾患と関連していることが知られている。
近年では、同様の構造的基盤を持ちながらも、生理的な役割を果たす 「機能性アミロイド(functional amyloids)」 の存在も発見されている。
アミロイド線維の基本構造は、タンパク質鎖が伸びた状態で規則的に積み重なった クロスβ構造 によって形成されている。
この繊維状の配置は非常に安定しており、一度タンパク質の誤った折りたたみ(ミスフォールディング)が始まると、同じ分子を次々と取り込みながら急速に大きな凝集体へと成長する。
その結果、アルツハイマー病やパーキンソン病など多くの神経変性疾患で観察されるアミロイド沈着は、ほぼ必然的に βシートを基盤とする線維構造をとる。
したがって、「なぜアミロイド病ではβシートが形成されるのか?」という問いに対しては、誤って折りたたまれたタンパク質が安定なクロスβ線維構造をとり、その結果として広範な凝集が生じるためであると説明することができる。
・Can you use amyloid β-sheets as materials?
PMC “The relationship between amyloid structure and cytotoxicity”
https://pmc.ncbi.nlm.nih.gov/articles/PMC4189889/
According to the link above, it is stated that the following can be utilized as materials
Tissue Repair and Cell Culture Scaffolds 組織修復および細胞培養用スキャフォールド
Utilizing amyloid fibrils as a scaffold can promote cell adhesion and proliferation via interactions with the cell membrane, making them highly effective in tissue engineering.
For example, composite hydrogels made from aloe vera and bovine serum albumin (BSA) have demonstrated strong wound-healing capabilities, and there are applications that combine 3D printing with bio-inks.
アミロイド線維をスキャフォールド(足場材料)として利用すると、細胞膜との相互作用によって細胞の接着や増殖を促進することができまる。そのため、組織工学の分野で非常に有効。
例:アロエベラとウシ血清アルブミン(BSA)から作られた複合ハイドロゲルは高い創傷治癒能力を示唆。3Dプリンティングとバイオインクを組み合わせた応用も報告されている。
Drug Delivery
The fibrous structure of amyloid can transport hydrophobic or positively charged molecules, which has led to its use in drug delivery systems. Reports include cases where methylene blue or riboflavin are effectively carried using amyloid-based materials.
アミロイドの繊維構造は、疎水性分子や正電荷を持つ分子を運搬する能力がある。 そのため、ドラッグデリバリーシステムへの応用が研究されている。 例:メチレンブルーやリボフラビンをアミロイド材料によって効率よく運搬できる。
Macroscale Fibers and Composites マクロスケール繊維および複合材料
Thanks to their mechanical strength and chemical stability, amyloid fibrils can be formed into hydrogels, macro-fibers, composite materials, and sensors. In addition, by incorporating conductive peptides, researchers aim to develop biodegradable and biocompatible conductive materials.
アミロイド線維は、機械的強度と化学的安定性が高い。
- ハイドロゲル
- マクロファイバー
- 複合材料
- センサー材料
などに加工することが可能
さらに、導電性ペプチドを組み込むことで、生分解性かつ生体適合性を持つ導電材料の開発も。
Amyloid as a Catalytic Material 触媒材料としてのアミロイド
Esterase Activity Enhancement エステラーゼ活性の向上
Introducing residues such as tyrosine or histidine within the amyloid fibril can create a microenvironment that displays esterase-like activity. For instance, peptide fibers containing histidine or hydrophobic side chains (e.g., leucine, isoleucine) can form reversible hydrogels with significant catalytic potential.
アミロイド線維の内部にチロシンやヒスチジンなどの残基を導入すると、エステラーゼ様活性を示す微小環境を形成することができる。
例えば、ヒスチジンや疎水性側鎖(ロイシン、イソロイシンなど)を含むペプチド線維は、可逆的なハイドロゲルを形成し、優れた触媒活性を示す可能性がある。
11. Design a β-sheet motif that forms a well-ordered structure.
(秩序だった構造を形成するβシートモチーフを設計しなさい。)
条件:
- Val (V) は βシートを作りやすい
- Thr (T) は極性があって並びを整えやすい
- 疎水性 / 親水性 が交互なのは、βストランドで良い
This motif has alternating hydrophobic and polar residues, which is favorable for β-sheet formation. Valine promotes β-sheet stability, and threonine helps create an ordered structure.
Part B. Protein Analysis and Visualization
In this part of the homework, you will be using online resources and 3D visualization software to answer questions about proteins. Pick any protein (from any organism) of your interest that has a 3D structure and answer the following questions:
I chose Green Fluorescent Protein (GFP) because I am interested in biological light emission and how light-producing mechanisms in living organisms work. GFP, originally derived from the jellyfish Aequorea victoria, absorbs light in the ultraviolet to blue range and emits green fluorescence. This unique property has made GFP widely used as a molecular marker in biological research to visualize proteins and cellular processes.
UniProt “P42212 · GFP_AEQVI”
[https://www.uniprot.org/uniprotkb/P42212/entry]
Colab
[https://colab.research.google.com/drive/1vlAU_Y84lb04e4Nnaf1axU8nQA6_QBP1?usp=sharing]
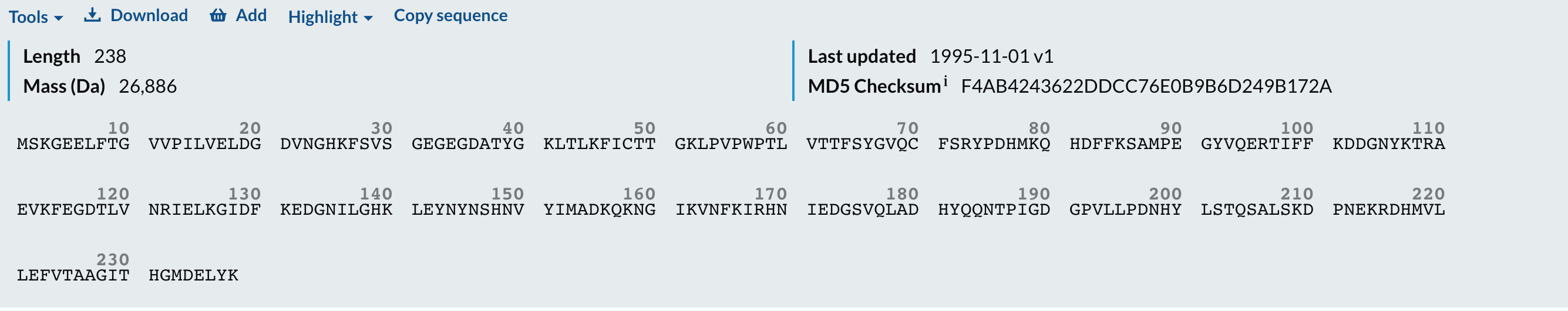


for amino, count in amino_acid_count.most_common():
print(amino, count)

sequence_length = len(cleaned_sequence)
print(f"The total length of the amino acid sequence is: {sequence_length} residues.”)

The amino acid sequence of GFP has a total length of 238 residues, and the most frequent amino acid is glycine (G), appearing 22 times
UniProt BLAST [https://www.uniprot.org/blast]
IDにGFP(Aequorea victoria)のUniProt ID:P42212を入力しRun BLASTする
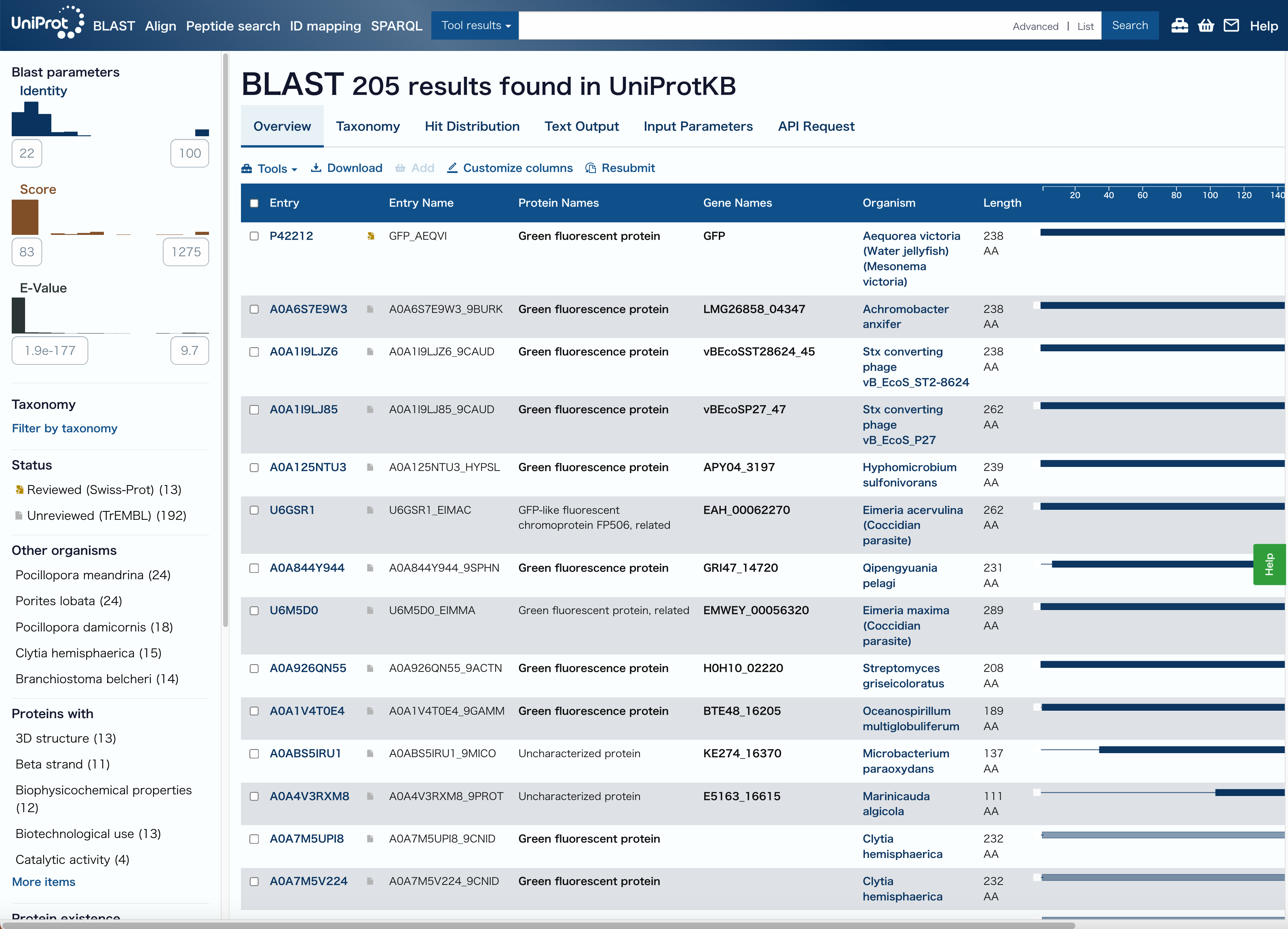
A BLAST search using UniProt identified 205 homologous protein sequences related to GFP across different organisms.
