Week 02 HW: DNA Read, White, and Edit
Part 1: Benchling & In-silico Gel Art
Enzymes Cuts Temp. 1.1 2.1 3.1 4/CS
BamHI 5 37°C 75* 100* 100 100*
EcoRI 5 37°C 25 100* 50 50*
EcoRV 21 37°C 10 50 100 10
HindIII 7 37°C 25 100 50 50
KpnI 2 — Not available for this vendor
SacI 2 — Not available for this vendor
SacII 4 37°C 10 100 10 100
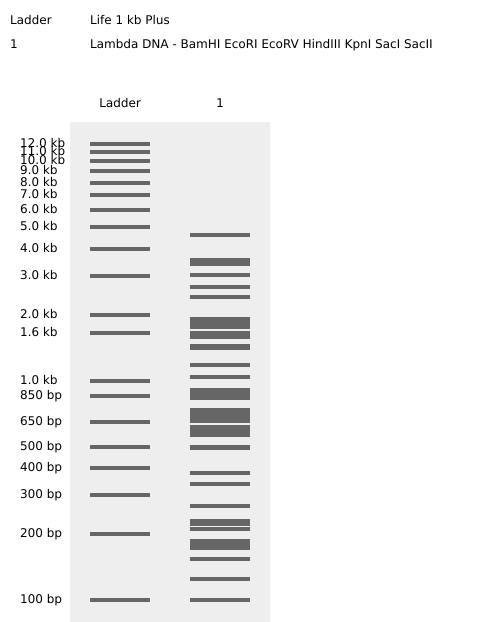

I have difficulty creating a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks. I will use this image as a placeholder for now. Sorry about that.
Part 3: DNA Design Challenge
3.1. Choose your protein.
I chose the cold shock protein CspA from Bacillus subtilis because it plays a role in cellular adaptation to low temperatures. Since my Week 1 homework focused on enhancing crop growth in short-season Canadian climates, selecting a protein involved in cold resilience aligns conceptually with my broader engineering interests.
I obtained the following protein sequence from Uniprot under the protein name Cold shock protein CspC, gene name cspC, BSU0520, CSPC_BACSU, P39158.
sp|P39158|CSPC_BACSU Cold shock protein CspC OS=Bacillus subtilis (strain 168) OX=224308 GN=cspC PE=1 SV=1 MEQGTVKWFNAEKGFGFIERENGDDVFVHFSAIQSDGFKSLDEGQKVSFDVEQGARGAQA ANVQKA
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
One result from Bioinformatics.org shows
reverse translation of sp|P39158|CSPC_BACSU Cold shock protein CspC OS=Bacillus subtilis (strain 168) OX=224308 GN=cspC PE=1 SV=1 to a 198 base sequence of most likely codons.
atggaacagggcaccgtgaaatggtttaacgcggaaaaaggctttggctttattgaacgc gaaaacggcgatgatgtgtttgtgcattttagcgcgattcagagcgatggctttaaaagc ctggatgaaggccagaaagtgagctttgatgtggaacagggcgcgcgcggcgcgcaggcg gcgaacgtgcagaaagcg
3.3. Codon optimization.
Codon optimization is necessary because different organisms preferentially use specific codons to encode the same amino acid. Although the genetic code is universal, codon usage bias affects translation efficiency and protein yield. I optimized the Bacillus subtilis expression sequence to improve expression efficiency in a soil bacterium relevant to my project proposed in Week 1 homework. The selection of Bacillus subtilis used help from GenAI. The tool is IDT Codon Optimization Tool https://www.idtdna.com/CodonOpt. The product type is gBlocks Gene Fragments. Restriction sites to avoid are Bsal, BsmBL, and Bbsl.
ATG GAG CAG GGC ACG GTT AAA TGG TTT AAT GCC GAG AAG GGC TTC GGT TTT ATC GAA AGA GAG AAT GGT GAT GAC GTT TTT GTG CAT TTT AGC GCA ATC CAA TCC GAC GGT TTC AAA TCT TTG GAT GAG GGT CAG AAG GTG TCT TTC GAT GTG GAA CAA GGC GCT CGC GGA GCG CAG GCT GCG AAT GTG CAG AAA GCG
3.4. You have a sequence! Now what?
I will explore the cell-free method, as it does not require living organisms, so that the system can be more controlled. In this way, cell-free expression systems could synthesize the protein directly from the DNA template without using living cells.
