Week 06 HW -Genetic Circuits Part I: Assembly Technologies
‘Week 6 — Genetic Circuits Part I: Assembly Technologies’
Documentation
Homework: Genetic Circuits Part I: Assembly Technologies
Assignment: DNA Assembly
Answer these questions about the protocol in this week’s lab:
https://www.youtube.com/watch?v=c07_5BfIDTw&t=115s
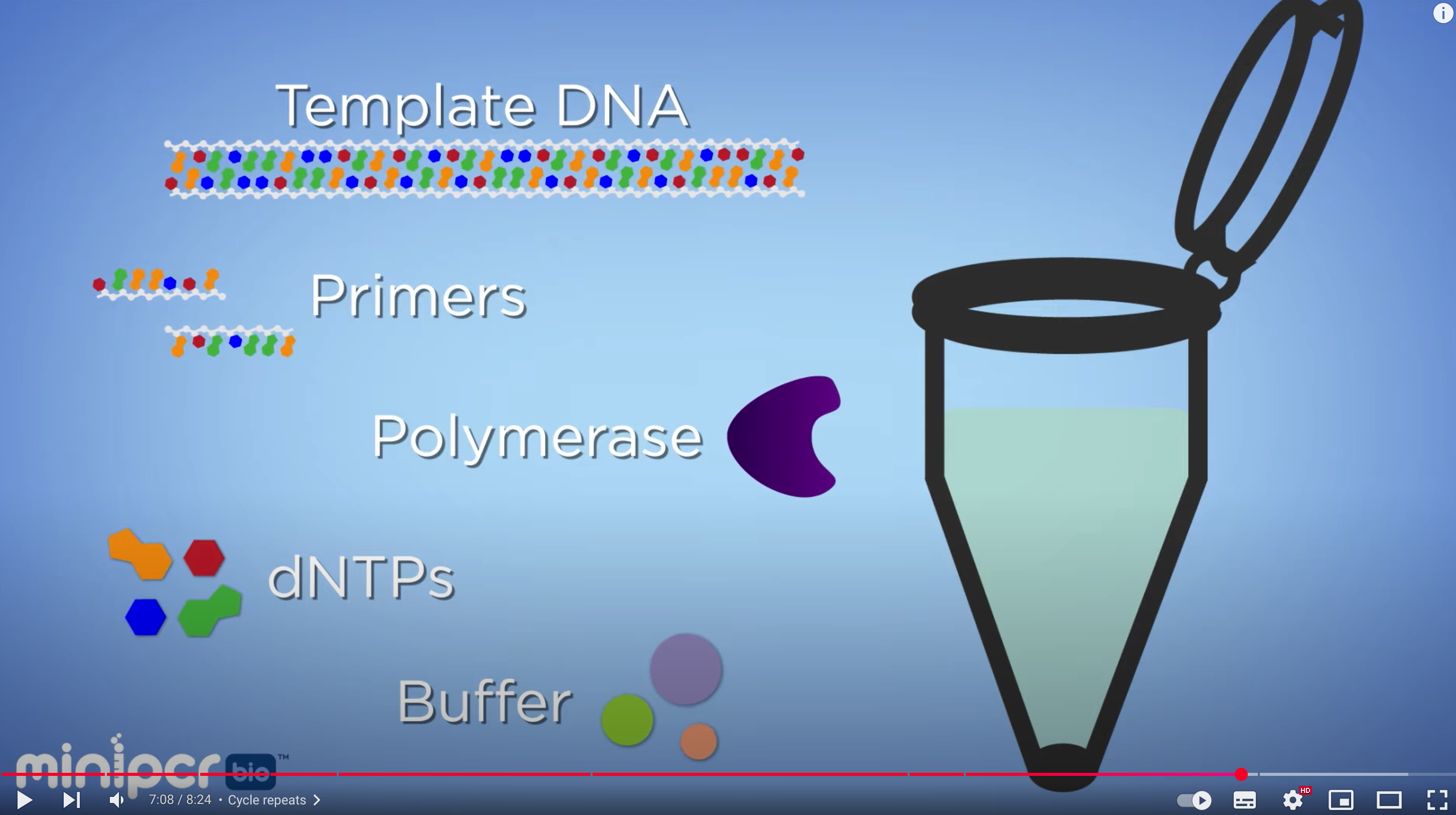
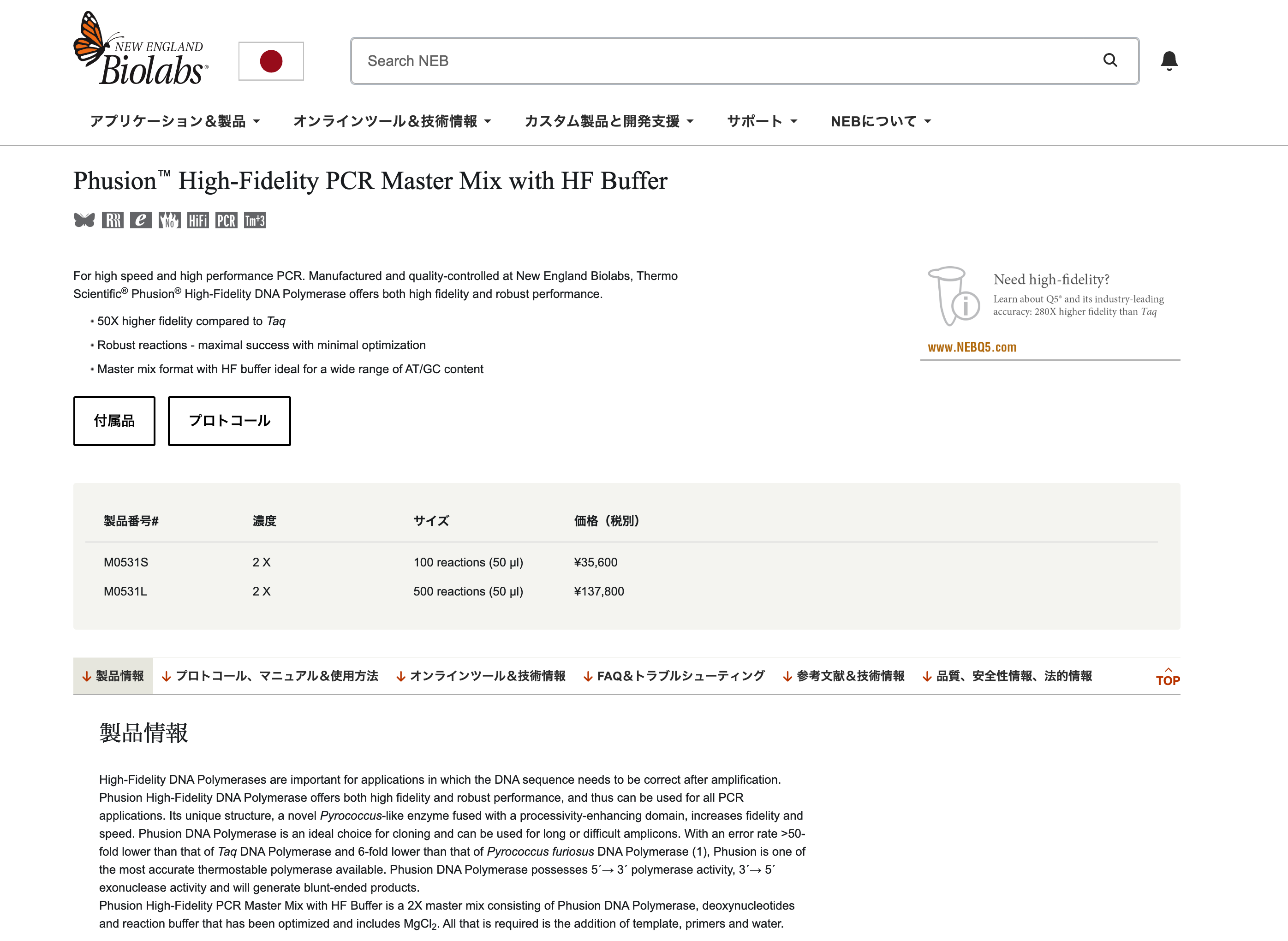


Phusion High-Fidelity DNA Polymerase
The enzyme that drives DNA synthesis with high fidelity. It has 3’→5’ exonuclease (proofreading) activity, which reduces the error rate during PCR.
dNTP(A, T, G, C)
