Subsections of Homeworks
Week 1 HW: Principles and Practices


First, describe a biological engineering application or tool you want to develop and why.
I want to develop a closed loop pipeline for peptide engineering that uses Feynman–Kac steering to control diffusion-based protein generation at inference time. The goal is to go beyond zero-shot prediction and instead build an automated engineering cycle that repeatedly:
- proposes peptide/mini-protein candidates,
- captures experimental readout (binding, activity, stability, etc.),
- turns those measurements into reward signals,
- uses FK steering to bias the next round of generative sampling toward better candidates without needing to retrain the underlying diffusion model
This is inspired by FK-steering approach which wraps a diffusion protein generator with a sampling scheme so trajectories are continuously reweighted toward user-defined rewards, which in this case, is the experimental readout.
Peptides are a good choice for this project as they are often fast to synthesize and test, making them compatible with iterative lab loops. However, many properties of peptides we care about (solubility, stability, expression, off-target behavior) can be hard to optimize from prediction alone so a wet-lab loop is attractive. Functionally, they can serve as binders, inhibitors, diagnostic reagents, or modular parts in synthetic biology pipelines.
As a concrete MVP within this class, I hope to learn how to perform the wet lab experiments associated to this project and finish at least 1 cycle. In the medium term, I would like to run comparisons between different computational approaches like simple finetuning or RL. In the long term, I would like to utilizie this method to discover therapeutic proteins.
Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals.
Closed loop design could be repurposed to create harmful biomolecules. Governance should reduce the probability of both deliberate misuse and accidental creation of dangerous function. Thus, one major goal would be to prevent misuse. As sub goals, the following may be good options:
Ensure the system does not optimize toward harmful or restricted targets/functions.
Reduce the chance that hazardous sequences are synthesized without review.
Ensure that there are audit trails and responsible-use norms.
Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”).
I propose three governance actions spanning institutional review, synthesis controls, and a logging infrastructure.
Option 1: Institutional Review
- Purpose: Add structured risk assessment before synthesis, target changes, or new reward functions in academic protein design projects.
- Design: One-page checklist covering target protein class, reward function, synthesis plan, and screening. Projects triggering high-risk criteria (regulated agents, virus optimization) require formal oversight.
- Assumptions: Small review gates and enforce good record keeping practices
- Risks: Could push students to under-report. If too strict, it may slow down R&D>
Option 2: Synthesis Controls
- Purpose: Require synthesis vendors to use functional or homology-based screening.
- Design: Institutions only purchase from vendors who screen orders and verify customers
- Assumptions: It is possible to do screening meaningfully well to reduce risk
- Risks: The screening needs to be highly accurate to catch edge cases which could have massive negative effects
Option 3: Logging Infrastructure
- Purpose: Create a secure shared database that tracks when AI tools generate protein designs
- Design: Logging of AI tools and cross-referencing of orders.
- Assumptions: Confidentiality and transparency is balanced
- Risks: Security or confidentiality concerns from hacking or from sensitive IP
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | 2 | 1 | 2 |
| • By helping respond | 1 | 2 | 1 |
| Foster Lab Safety | |||
| • By preventing incident | 1 | 2 | 3 |
| • By helping respond | 1 | 2 | 1 |
| Protect the environment | |||
| • By preventing incidents | 2 | 2 | 3 |
| • By helping respond | 2 | 2 | 1 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 2 | 2 | 2 |
| • Feasibility? | 1 | 2 | 3 |
| • Not impede research | 1 | 2 | 1 |
| • Promote constructive applications | 1 | 2 | 2 |
Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties.
In order of priority:
- Option 1: This option can arguably be implemented the fastest. MIT already has the safety infrastructure (IBC, EHS) to build on. As a leading institution in AI protein design, MIT can set standards that others follow. A well-designed, lightweight review process could become a widely adopted model.
- Option 2: The existing government framework provides a strong template with vendor screening, customer verification, and reporting requirements. However, this depends on federal action and industry cooperation beyond MIT’s control. MIT can help by researching better screening algorithms and influencing governement gold standards.