[https://www.uniprot.org/uniprotkb/P42212/entry]
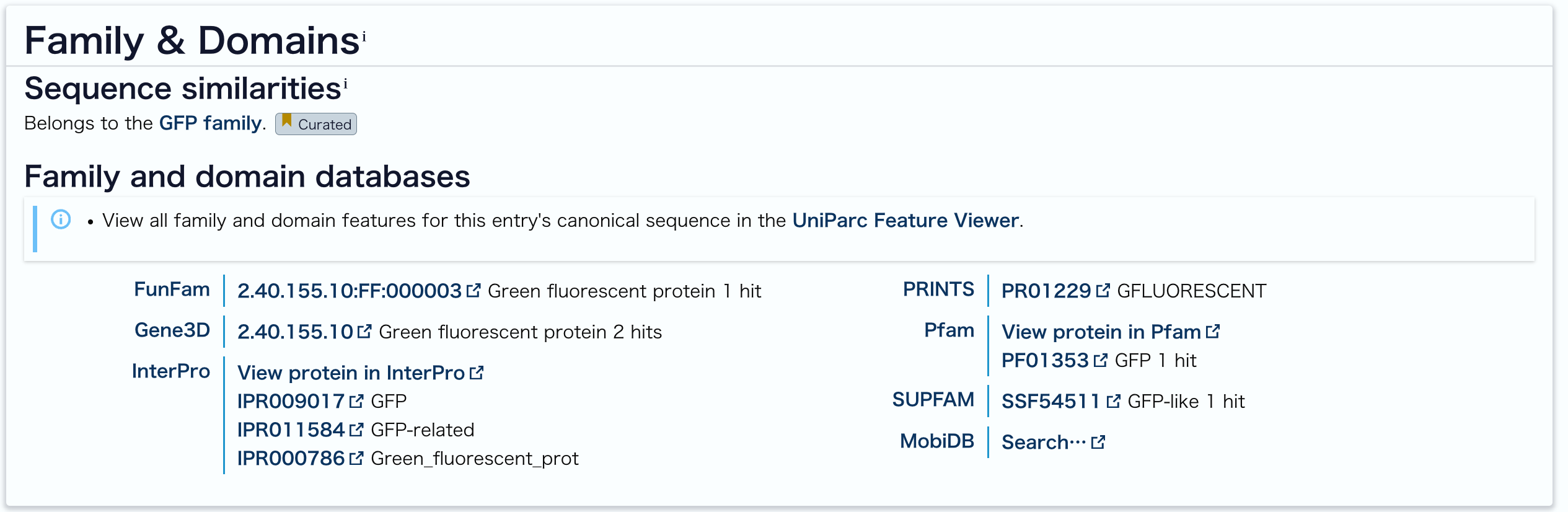
Belongs to the GFP family
[https://www.rcsb.org/]
ここで、Green fluorescent proteinで検索すると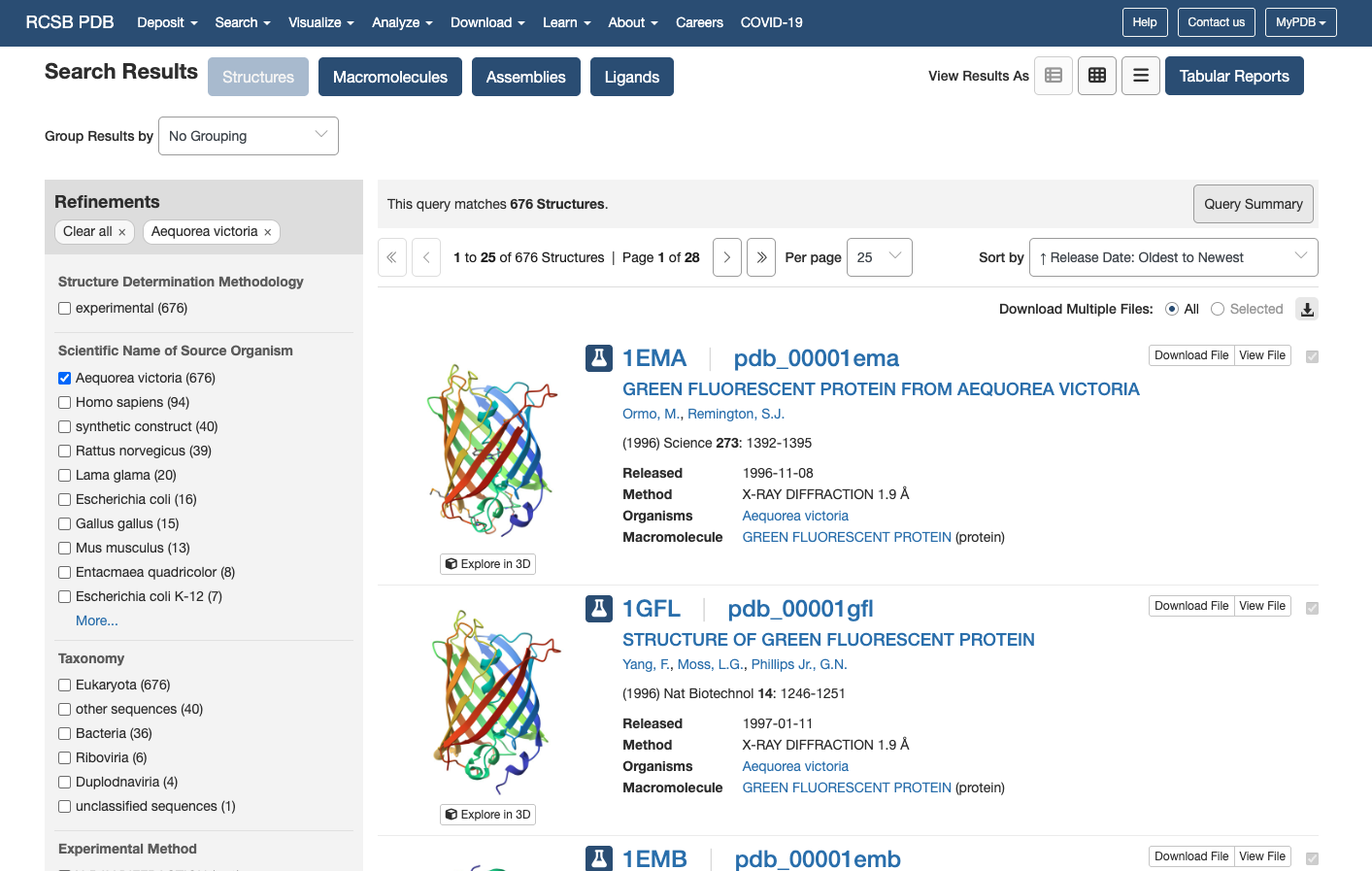
一番上の1EMAが出てくる
GREEN FLUORESCENT PROTEIN FROM EQUOREA VICTORIA
[https://www.rcsb.org/structure/1EMA]
- Classification: Fluorescent Protein
- Organism(s): Aequorea victoria(クラゲ)
- Method: X-ray crystallography
- Resolution: 1.9 Å
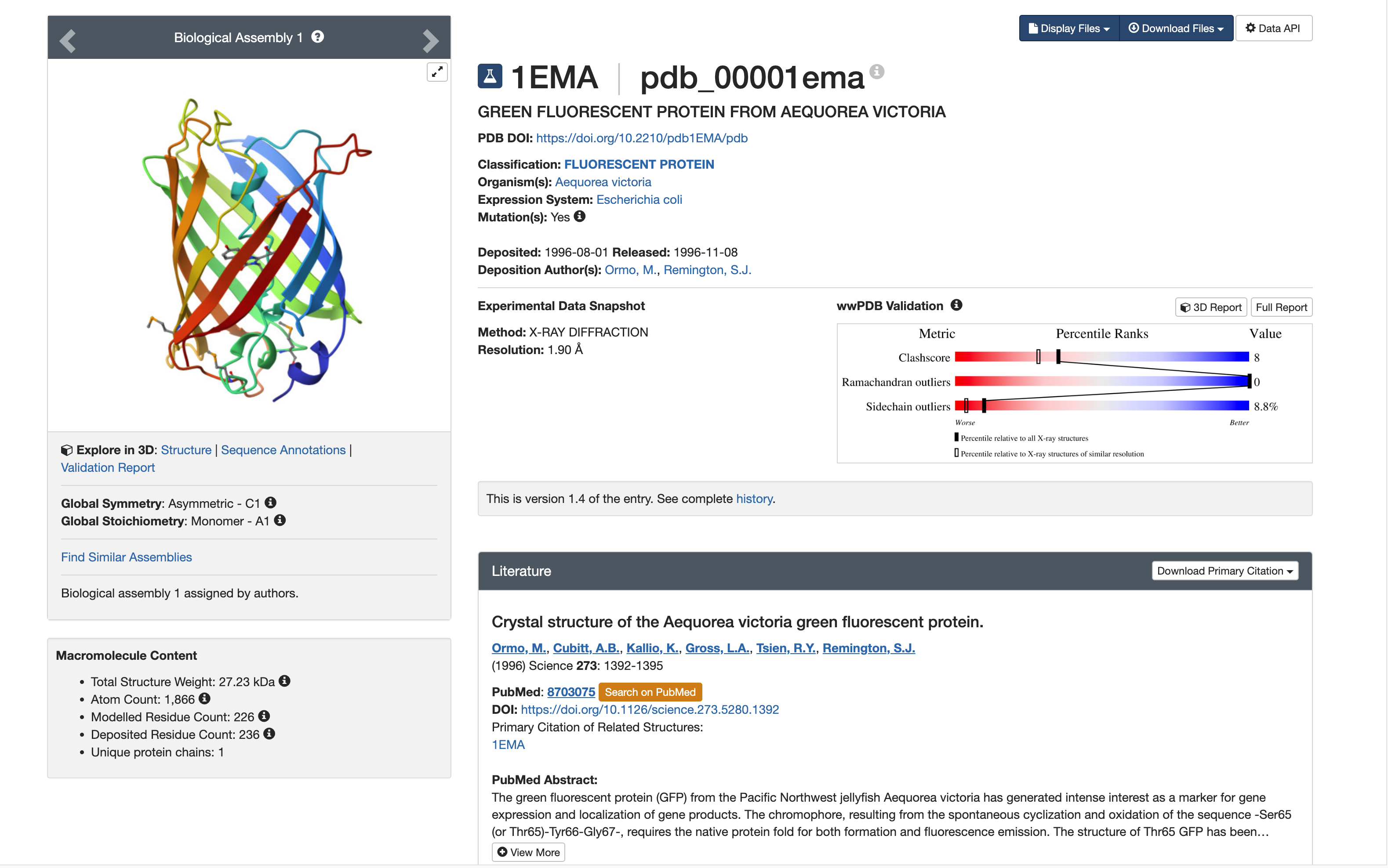
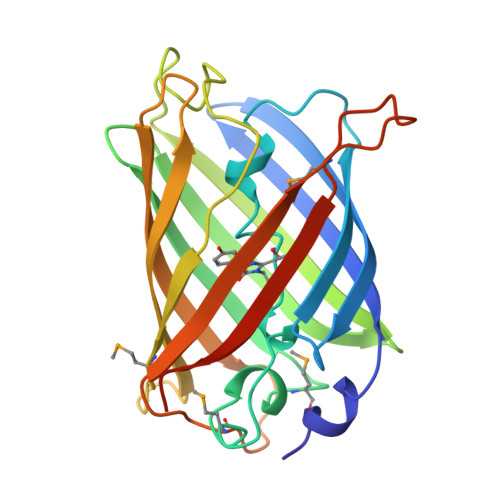
[https://www.rcsb.org/structure/1EMA]
- Released: 1996-11-08
- Resolution: 1.90 Å
Yes, it is a good quality structure.