These are the nucleotide building blocks used to synthesize new DNA strands during PCR.
HF Buffer(High-Fidelity Buffer)
This is an optimized reaction buffer that maintains the proper pH and ionic conditions for high-fidelity DNA amplification.
MgCl₂
Magnesium ions are an essential cofactor for DNA polymerase activity and are provided at an optimal concentration (1.5 mM in the 1X final reaction).
Together, these components provide the enzyme, substrates, and chemical environment needed for accurate DNA amplification.
https://www.nippongene.com/siyaku/product/pcr/cat_pcr.pdf

The primer annealing temperature should be set about 5°C lower than the Tm, typically around 55–60 (50- 65?)°C. Higher annealing temperatures increase specificity. At a primer concentration of 0.2 µmol/L, annealing occurs within a few seconds
The extension reaction is commonly carried out at 72°C, and depending on other reaction conditions, the synthesis rate is approximately 35–100 nucleotides per second
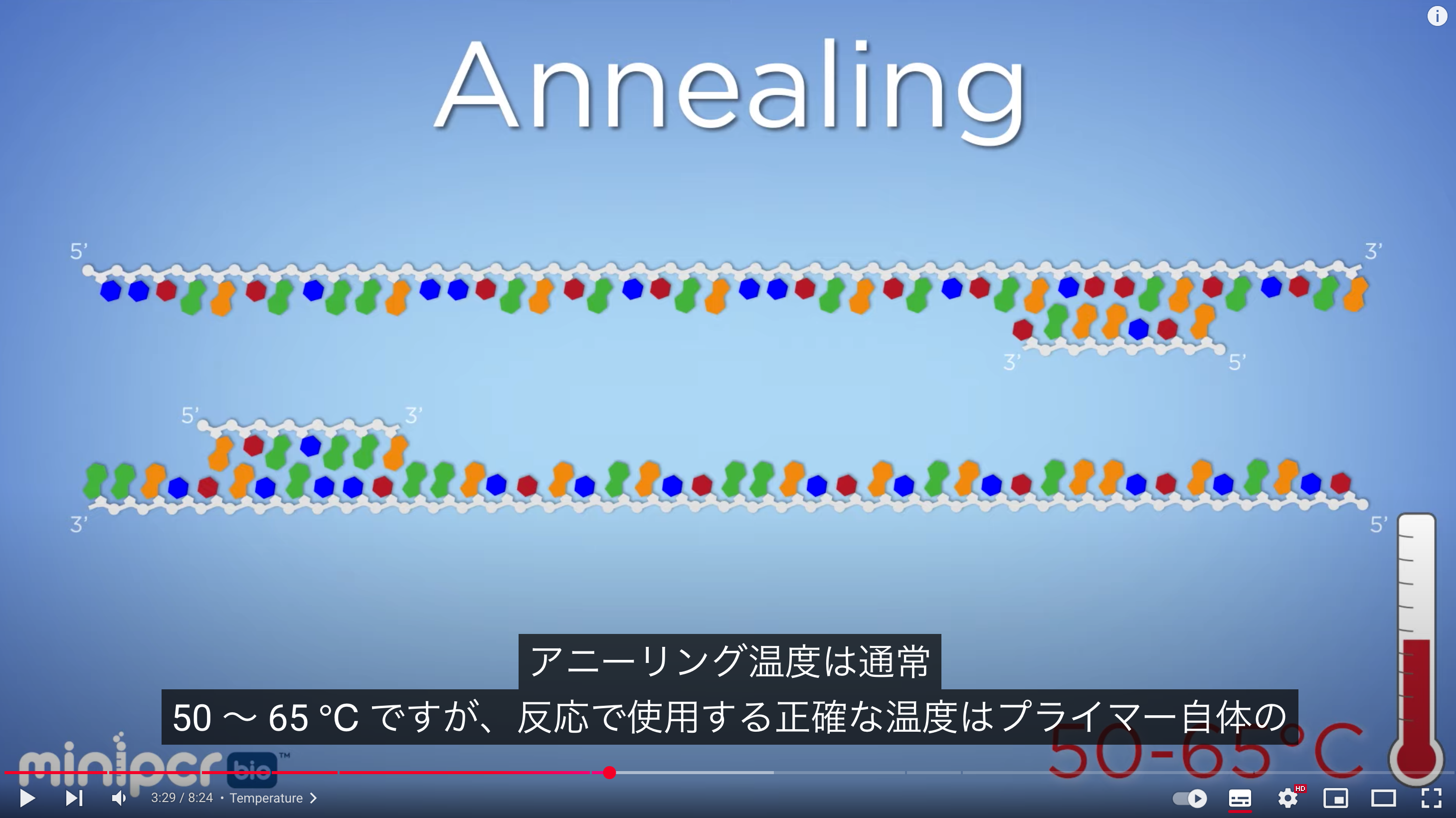
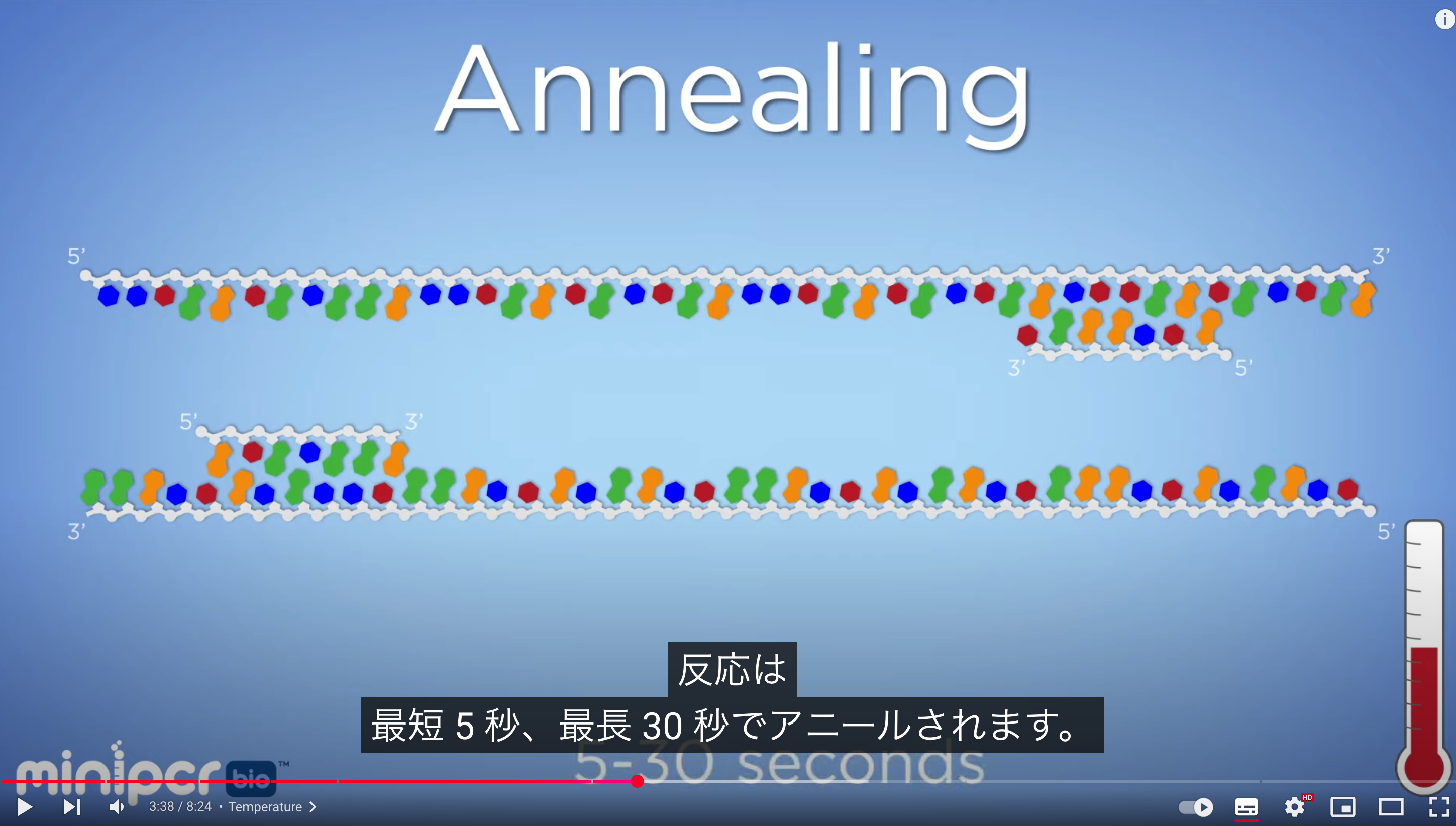

PCR obtains a desired DNA fragment by using primers to flank and amplify a specific region, requiring primers, dNTPs, and polymerase
Restriction enzyme digestion involves enzymes (e.g., EcoRI) that recognize specific restriction sites and cut the DNA