- Option 3: If this project becomes a widely used system, tracking who designed what becomes relatively easy. However, the system will have to be designed extremely well to be scalable, secure, transperent yet confidential.
Tradeoffs:
- Speed vs. safety
- Open science vs. closed science
- Transparent vs. confidential
Key Uncertainties:
- How manageable it is to manually gate research directions.
- How well screening actually works against deliberate misuse.
- How feasible it is to design a logging system everyone is happy with.
Reflecting on what you learned and did in class this week, outline any ethical concerns that arose, especially any that were new to you. Then propose any governance actions you think might be appropriate to address those issues. This should be included on your class page for this week.
Unfortunately, I was ill this week so I was not able to attend class.
Week 2 HW: DNA Read, Write, & Edit
Gel Electrophoresis Designs
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks
I have created an image of mount fuji with clouds in the sky. I have inverted the image so it is easier to visualize.
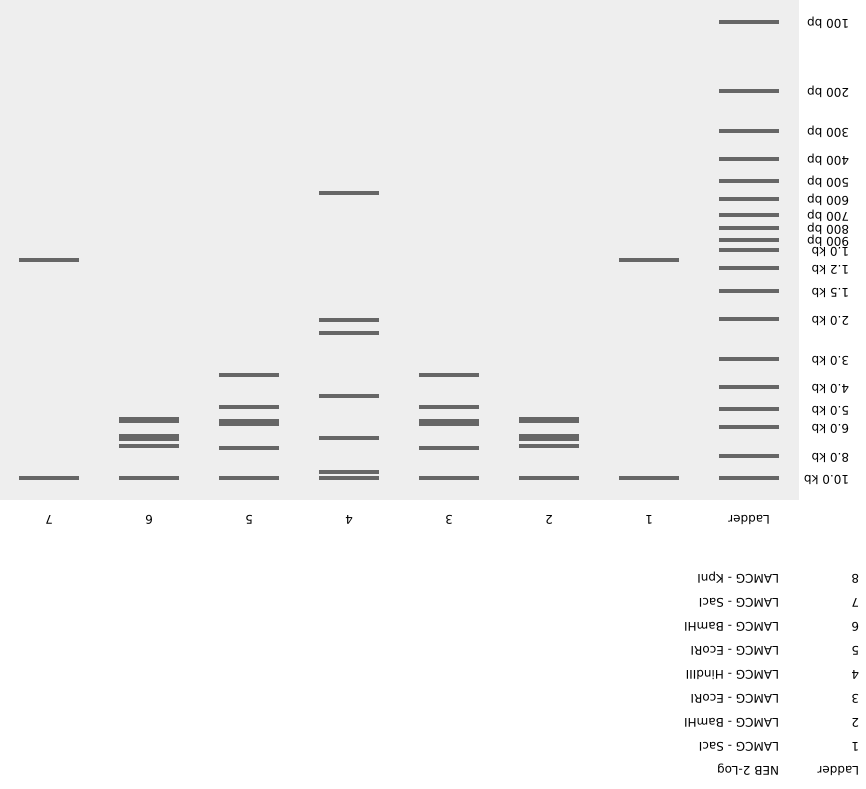

Note: Since we worked in groups during lab this week, we created a different design than the one shown above for the lab activity.
DNA Design Challenge
Choose your protein.
RES-701-3 is a tiny natural protein made by soil bacteria (Streptomyces). It belongs to a family called lasso peptides, named because their structure looks like a lasso or slipknot. The tail of the protein threads through a loop, creating a knot that is extremely hard to unravel.
This knotted shape makes lasso peptides unusually tough. They resist being broken down by digestive enzymes, heat, and harsh chemical environments. These are properties that most proteins lack, and that make them attractive as potential drugs.
RES-701-3 blocks a receptor on the surface of blood vessel cells called the endothelin type B receptor (ETB). The endothelin system controls blood vessel tightening and relaxation, and becomes dysregulated with age, contributing to high blood pressure and vascular disease. RES-701-3 acts as an inverse agonist, meaning it blocks the receptor and pushes toward a less active state than its resting baseline.