[https://www.rcsb.org/structure/1EMA]
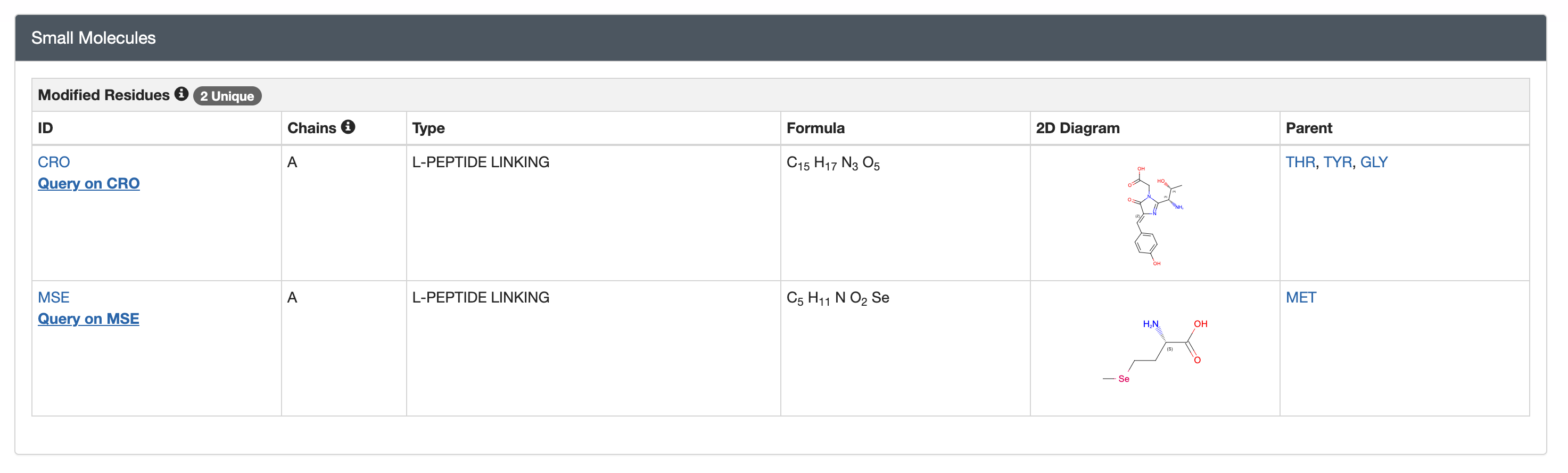
UnitProt “P42212・GFP_AEQVI” [https://www.uniprot.org/uniprotkb/P42212/entry]


[https://www.ebi.ac.uk/pdbe/scop/term/8096045]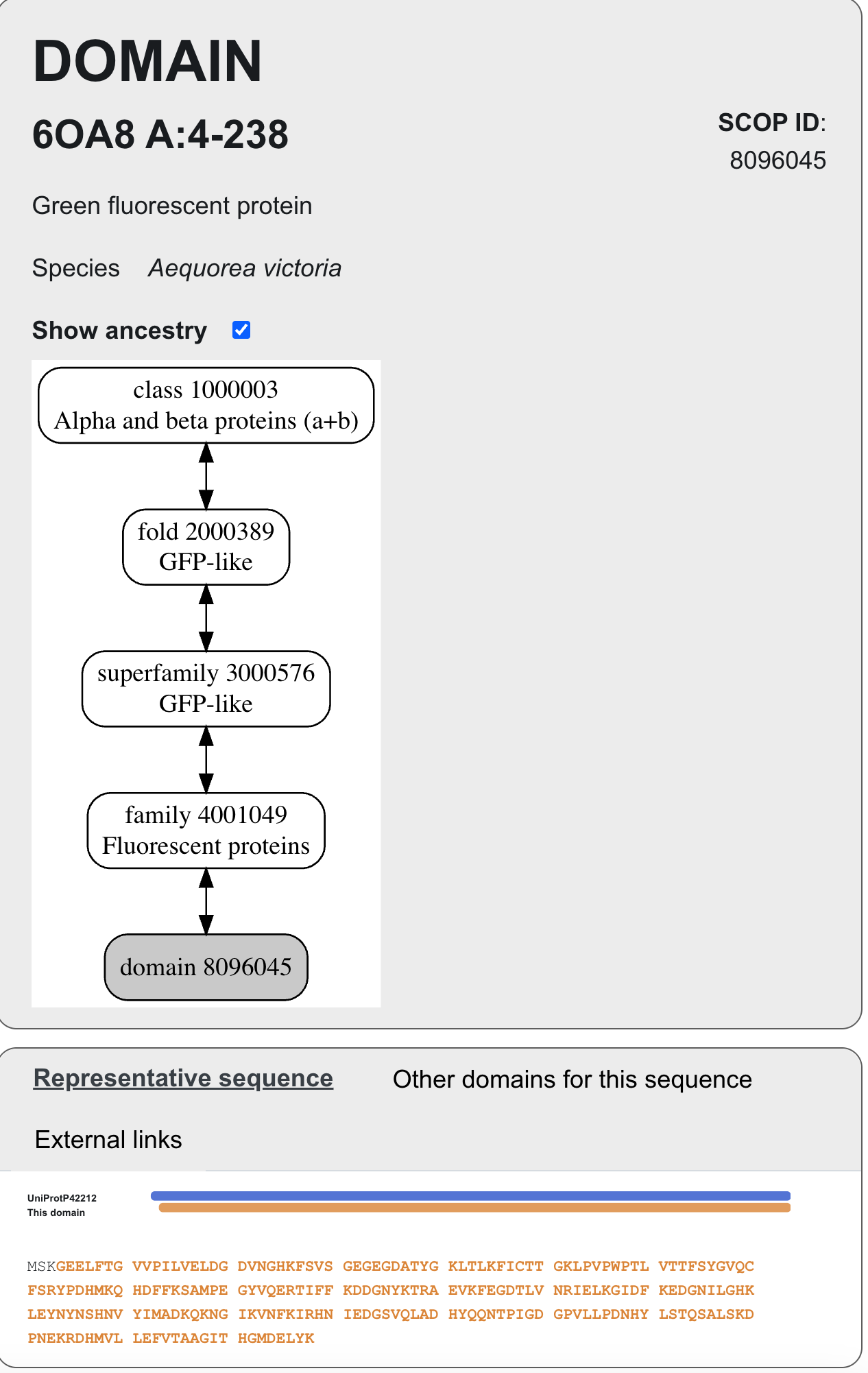
Alpha and beta proteins ⇔ GFP-like ⇔ GFP-like ⇔ Fluorescent proteins
GFP belongs to the SCOP structural classification family as follows:
Class: Alpha and beta proteins (α+β)
Fold: GFP-like
Superfamily: GFP-like
Structure Family: Fluorescent proteins
According to the SCOP structural classification database, GFP belongs to the fluorescent protein family.