If no such sites exist in the vector or DNA sequence, digestion is not possible
Thus, PCR is used when the target restriction site is absent, while restriction digestion is preferred if the vector already contains suitable restriction sites and high reproducibility is desired
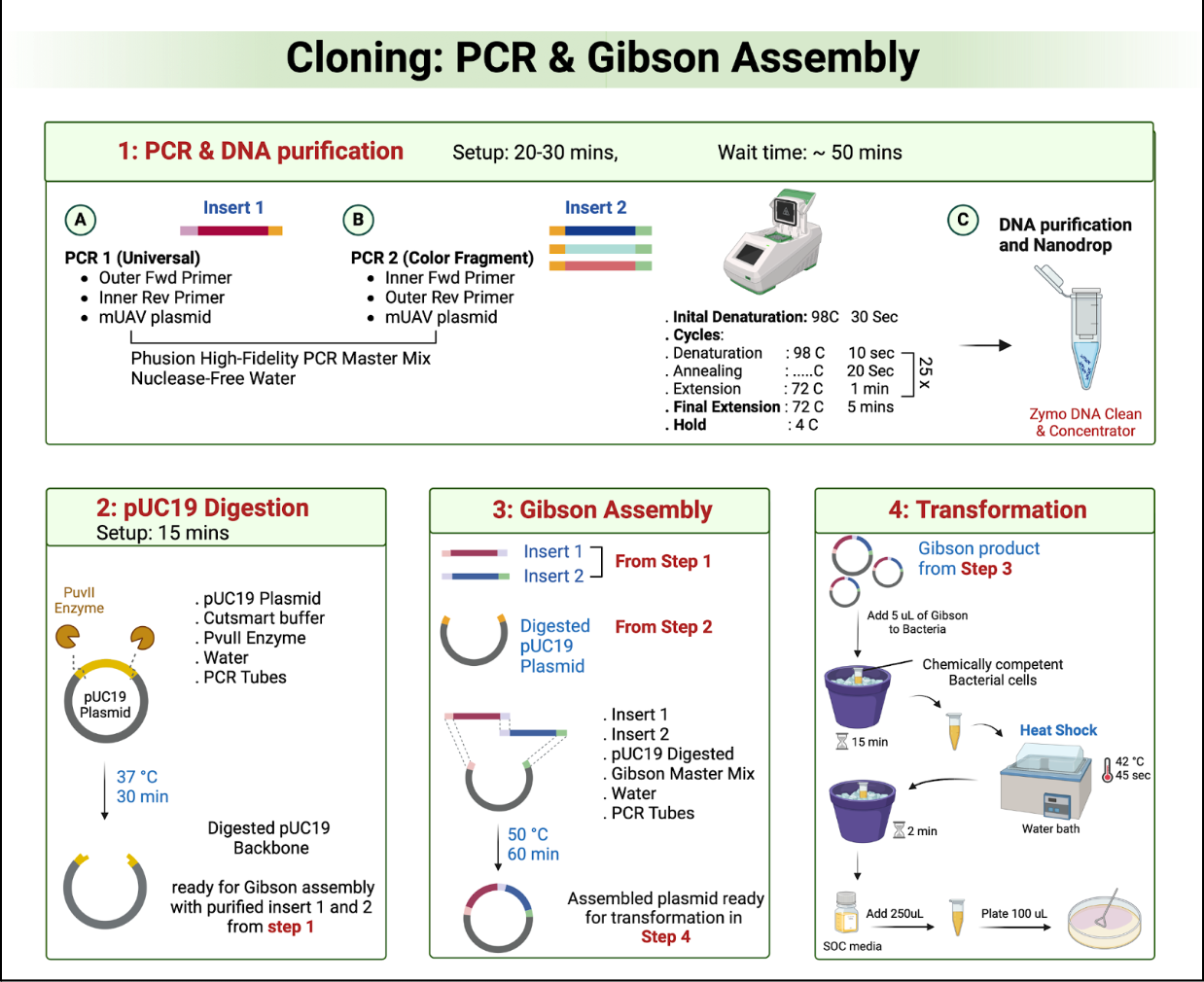
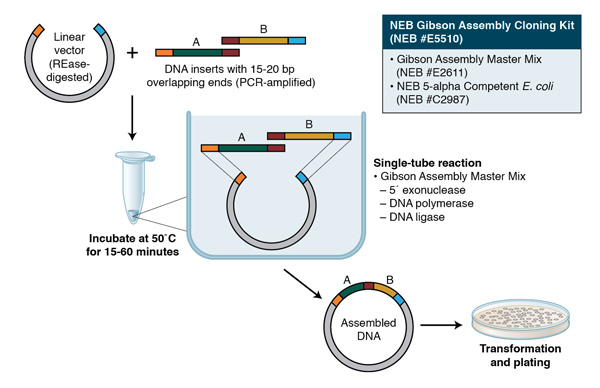

Overlap Design
Confirm that each fragment has the correct 20–40 bp overlaps required for Gibson Assembly
Fragment Size and Sequence Accuracy
Check via gel electrophoresis (for size) and, if necessary, sequencing (for the correctness of overlap regions or introduced mutations)