Part 4: Prepare a Twist DNA Synthesis Order
https://benchling.com/s/seq-zDyiwqJu2nW3ipK5tFql?m=slm-UFVFDHppIaBpufyDyHeC
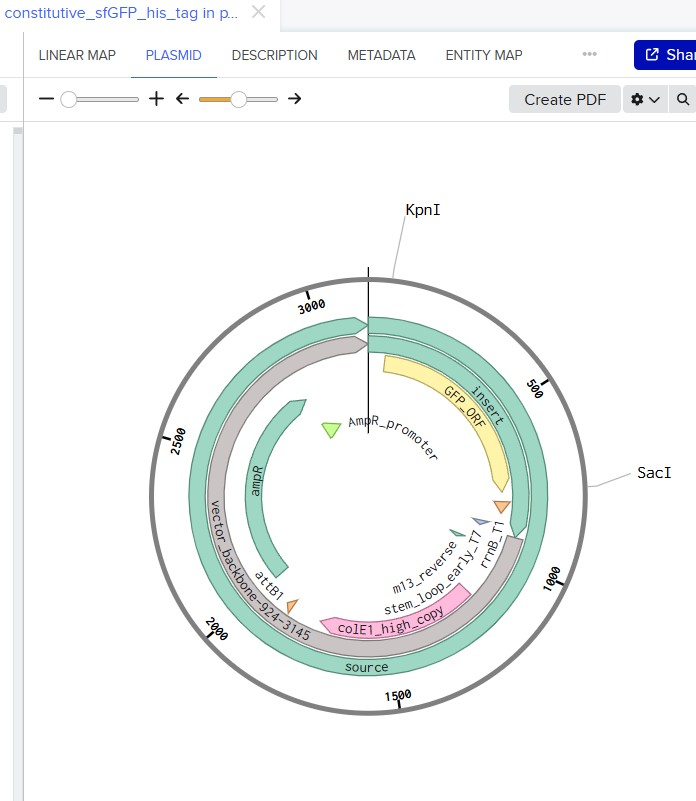

Part 5 — DNA Read / Write / Edit
I will have to revisit this part, once I go through the lecture again. Currently, the answers have relied heavily on Google and GenAI.
5.1 DNA Read
(i) What DNA would you want to sequence and why?
I would want to sequence DNA used in DNA-based digital data storage, which was used in the homework example. This would allow verification of synthesis accuracy, detection of errors introduced during storage or replication, and complete recovery of the original encoded information.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
(I sought help from ChatGPT to answer this question.) Illumina Sequencing by Synthesis (SBS) due to the parallel sequencing of fragment libraries and accuracy.
Also answer the following questions:
Is your method first-, second- or third-generation or other? How so?
- Second-generation due to the generated short reads.
What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
- DNA molecules (synthetic oligo libraries) as input, adapter ligation to both ends, and PCR to create clusters.
What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
- DNA fragments bind to a flow cell and form clusters by bridge amplification.
What is the output of your chosen sequencing technology?
- FASTQ files with short reads.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I would synthesize DNA constructs encoding digitally compressed cultural content because it enables long-term digital archiving.
(ii) What technology or technologies would you use to perform this DNA synthesis and why? Also answer the following questions:
I would use chip-based solid-phase oligonucleotide synthesis, such as the platforms developed by Twist Bioscience and others. These use phosphoramidite chemistry in a massively parallel microarray format.
What are the essential steps of your chosen sequencing methods? -A DNA chain, protected nucleotides, and deprotection and coupling cycles.
What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability? -error and length
5.3 DNA Edit
(i) What DNA would you want to edit and why?
I would like to continue my proposed project in Week 1 homework and add this week’s work on Bacillus subtilis.
(ii) What technology or technologies would you use to perform these DNA edits and why? Also answer the following questions:
I would like to employ CRISPR-Cas9, as mentioned in Professor Church’s slide. It is a tool that is frequently mentioned in the lecture, and I wish to explore more.
How does your technology of choice edit DNA? What are the essential steps? -CRISPR-Cas9 uses a short guide RNA (gRNA) to direct Cas9 to a specific DNA sequence. Once bound, Cas9 introduces a double-strand break at the target site.
What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing? -1) Guide RNA(s) targeting loci involved in repair or replication; 2) Cas9 or base/prime editor constructs; 3) Donor DNA templates for HDR when introducing specific sequences; and 4) Competent cells capable of taking up editing components
What are the limitations of your editing methods (if any) in terms of efficiency or precision? -Unintended edits can occur at similar sequences.
Week 1 Homework: Lecture 2 Preparation:
Professor Jacobson’s lecture:
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
The error rate is 1:100,000 or 1,000,000 throughput: 10mS per Base Addition (according to the slide and Google).
Human Genome Length: A single human diploid cell contains roughly 6 million base pairs. The discrepancy without correction, if the error rate were 100,000, every cell division would introduce roughly 100,000-600,000 errors. This is drawn from Google AI Overview.
One way biology deals with the discrepancy is intrinsic proofreading (3’-5’ exonuclease), according to Google.
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
(Directly from Google search)
Different ways: synonymous codons, mathematical combinations, and redundant genetic code.
Reasons not working: condon usage bias, protein folding defects, mRNA stability and decay, mRNA structure, and transcriptional efficiency.
Dr. LeProust’s lecture:
What’s the most commonly used method for oligo synthesis currently?
Solid-phase phosphoramidite chemistry. (Google)
Why is it difficult to make oligos longer than 200nt via direct synthesis?
It is difficult because the product’s yield and purity decrease exponentially with each added base. (Google)
Why can’t you make a 2000bp gene via direct oligo synthesis?
This is because conventional automated phosphoramidite synthesis is limited by the efficiency of nucleotide coupling and error accumulation. (Google)
George Church’s lecture (choose one to answer):
(Using Google & Prof. Church’s slide #4) What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”
The ten essential amino acids are Arginine, Histidine, Isoleucine, Leucine, Lysine, Methionine, Phenylalanine, Threonine, Tryptophan, and Valine. The effect is that humans may be able to control nature if they control these ten amino acids, which is not true. (Google)
Note: I apologize for heavily relying on Google, but I am struggling to understand the material by myself.