CHAT GTP Auto
</> pymol
fetch 1ema
cartoon
hide everything
show cartoon
ribbon
hide everything
show ribbon
ball and stick
hide everything
show stick show spheres
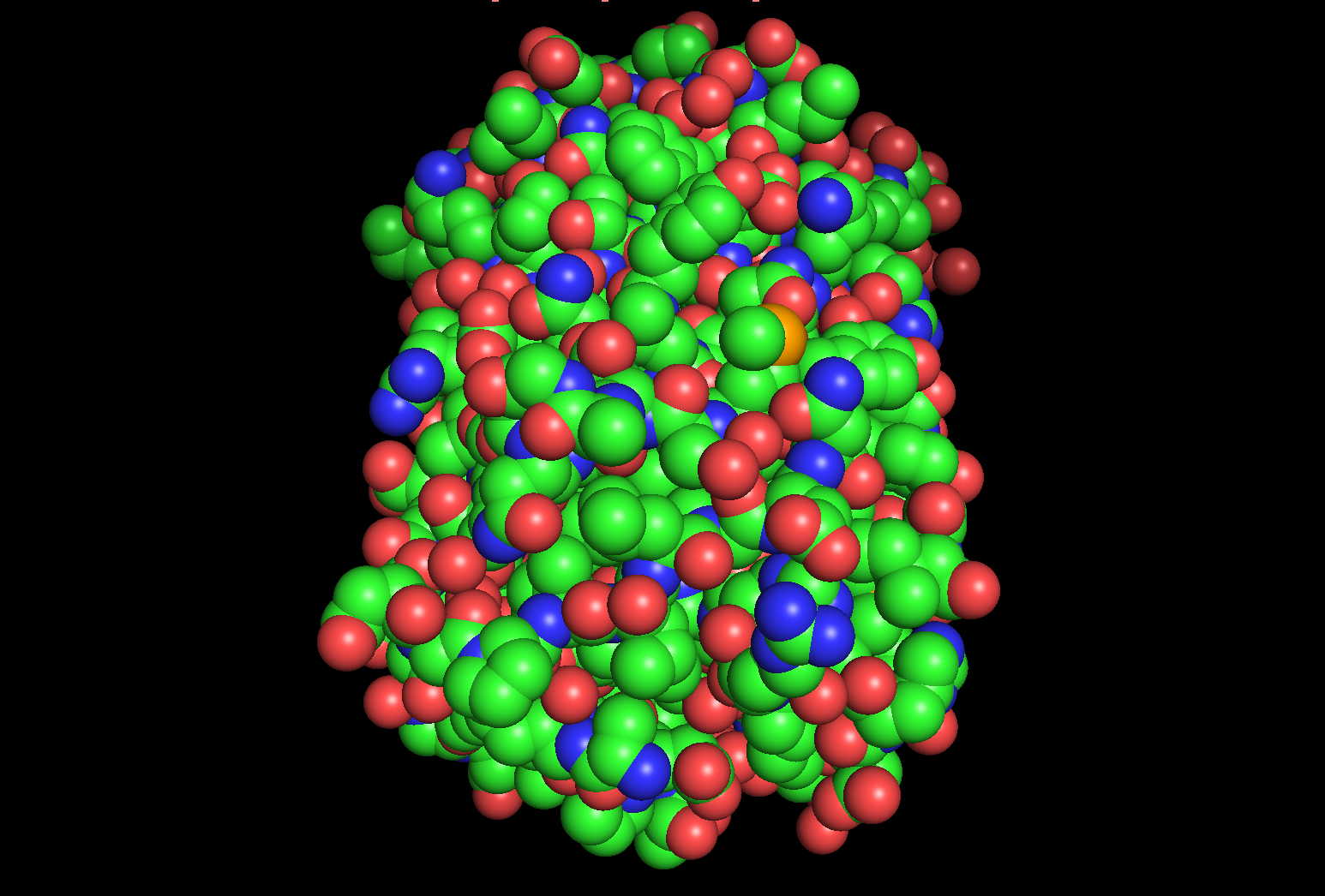

hide everything show cartoon
color red, ss h # αヘリックスを赤色で表示
color yellow, ss s # βシートを黄色で表示
color green, ss l+’’ # ループ(コイル部分)を緑色で表示
α-helix: red
β-sheet: yellow
coil: green
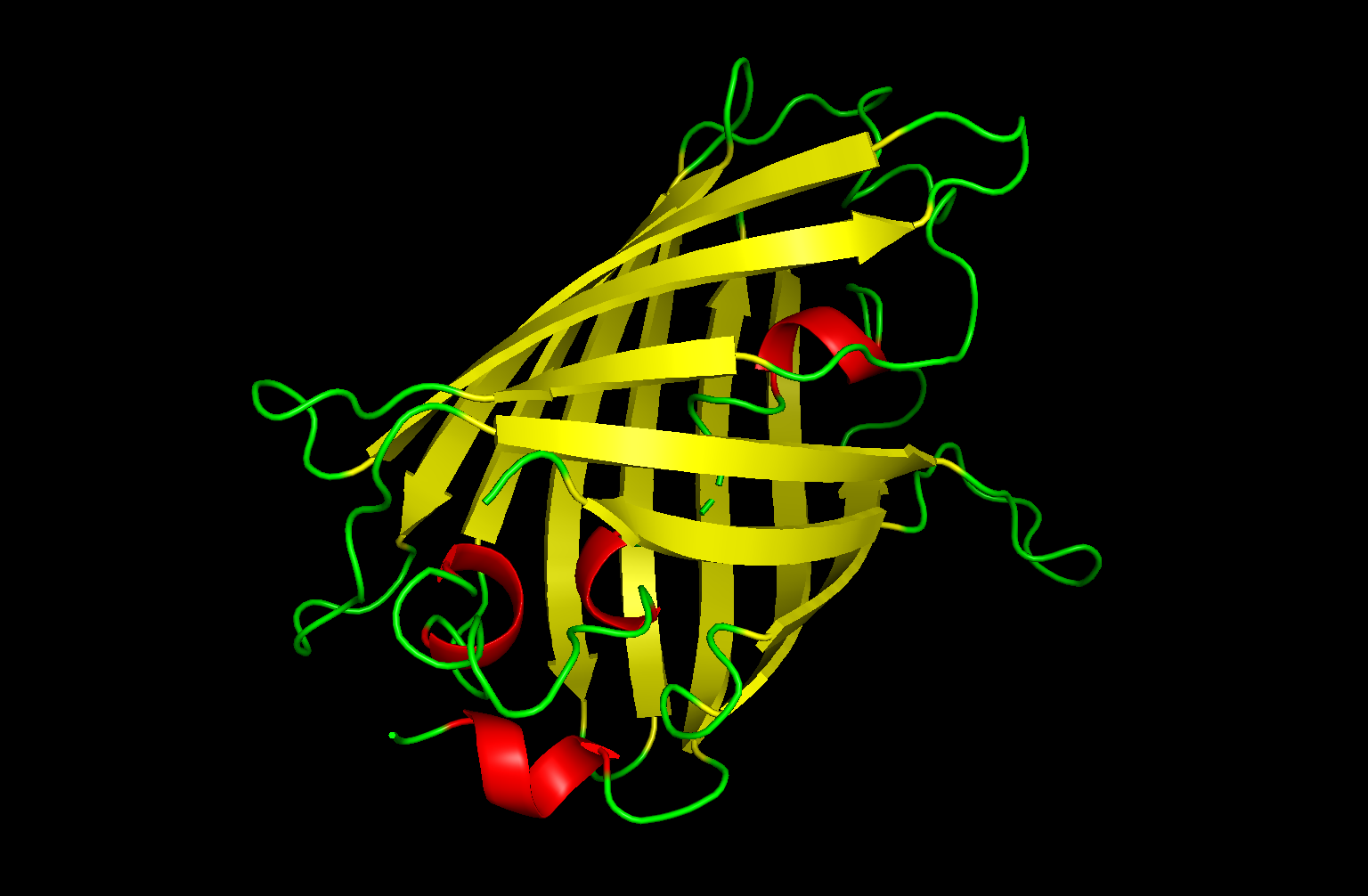
The structure has more yellow regions, indicating that it is rich in β-sheets.
#まずタンパク質をリセットします
hide everything
show cartoon
#疎水性アミノ酸をオレンジに色付け
color orange, resn ALA+VAL+ILE+LEU+MET+PHE+TYR+TRP+PRO
#親水性アミノ酸をシアン色に色付け
color cyan, resn ARG+ASN+ASP+GLN+GLU+HIS+LYS+SER+THR+CYS
#中性または特定が難しいグリシン(G)を白に色付け
color white, resn GLY
Hydrophobic residues : orange
Hydrophilic residues in : cyan
Neutral or ambiguous residues (such as glycine) : white