Orientation and Molar Ratios
Ensure each fragment is in the proper 5′→3′ orientation and use an appropriate insert-to-vector molar ratio (commonly 2:1) to maximize assembly efficiency
In order to introduce DNA into cells, it is necessary to temporarily increase the permeability of the cell membrane
Generally, there are two main methods for transforming E. coli: chemical transformation (e.g., the CaCl₂ method) and electroporation
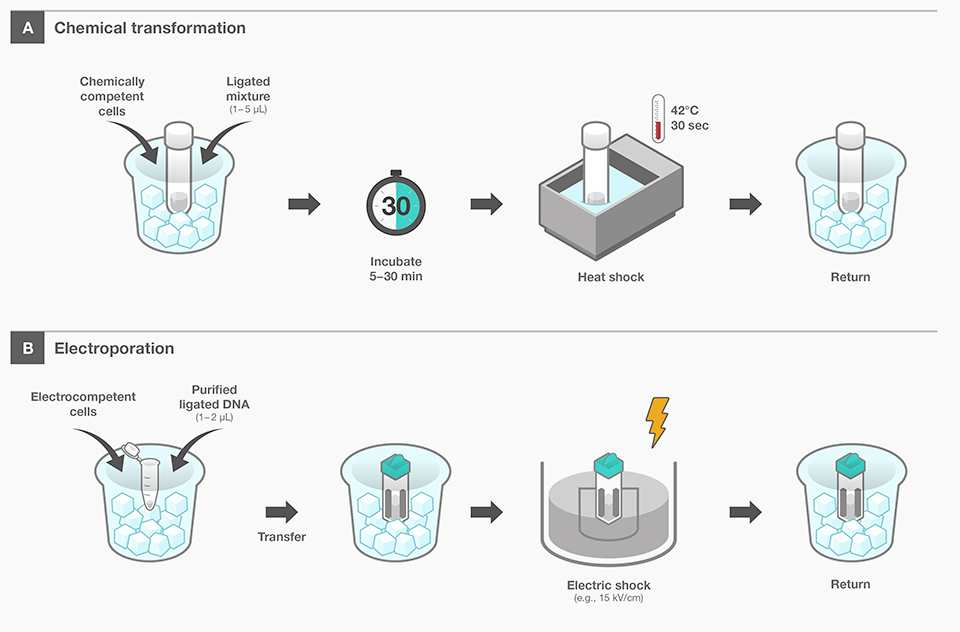
(A) Chemical Transformation
- Mix chemically competent E. coli cells (prepared with CaCl₂, etc.) with ligated DNA and incubate for a set period