In nature, the bacteria makes this peptide in two parts:
- Leader section:
MSDITLTPMDLLDLDELAAGGGRSTARE - Core peptide sequence:
GNWHEPEIDGWNPHGW
The core is removed from the leader with an enzyme, which makes it active.
Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The nucleotide sequence of the leader and the core is shown respectively.
- Leader:
ATGAGCGATATTACCCTGACCCCGATGGATCTGCTGGATCTGGATGAACTGGCTGCTGGTGGTGGTCGTAGCACCGCTCGTGAA - Core:
GGTAACTGGCATGAACCGGAAATTGATGGTTGGAACCCGCATGGTTGGTAA
Codon optimization.
Due to evolution, different species have different codons it uses frequently and has abundant matching transfer RNAs for, and codons it rarely uses and has few tRNAs for. RES-701-3 comes from Streptomyces and strongly prefers codons loaded with G and C. Twist has a Streptomyces coelicolor for codon optimization.
However, it’s worth mentioning that in a 2025 paper by Shihoya et al. paper, they used Streptomyces venezuelae as organism and achieved the highest reported yields. If I was in a real drug development setting, I might go with this.
Here is the codon optimized variant for both leader and core together:
You have a sequence! Now what?
I have listed the Promoter, RBS, Start Codon, Coding Sequence, His Tag, Stop Codon, Terminator as well as the reagents needed below.
Promoter: The **ermE*p promoter is supposed to be the most widely used for gene expression in Streptomyces.
Ribosome Binding Site: We’re using Shine Dalgarno (SD) sequence, AAGGAG, which is supposed to be a good RBS for streptomyces with leaders. It is supposed to be positioned 6 to 10 nucleotides upstream of the start codon, so we will use 7 nucleotides. We’re going to put two spacers before and after the SD sequence, CGACG and ACAC.
Start Codon: This is just going to be the usual ATG.
Coding Sequence: We are going to put both of our leader and core peptide sequence together here.
His tag: This is a short string of six histidine amino acids added to the protein so you can fish it out of a mixture using a nickel column. The histidines stick to nickel, letting you pull your protein out of everything else the cell makes. However, in practice, apparently this is not actually good to put on for RES-701-3 because it would interfere with binding the ETB receptor.
Stop Codon: TGA tells the ribosome to stop building the protein here. TGA is the preferred stop codon in Streptomyces because it is relatively speaking, GC-rich, matching the organism’s DNA preferences as discussed before. For example, typical stop codon is TAA.
Terminator: Tells the cell’s RNA-copying machinery to stop making mRNA. Without it, the cell would keep reading past your gene into random neighboring DNA. We’re using the fd terminator from a bacteriophage which is commonly used in Streptomyces expression vectors.
Reagents
In order to produce these proteins we also need to use some enzymes to be used as reagents, namely, LasB1, LasB2 and LasC. For this lasso peptide, LasB1 binds the leader, delivers the whole precursor to LasB2 which cuts the leader off, and then LasC closes the ring on the core. It doesn’t seem easy to order the reagents so it seems like this peptide wouldn’t be a great choice for the class. In addition, the yield is optimized by using Streptomyces venezuelae, which is also not too common.