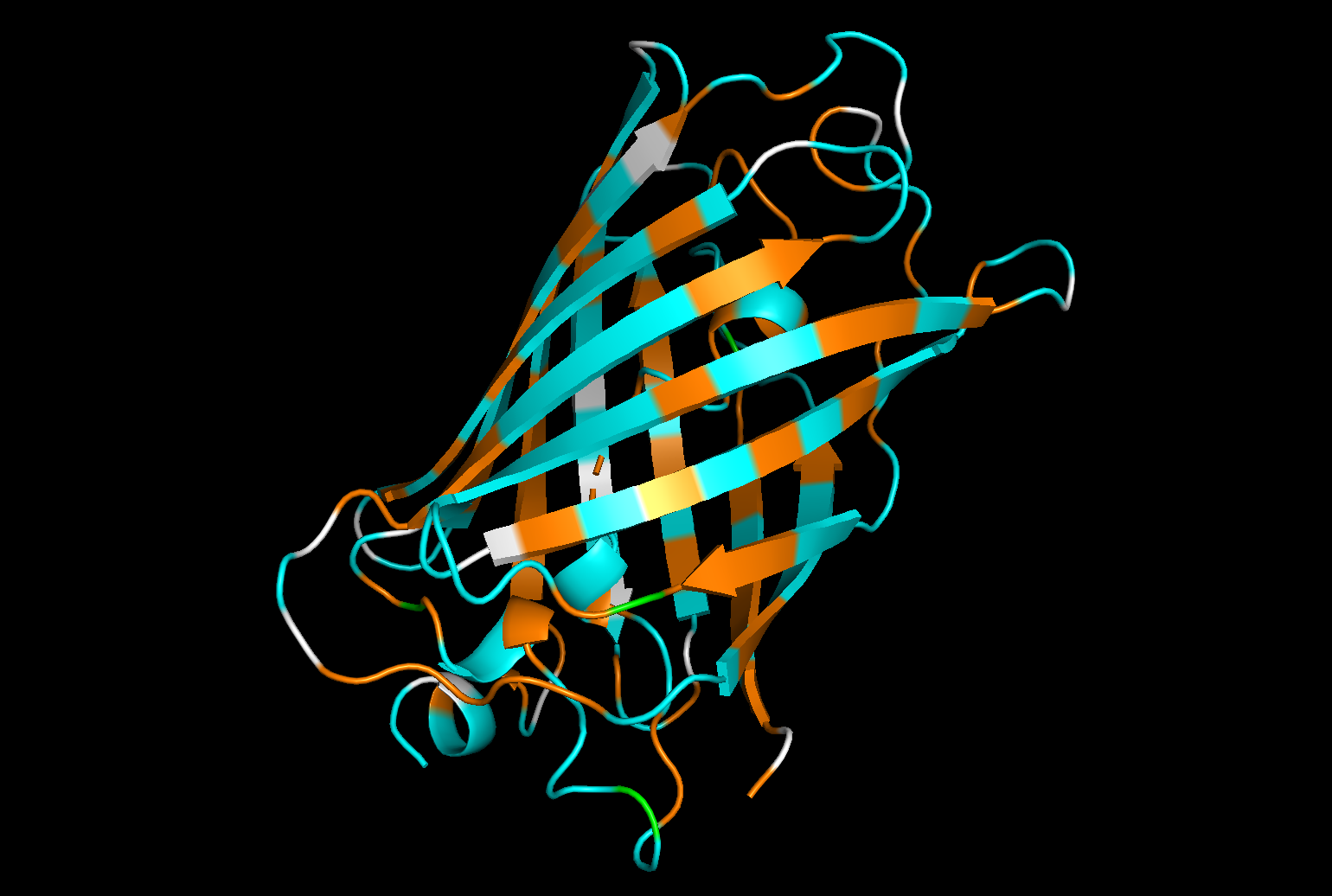
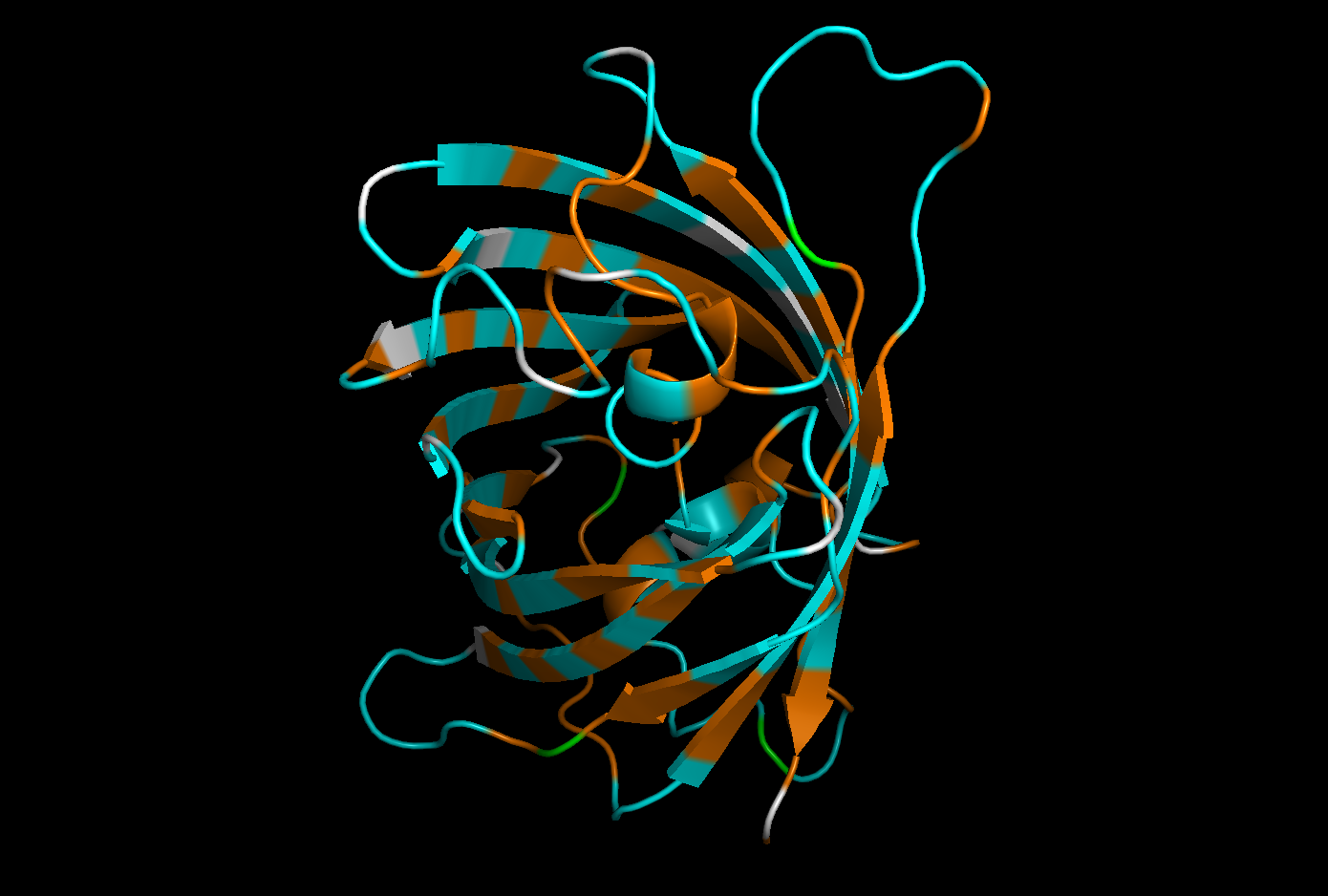
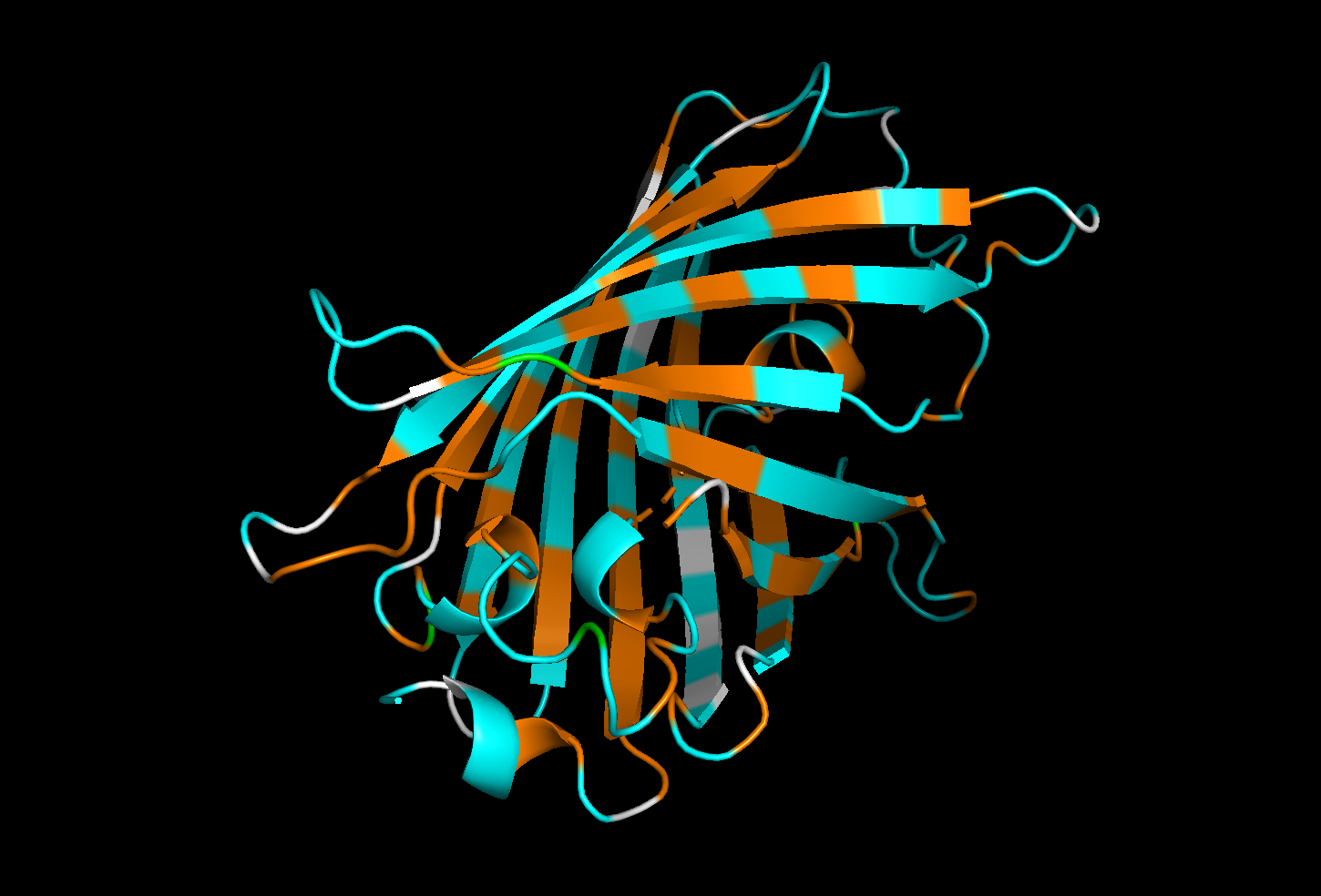
The distribution of hydrophobic and hydrophilic residues in GFP appears to be relatively balanced, with both types of residues evenly distributed throughout the structure
hide everything show surface set transparency, 0.2
The protein surface visualization reveals clear binding pockets or “holes” on the protein surface.
Part C. Using ML-Based Protein Design Tools
In this section, we will learn about the capabilities of modern protein AI models and test some of them in your chosen protein.
Colob [https://colab.research.google.com/drive/1L6ok_BjbmiIM_xB99dT4k0mHO9RnNSBy#scrollTo=33580eea]
For this section I continued using Green Fluorescent Protein (PDB ID: 1EMA), which I selected earlier from the Protein Data Bank.
C1. Protein Language Modeling
a.