- Briefly subject the mixture to a high temperature (e.g., 42°C) for heat shock, creating temporary pores in the cell membrane through which the DNA can enter
- Transfer the cells into a recovery medium afterward to allow them to recover
(B) Electroporation
- Add purified DNA to electrocompetent E. coli cells
- Apply a high-voltage pulse (e.g., ~15 kV/cm), which forms transient micro-pores in the cell membrane, enabling the DNA to enter
- Finally, move the cells to a recovery medium to allow them to recuperate
https://www.youtube.com/watch?v=NzQdLQ44I7w
https://www.youtube.com/watch?v=EpHeu44hitI
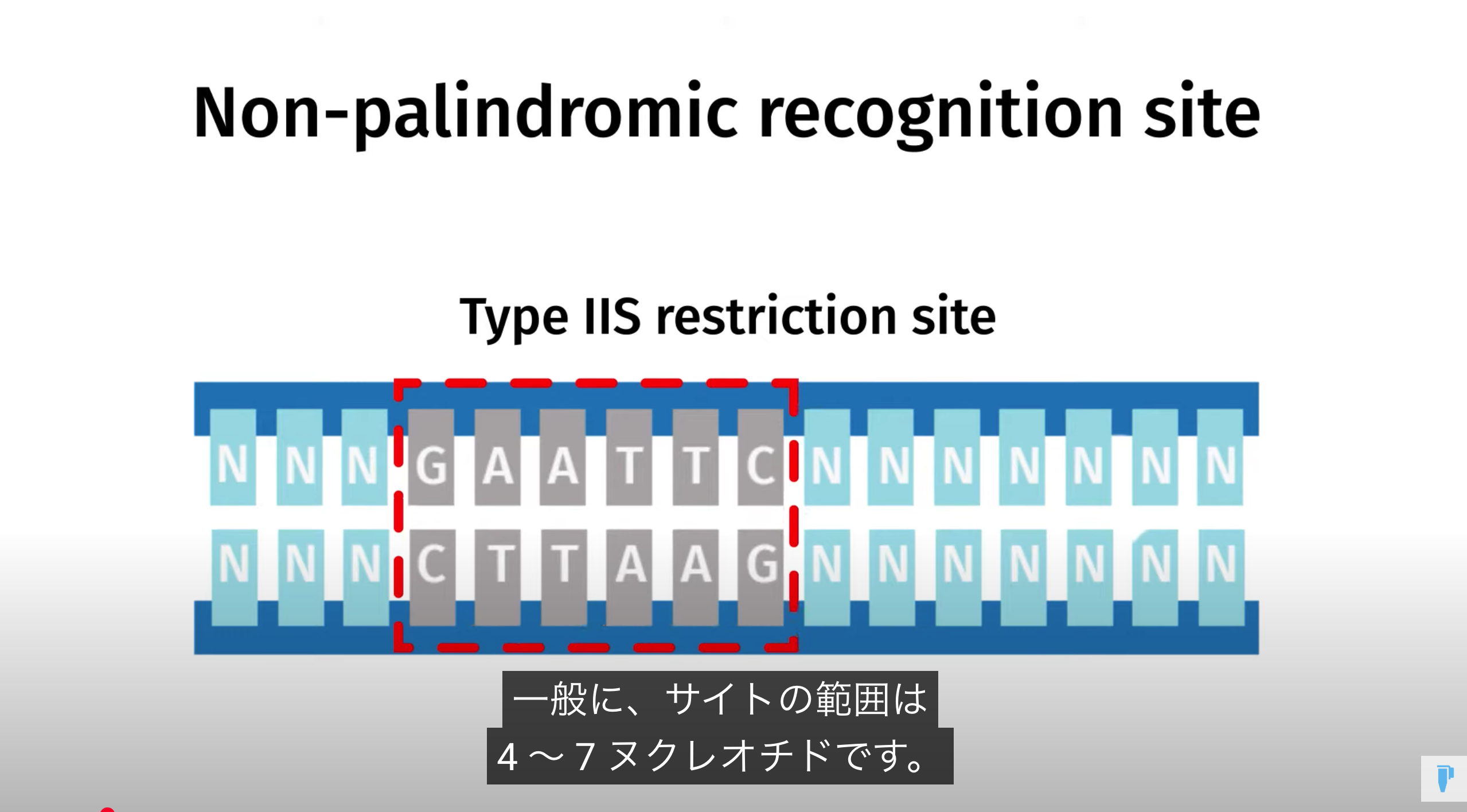
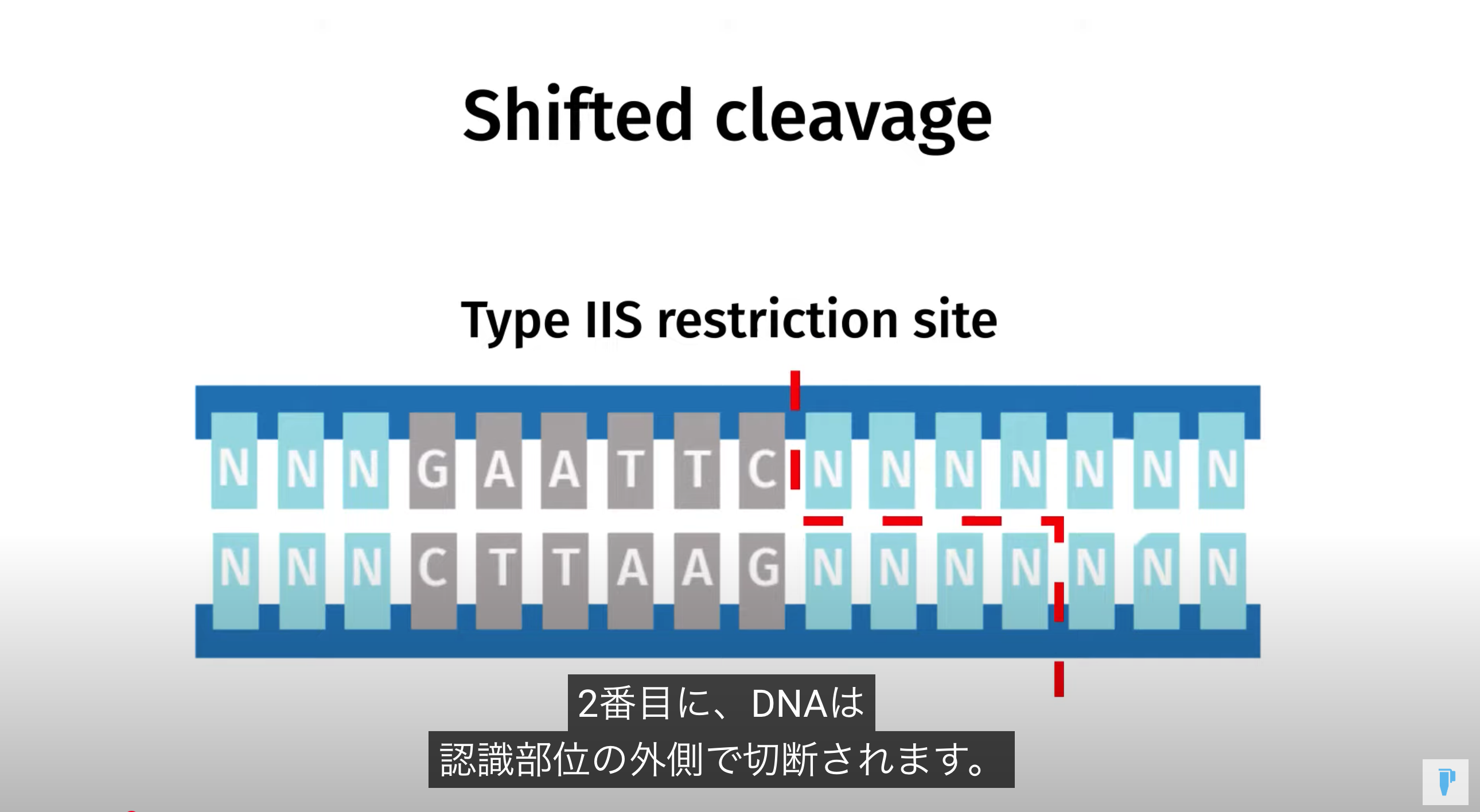