Prepare a Twist DNA Synthesis Order
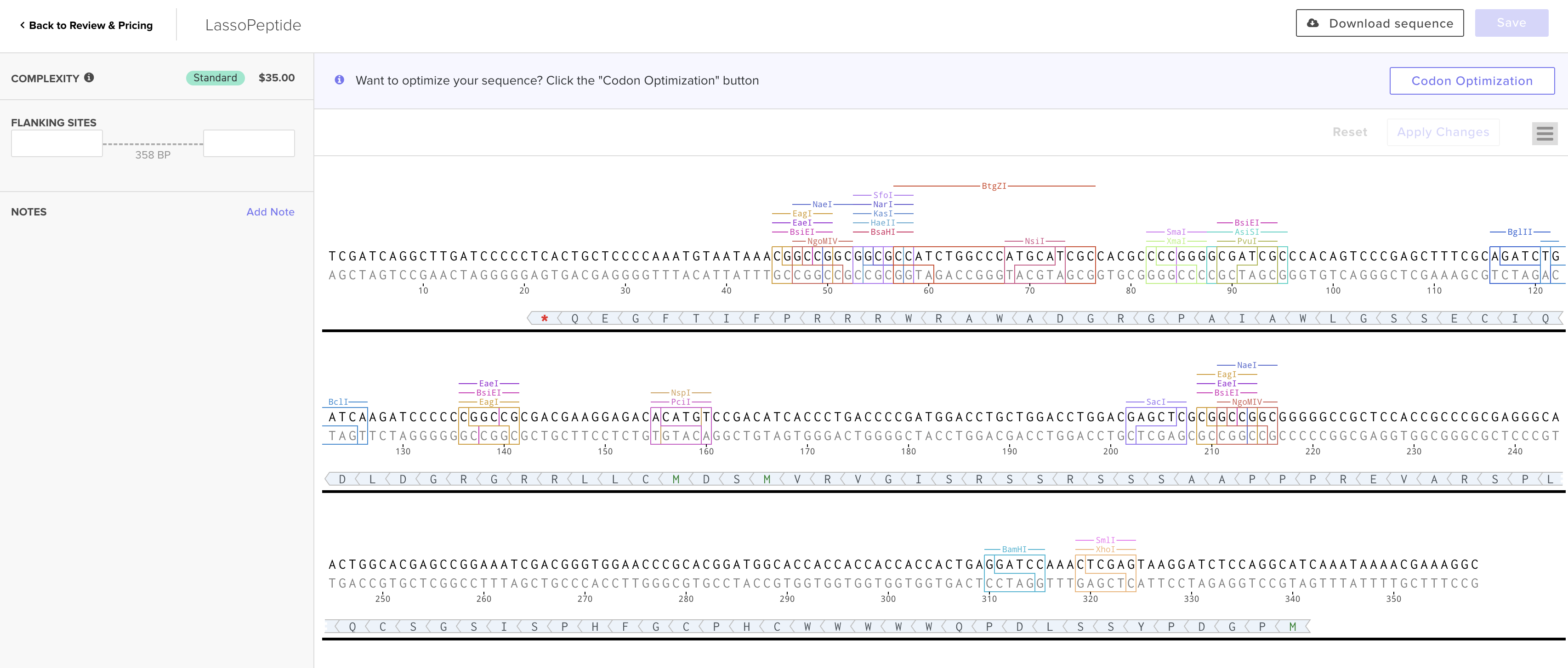
I prepared the lasso peptide order. Here is a picture of the expression cassette below in benchling.
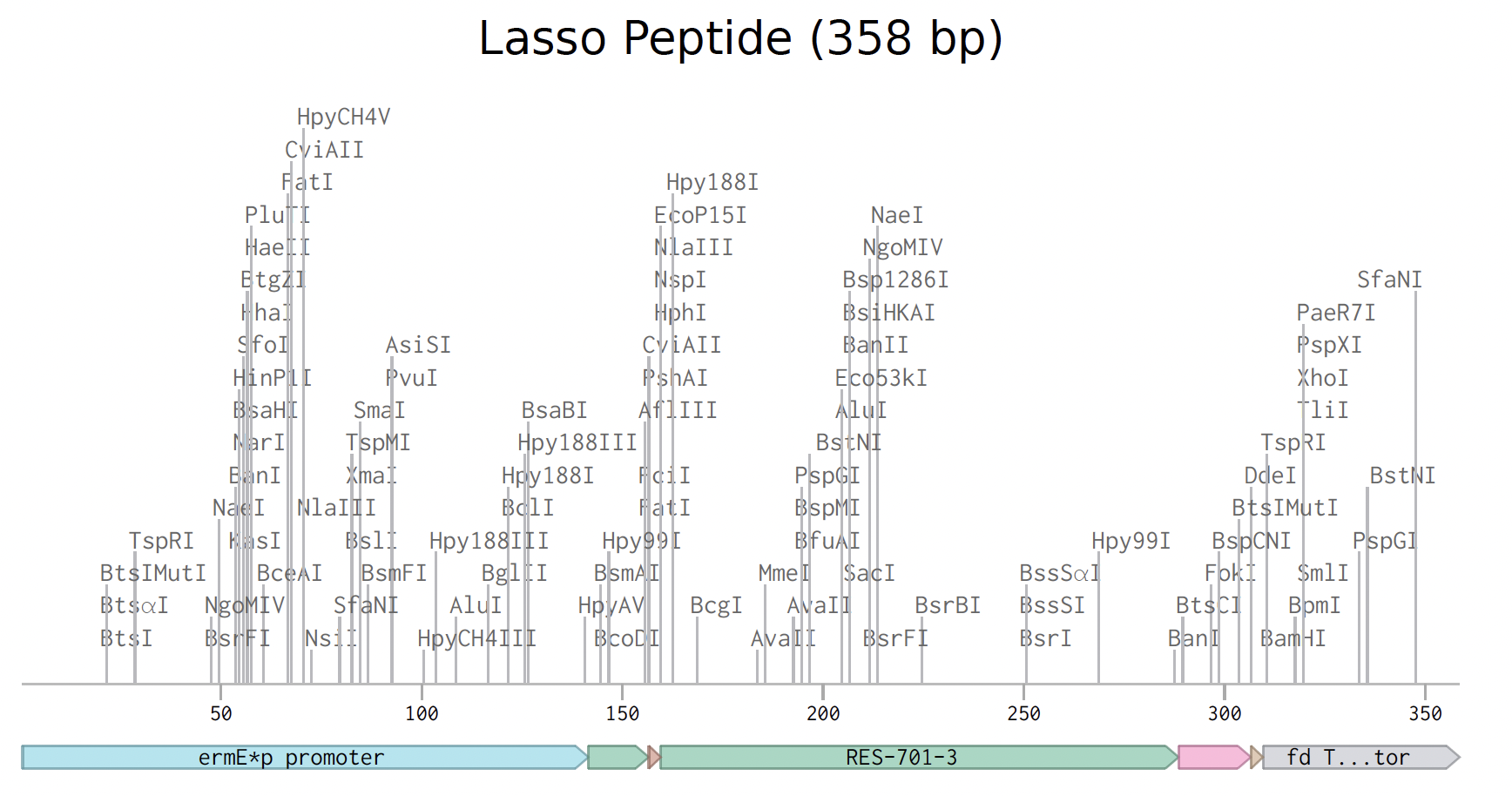

Instead of a clonal gene, I used gene fragments because they work better Streptomyces as an organism rather than e coli, which are the standard cloning vectors.

DNA Read/Write/Edit
5.1 DNA Read
What DNA would you want to sequence (e.g., read) and why?
I would want to sequence the whole genomes of all ~6,000 mammalian species. The largest current collection of mammalian genomes is the Zoonomia project, which contains around 250 whole genomes along with known maximum lifespan data for most of these species. However, expanding this to cover all mammals—paired with their maximum lifespan records—would allow us to train computational models that identify DNA patterns predicting how long a species can live. In short, more genomes means better predictions about which parts of DNA are linked to longevity.
In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Illumina short-read sequencing (second-generation): This produces highly accurate short reads (~150–300 base pairs) and is great for spotting small genetic differences between species.
Is your method first-, second-, or third-generation?
I am using both second-generation Illumina. First-generation refers to older Sanger sequencing, which reads one fragment at a time and is too slow and expensive for whole genomes. Second-generation sequences millions of short fragments in parallel, making it fast and cheap.
What is your input? How do you prepare your input?
The input is genomic DNA extracted from tissue or blood samples of each mammalian species. The essential preparation steps are:
- DNA extraction: Isolate high-quality DNA from the biological sample.
- Fragmentation: Break the DNA into smaller pieces.
- Adapter ligation: Attach short known DNA sequences adapters to the ends of each fragment so the sequencing machine can recognize and handle them.
- PCR amplification (Illumina): Make many copies of each fragment to boost the signal.
- Quality check: Verify the library is the right size and concentration before loading it onto the sequencer.
What are the essential steps of your chosen sequencing technology? How does it decode bases (base calling)?
Fragmented DNA is attached to a glass surface flow cell, amplified into clusters, and then sequenced one base at a time. In each cycle, a fluorescently labeled nucleotide is added, a camera captures which color lights up at each cluster where each of the four bases has a different color, and the machine records the base. This process repeats hundreds of times to read out each fragment.