GFPのヒートマップを作る
[https://colab.research.google.com/drive/1L6ok_BjbmiIM_xB99dT4k0mHO9RnNSBy#scrollTo=09FwbZ6v1AUs]
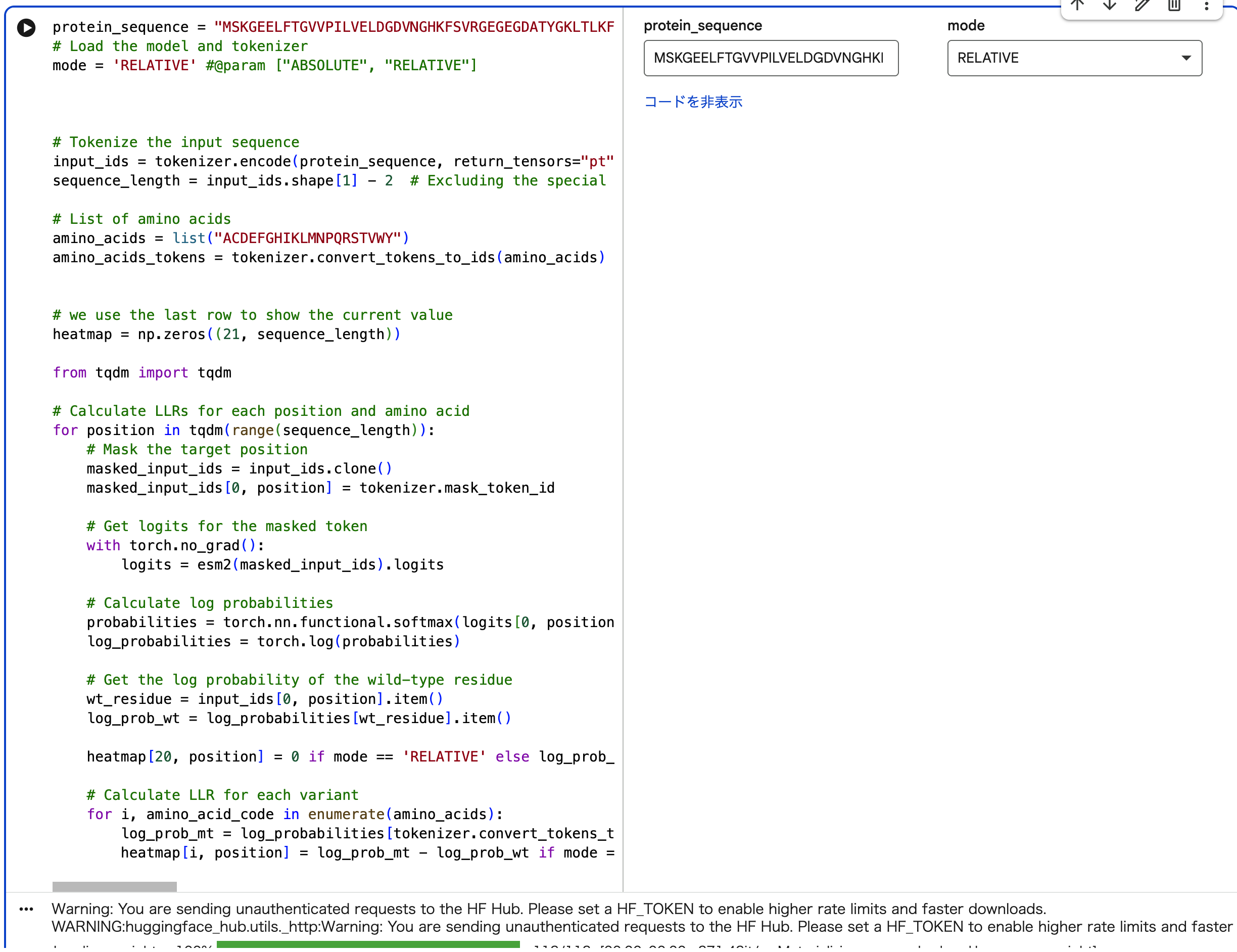
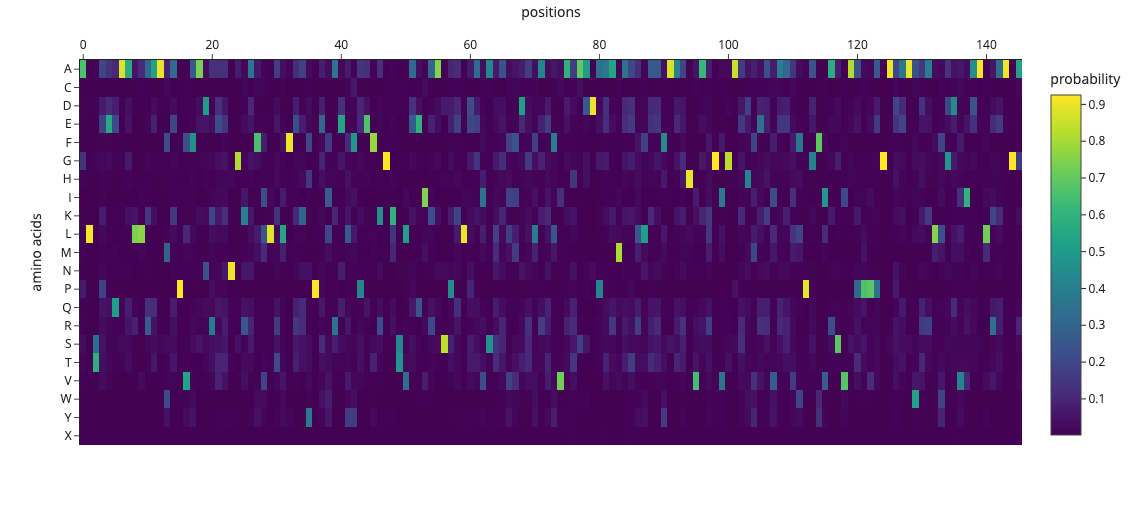
An unsupervised deep mutational scan of GFP was generated using the ESM2 language model. The resulting heatmap shows the predicted likelihood of all possible amino acid substitutions across the protein sequence.
b.
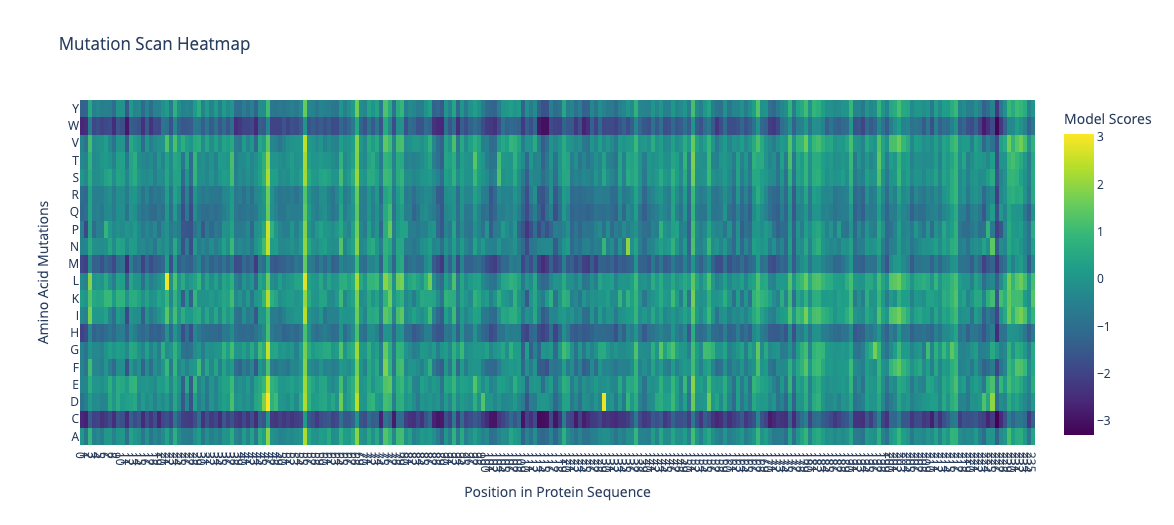
c. (Bonus) NA
a.
Protein sequences from the provided dataset were embedded using the ESM2 language model and projected into a reduced three-dimensional latent space using t-SNE. This visualization allows relationships between protein sequences to be explored.
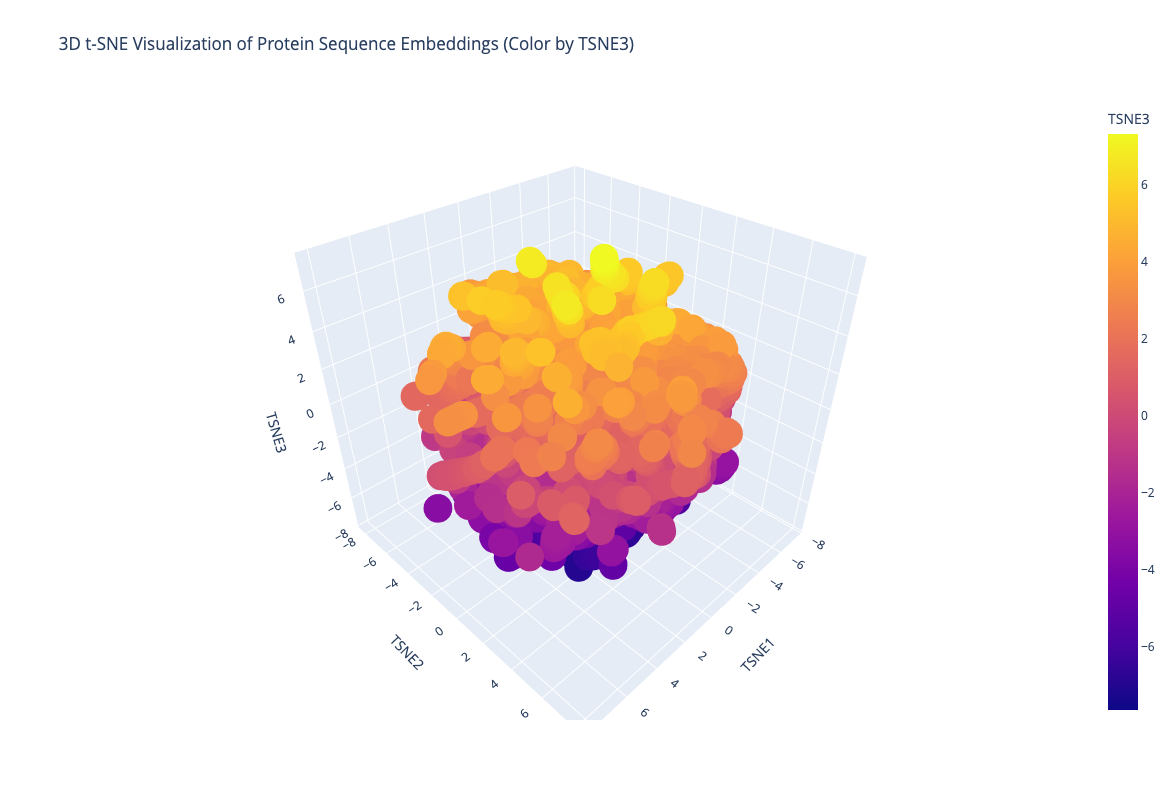
b. The visualization shows that proteins form neighborhoods in the latent space. This suggests that the language model captures meaningful relationships between sequences, as similar proteins tend to cluster together.
C.