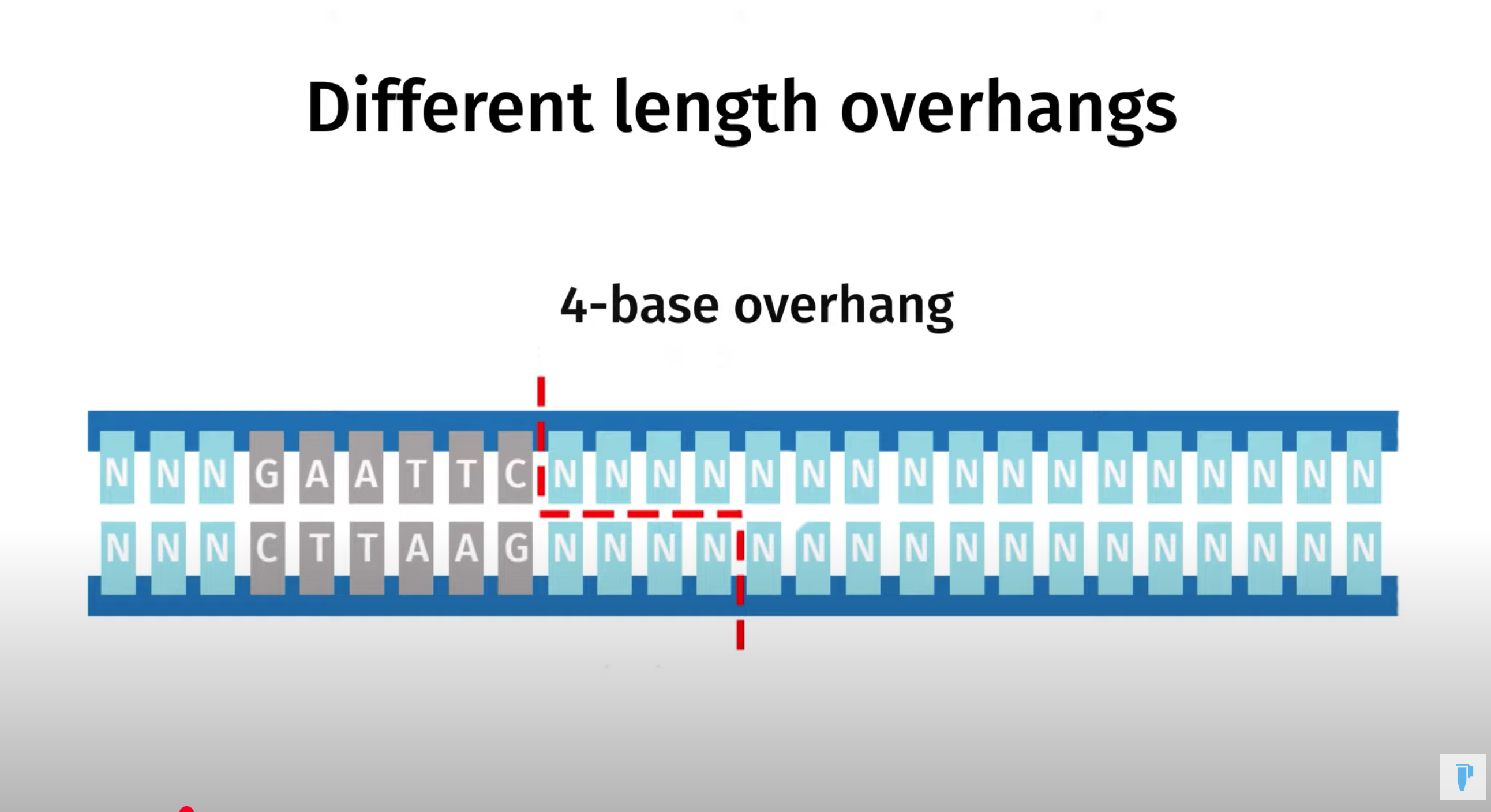
Golden Gate Assembly is a method that employs Type IIS restriction enzymes to cleave DNA and efficiently ligate multiple fragments using custom-designed overhangs
※ An overhang refers to the single-stranded extension of DNA that protrudes from one strand when the DNA is cleaved
Prepare each fragment so that it contains a Type IIS site—using primer design and PCR—ensuring the desired overhangs appear upon enzyme digestion
Next, combine all fragments in a single tube with the Type IIS restriction enzyme (e.g., BsaI) and T4 DNA ligase, enabling digestion and ligation to occur simultaneously
