Week 06 HW: Genetic Circuit Part 1
Part 1: DNA Assembly
What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The key components include Phusion DNA polymerase (providing extreme fidelity), optimized reaction buffer (providing the necessary environment for enzyme activity), Magnesium Chloride (supporting DNA polymerase activity), and Deoxynucleotide Triphosphates (dNTPs - building new DNA strands). The purpose of the master mix is to reduce the risk of contamination, save time, and minimize mixing errors.
What are some factors that determine primer annealing temperature during PCR?
Primer melting temperature is the most crucial factor. Primer length, Guanine and Cytosine content, primer concentration, and primer pair match are also primary factors that determine primer annealing temperature during PCR.
There are two methods from this class that create linear fragments of DNA: PCR and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Both PCR and restriction enzyme digests are fundamental to DNA manipulation and analysis. In general, unless PCR, Restriction Enzyme Digest doesn’t have Magnesium or dNTP as reagents, add the enzyme last in a reaction setup, require a lower temperature, and last shorter than RCR. In general, PCR is chosen when the target sequence is unknown or needs to add specific restriction sites. Restriction Digests are chosen when there is already a cloned construct and a need for high fidelity (can’t risk introducing mutations)
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
There are a few considerations of primer design, including high-fidelity amplification, purification, and accurate quantification to ensure optimal overlap and molar ratios.
How does the plasmid DNA enter the E. coli cells during transformation?
By passing through transient pores created in the cell membrane. A heat shock creates a thermal imbalance that facilitates the movement of negatively charged DNA through the pores.
Describe another assembly method in detail (such as Golden Gate Assembly)
Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
This response is based on SnapGene Academy. Golden Gate Assembly is a one-pot, one-reaction cloning procedure created by Carola Engler et al. in 2008. The method takes advantage of Type IIS restriction enzymes (e.g. BsaI), which cleave DNA outside their recognition sequences. The result is an ordered assembly of a vector and one or more DNA fragments.
Golden Gate assembly can be split into two distinct steps that occur within the same reaction: 1) Type IIS restriction enzyme digestion, and 2) DNA ligation. It works by mixing the following components into a single reaction tube: Destination vector DNA insert(s) (e.g. amplicon or pre-cloned), Type IIS restriction enzyme (e.g. BsaI), T4 DNA ligase, and Reaction buffer. Golden Gate Assembly.jpg
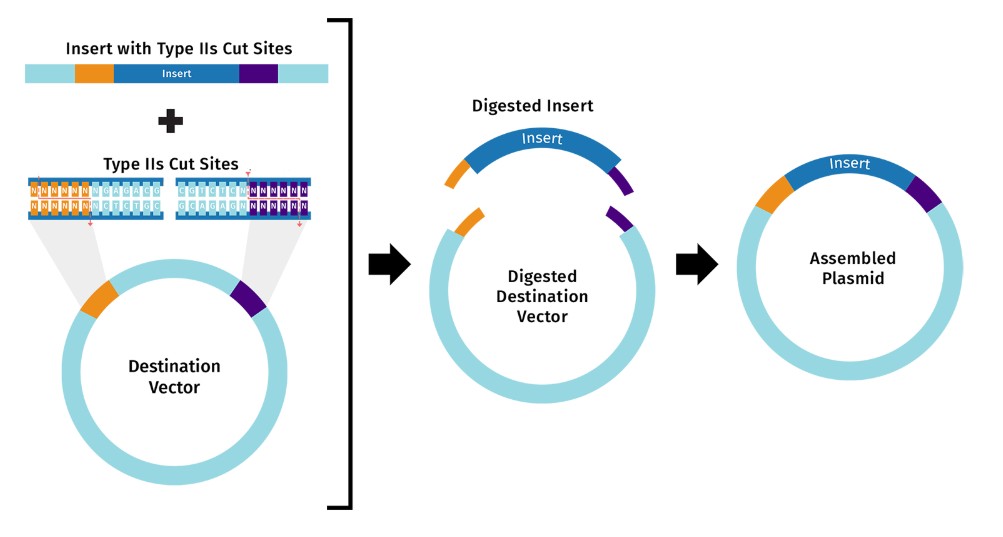

Model this assembly method with Benchling or Asimov Kernel!
Modeling with benchling is based on the response from Google AI Overview. Modeling Golden Gate Assembly in Benchling is streamlined through their built-in Assembly Wizard, which supports the one-pot, scarless assembly of multiple DNA fragments using Type IIS restriction enzymes (e.g., BsaI, BsmBI). Benchling, in collaboration with New England Biolabs (NEB), allows to simulate digestion and ligation, generate specific 4-base overhangs, and design the necessary primers to add Type IIS sites to inserts.
Part 2: Asimov Kernel
The results are recorded in my notebook “SK”. I am not sure about the third construct, as the results from the first two are not satisfactory.