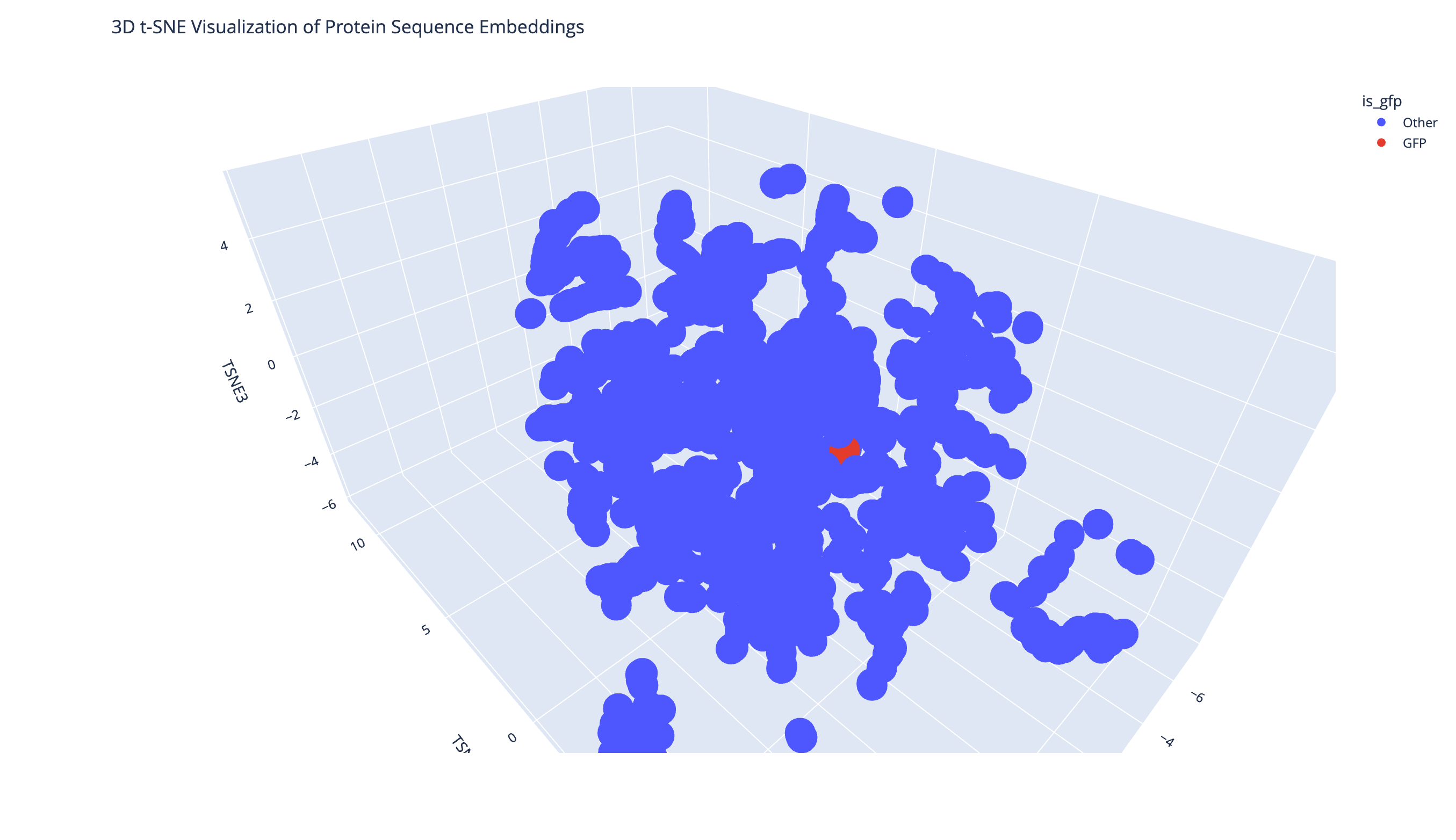
GFP appears as a single point in the latent space because only one GFP sequence was added. Its position within a dense neighborhood suggests that GFP shares meaningful sequence-level features with nearby proteins.
C2. Protein Folding
Folding a protein
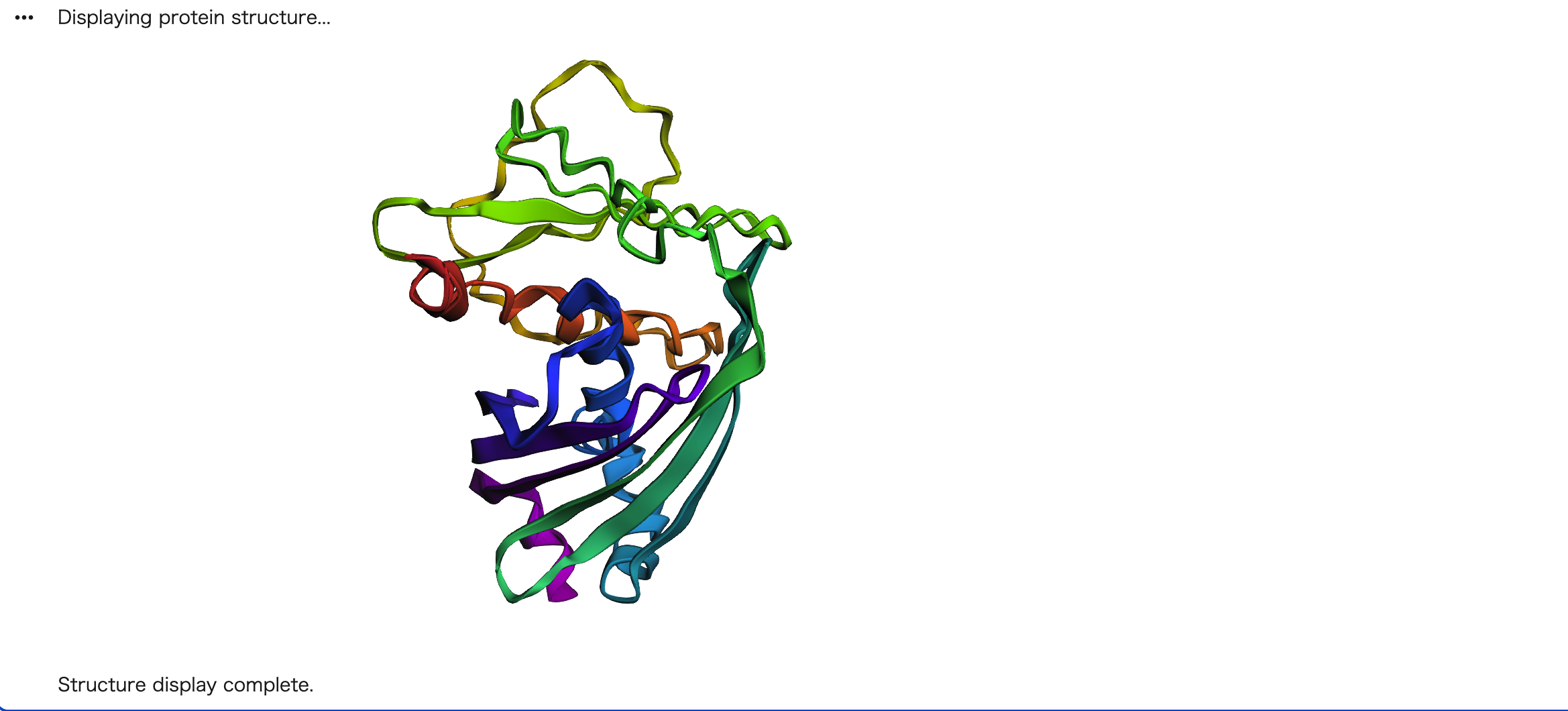
PDB: 1EMA
[https://www.rcsb.org/structure/1EMA]

The ESMFold-predicted structure of GFP closely matches the original crystal structure (PDB: 1EMA). Both show the characteristic 11-stranded β-barrel, which is the hallmark of GFP. However, the predicted structure appears slightly less compact compared to the experimentally determined structure.
First, I tried a small mutation by changing the last amino acid from K (Lysine) to A (Alanine). As shown in the figure below, the overall structure remained almost identical to the original.
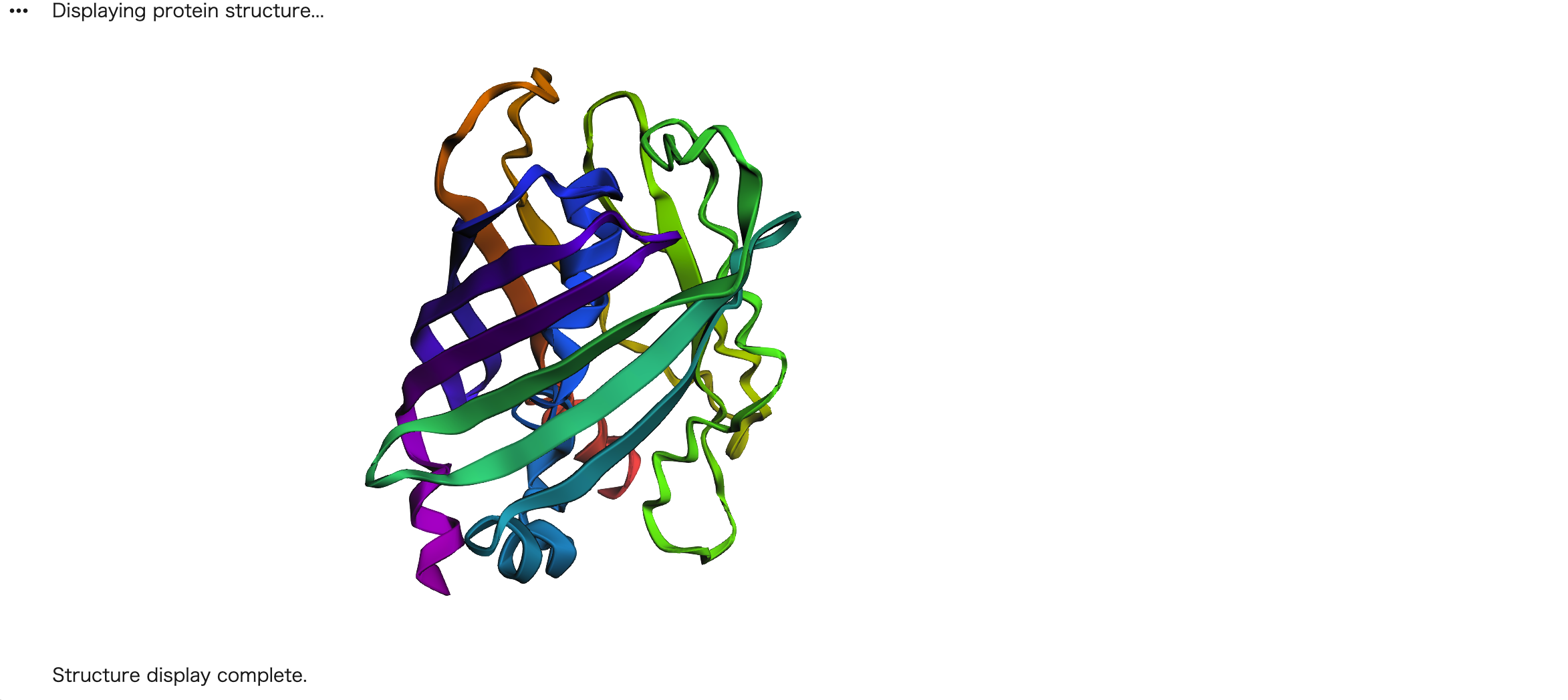
Next, I tried a larger mutation by replacing the last 9 amino acids with Alanine (AAAAAAAA). Again, the β-barrel structure was largely preserved. These results suggest that GFP is highly resilient to mutations, especially at the C-terminal region. The core β-barrel scaffold is robust and maintained even when significant sequence changes are introduced at the terminal end.