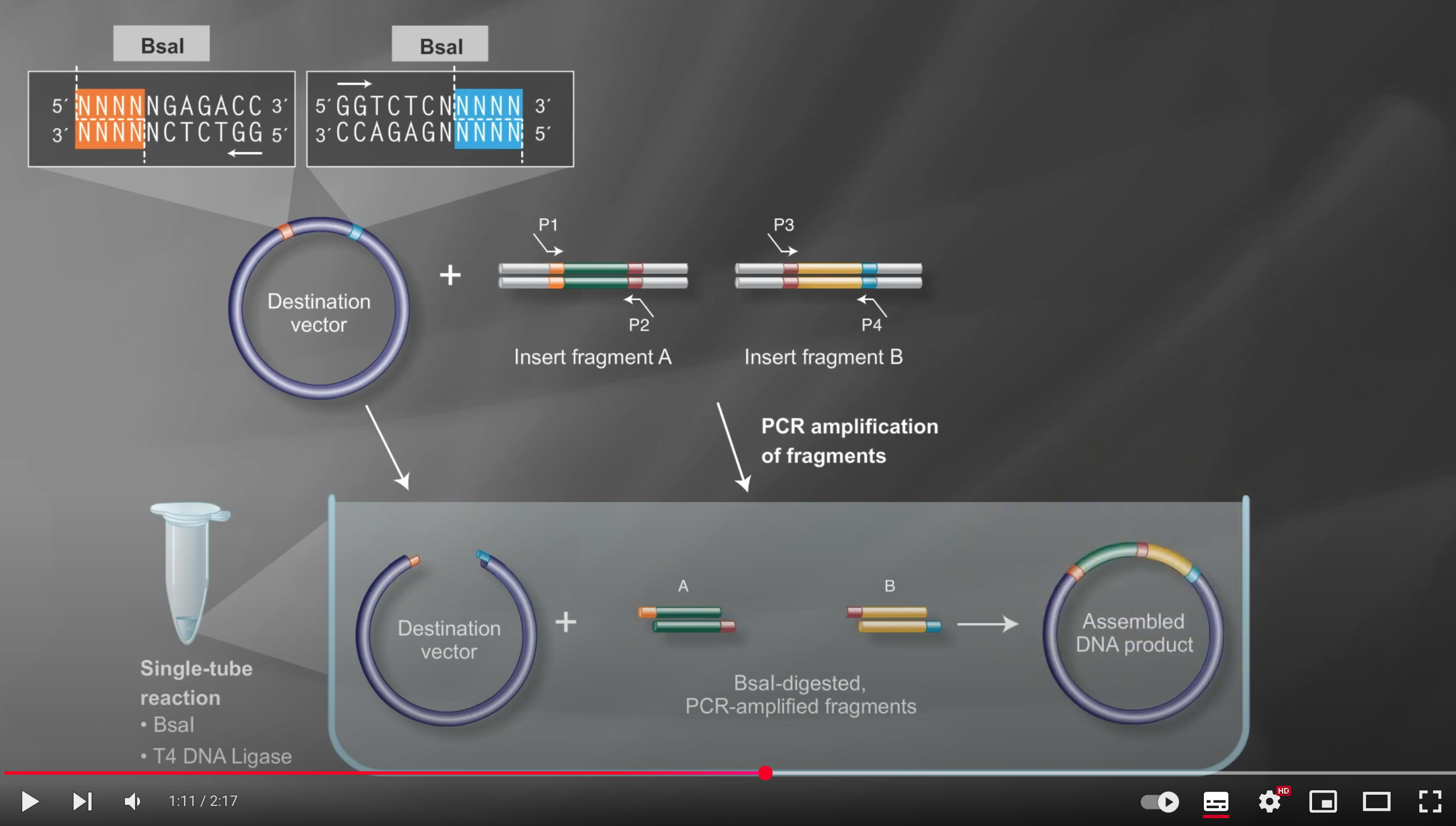
Under these reaction conditions, the enzyme repeatedly cuts the DNA to create transient overhangs, which then anneal, and the ligase seals the nicks
Incorrect assemblies are recut, so only the properly matching fragments remain ligated, resulting in an efficient multi-fragment assembly in the correct order
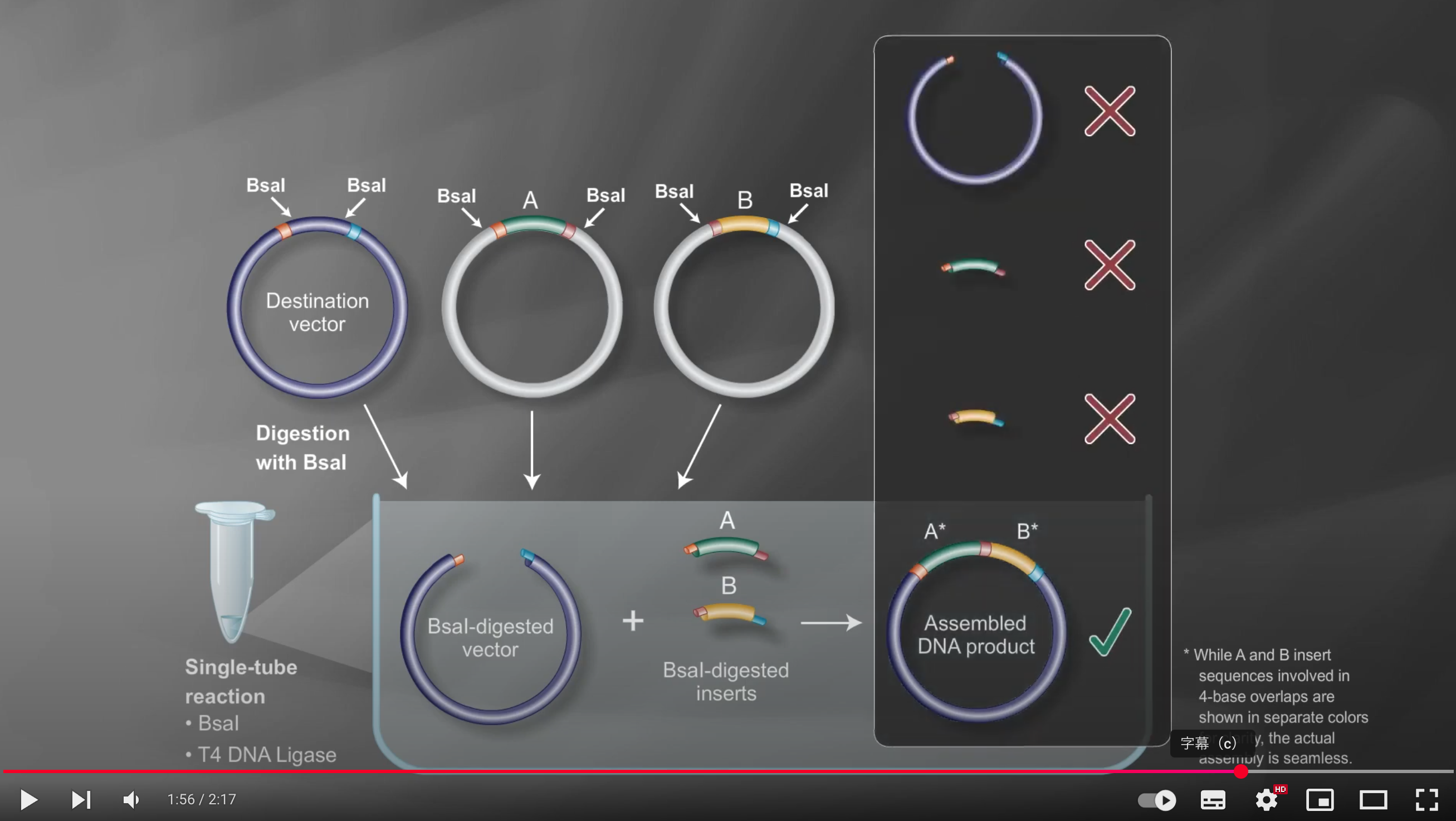
Because Type IIS enzymes cleave outside their recognition sequences, the final product is “scarless,” with no leftover restriction sites
After the reaction, transform the assembled plasmid into E. coli, and confirm the intended construct by sequencing if necessary