What is the output?
The output is digital sequence files, typically in FASTQ format, containing millions of reads—short or long strings of A, T, C, and G letters—along with quality scores indicating how confident the machine is about each base call. These reads are then assembled and aligned computationally to reconstruct each species’ complete genome.
5.2 DNA Write
What DNA would you want to synthesize (e.g., write) and why?
Based on the sequencing data above, I would use trained computational models to predict specific DNA sequences associated with high maximum lifespan. I would then synthesize these predicted longevity-linked sequences—for example, specific gene variants or regulatory elements found in long-lived species like bowhead whales or naked mole-rats—so they can be tested in cell cultures or animal models. The goal is to move from computational prediction to experimental validation: do these DNA sequences actually promote cellular health and longevity?
What technology or technologies would you use to perform this DNA synthesis and why?
- Oligonucleotide synthesis from Twist Bioscience: For building short to medium DNA fragments (up to a few thousand base pairs). These companies use chemical synthesis on microchips to build many sequences in parallel, making it fast and affordable.
- Gibson Assembly or Golden Gate Assembly: For stitching shorter synthesized fragments together into larger constructs. These are molecular cloning methods that use enzymes to join DNA pieces seamlessly.
What are the essential steps of your chosen synthesis method?
- Sequence design: Use computational models to design the target DNA sequences, optimizing codon usage for the target organism and avoiding problematic features (e.g., long repeats, extreme GC content).
- Oligonucleotide synthesis: Short single-stranded DNA pieces (oligos, ~50–200 bases) are built base by base using chemical reactions on a solid support. Each cycle adds one nucleotide at a time.
- Assembly: Overlapping oligos are combined and joined enzymatically into longer double-stranded fragments (a few hundred to a few thousand base pairs).
- Cloning: The assembled fragments are inserted into a circular DNA carrier (plasmid vector) and introduced into bacteria, which copy the DNA as they grow.
- Verification: The final constructs are sequenced to confirm the correct sequence was built.
- Large construct assembly: Multiple verified fragments are stitched together using Gibson Assembly or Golden Gate Assembly to create larger genetic constructs.
What are the limitations of your synthesis method in terms of speed, accuracy, and scalability?
- Speed: Synthesizing and assembling long constructs (>10,000 base pairs) can take weeks, since each fragment must be built, verified, and then joined together step by step.
- Accuracy: Chemical synthesis introduces errors at a rate of roughly 1 in 200 bases per oligo. While these errors are corrected through screening and verification, it adds time and cost.
- Scalability: Very long or repetitive sequences are difficult to synthesize because the oligos may misassemble or fold in unwanted ways. Sequences with extreme GC content are also harder to build reliably.
5.3 DNA Edit
What DNA would you want to edit and why?
I would want to edit specific genes in model organisms (such as mice) to replace their native sequences with the longevity-associated sequences identified from the analysis above. For example, if the computational model predicts that a certain variant of a DNA repair gene is linked to longer lifespan in mammals, I would edit a mouse’s genome to carry that variant. This would let us test whether swapping in these predicted “long-life” DNA variants actually extends lifespan or improves age-related health outcomes like cancer resistance or cellular repair.
What technology or technologies would you use to perform these DNA edits and why?
I would use CRISPR-Cas9 gene editing, because it is the most precise, versatile, and widely used genome editing tool available. It can make targeted changes at specific locations in the genome of living cells and organisms, and it works well in mammalian systems including mice.
How does your technology edit DNA? What are the essential steps?
- Target selection: Identify the exact location in the genome you want to edit.
- Guide RNA design: Design a short RNA sequence that matches the target DNA site.
- Cutting: The Cas9 protein, guided by the RNA, binds to the matching DNA site and makes a double-strand break.
- Repair: The cell’s natural repair machinery fixes the break. If a DNA template with the desired new sequence is provided alongside the CRISPR components, the cell can use it as a blueprint to incorporate the new sequence, called homology-directed repair.
- Screening: Edited cells are sequenced to confirm the desired change was made correctly.