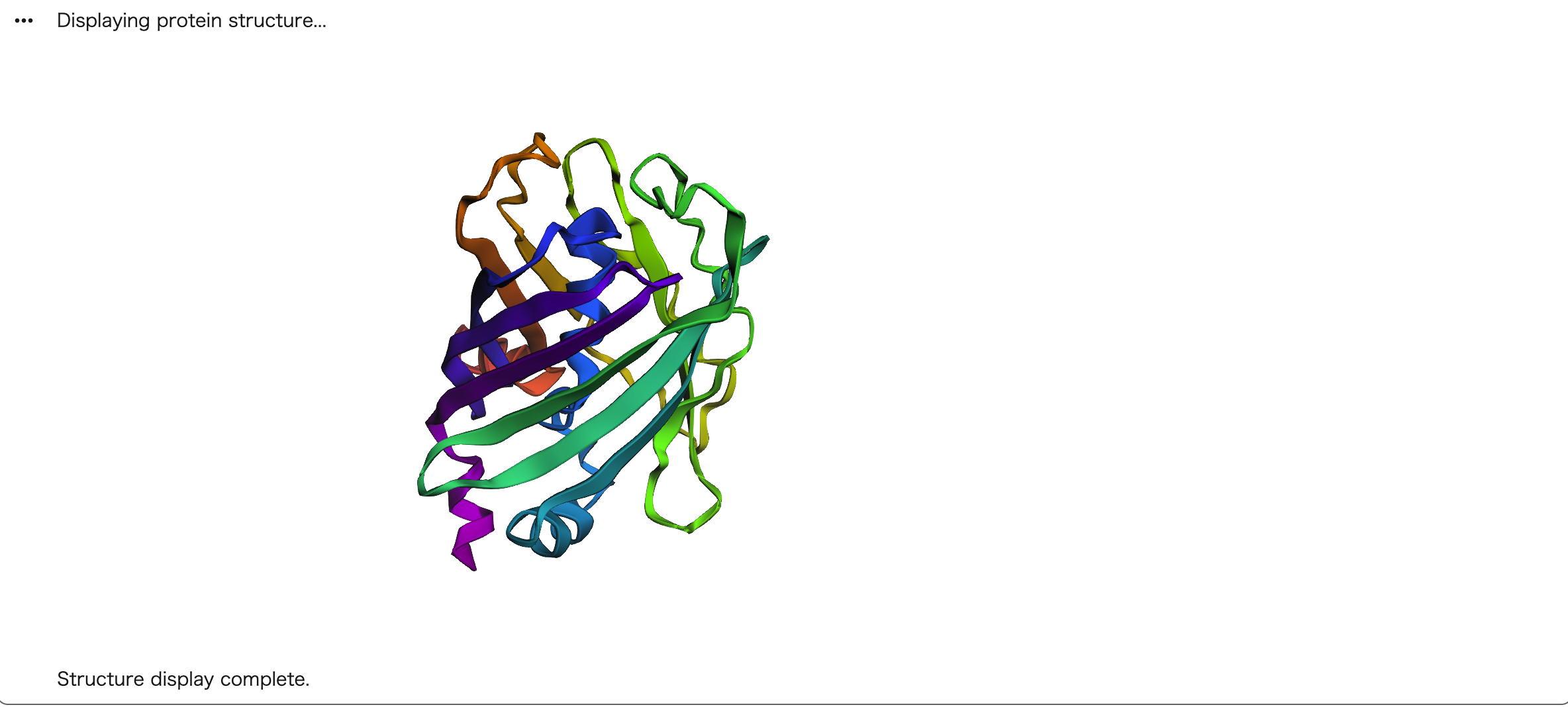
C3. Protein Generation
Inverse-Folding a protein: Let’s now use the backbone of your chosen PDB to propose sequence candidates via ProteinMPNN (選んだPDBの骨格構造を使って、ProteinMPNNで配列の候補を提案する)
sequence probability heatmap
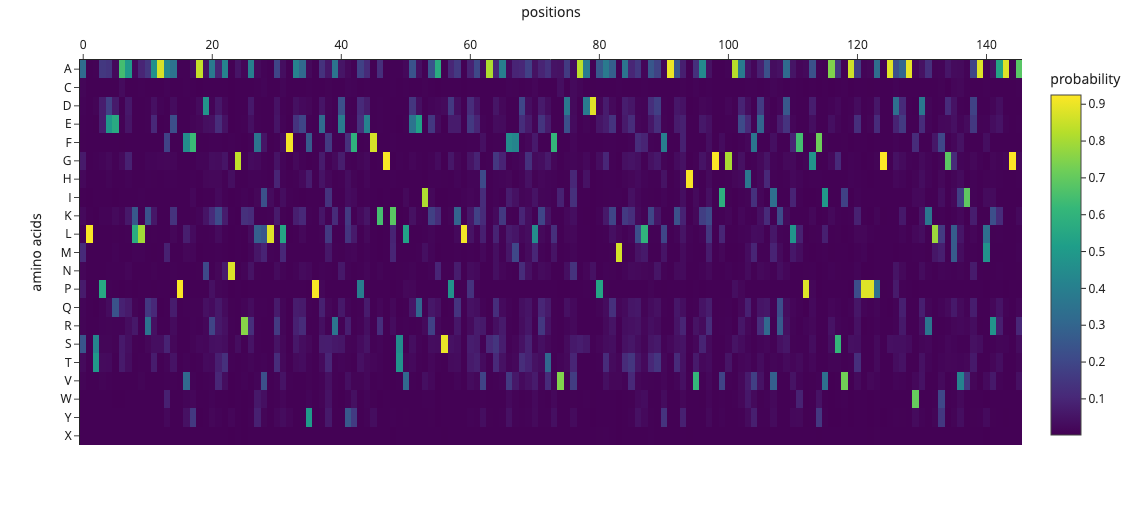
横軸(positions) → タンパク質の各位置(0〜150番目のアミノ酸) 縦軸(amino acids) → 20種類のアミノ酸(A, C, D, E, F…) 黄色 → 確率高い(0.9以上)→ ProteinMPNNが強く推薦
Colabより
New Sequence:
ALTPEEAAKLAAAWAPVAANAAANGKAFILTLFEKYPEIAEKFPEFKGKSLEEIKASPKLPAISSAFFATLDTLVAVADDAAKMAALLDALAKAHVALGIGAEDFEKVRAIFPGFVASIAPPPAGADAAWDKLFGDIIAALRAAGA
この配列を使って、Colabに入れ直す
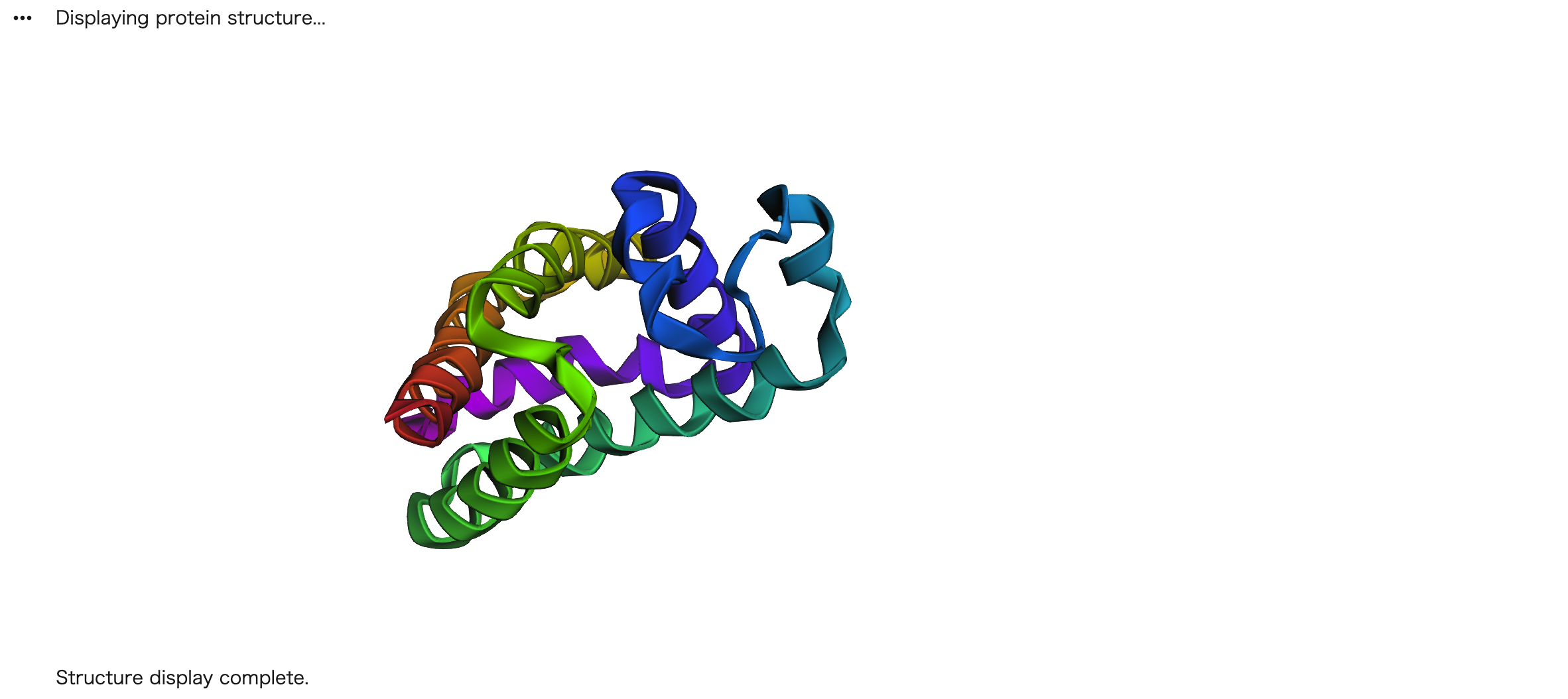
The new sequence folded into alpha-helices instead of the original β-barrel structure.
I input the ProteinMPNN-proposed sequence into ESMFold and compared it to the original GFP structure (1EMA). The original GFP folded into a characteristic β-barrel structure, while the new sequence predicted by ProteinMPNN folded into α-helices.