What preparation do you need to do, and what is the input?
- Design inputs: The target DNA sequence, a custom guide RNA matching that sequence, and a DNA donor template carrying the desired new sequence flanked by regions that match the area around the cut site.
- Molecular inputs: Cas9 protein or mRNA, synthesized guide RNA, donor template DNA, and delivery reagents.
- Biological inputs: Target mouse cell.
What are the limitations of your editing method in terms of efficiency or precision?
- Off-target edits: The guide RNA can sometimes bind to similar sites elsewhere in the genome, causing unintended cuts and mutations.
- Low HDR efficiency: Only a fraction of edited cells may carry the precise desired change, requiring extensive screening.
- Delivery challenges: Getting CRISPR components into every target cell efficiently, especially in living animals, remains difficult. Some tissues are harder to reach than others.
Week 3 HW: Lab Automation
I have included my OpenTron work, answers to post-lab questions and 3 early stage project ideas in the Week 3 lab section.
Week 4 HW: Protein Design
Part A: Conceptual Questions
Why do beta-sheets tend to aggregate?
A beta-strand forms when a protein’s backbone — the repeating NH–Cα–CO chain shared by every amino acid — stretches out into a nearly flat zigzag. When two or more of these strands line up next to each other and link through hydrogen bonds (where an N–H on one strand bonds to a C=O on the neighbor), you get a beta-sheet.
The strands on the outer edges still have a full row of exposed N–H and C=O groups, allowing another strand to be added, and so on — this is why beta-sheets tend to aggregate.
What forces pull sheets together?
Hydrophobic effect — the biggest driver. In a beta-strand, side chains stick out alternately above and below the sheet. Since many side chains are hydrophobic, two sheets stack such that the greasy surfaces face inward.
Hydrogen bonding — gives the structure its regularity. Each new strand that joins the sheet edge contributes roughly one H-bond per amino acid along its length. Individually, H-bonds in water are not enormously strong (breaking one with a neighbor just lets you form one with water instead), but across a strand of ten or more residues they add up meaningfully.
Van der Waals packing — stabilizes sheets that have stacked together. These forces are much weaker and shorter-range, arising from temporary, fluctuating dipoles.
Part B: Protein Analysis and Design
Briefly describe the protein you selected and why you selected it.
I selected a monoclonal antibody for the following reasons:
- Ability to target specific proteins on cell surfaces with extreme precision, directly applicable to therapeutics
- Ability to recruit the immune system (via Fc region) to destroy tagged cells, combining specificity with immune effector functions
- Can be engineered with ML and computational methods for improved binding affinity and reduced immunogenicity
- Compared to small molecule drugs, highly specific to their target with fewer off-target effects
For this exercise, I chose trastuzumab, famous for revolutionizing the treatment of HER2-positive breast cancer. It is a humanized IgG1 monoclonal antibody that binds to the extracellular domain IV of HER2 (human epidermal growth factor receptor 2), blocking receptor dimerization and downstream signaling that drives tumor growth.
How long is it? What is the most frequent amino acid?
The full trastuzumab IgG has 2 heavy chains (449 aa each) and 2 light chains (214 aa each), for a total of ~1,326 amino acids and ~148 kDa.
However, the crystal structure (PDB: 1N8Z) contains only the Fab fragment (the antigen-binding portion), which includes:
Heavy chain Fab (chain B, 220 aa):
Light chain (chain A, 214 aa):
- Combined Fab length: 434 amino acids
- Most common amino acid: S (Serine), appearing 60 times
How many protein sequence homologs are there for your protein?
Because trastuzumab is a humanized antibody with conserved IgG1 framework regions, BLAST returns a very large number of homologs — antibodies share ~70–90% identity in their framework regions. A BLAST search of the heavy chain Fab against UniProt returns over 250 homologs. The variable CDR (complementarity-determining region) loops are what make trastuzumab unique in its HER2 specificity.
When was the structure solved? Is it a good quality structure?
Good quality = good resolution. Smaller is better (benchmark: 2.70 Å).
- Deposited: 2002-11-21
- Released: 2003-02-18
- Published: Cho et al., Nature (2003) 421: 756–760
- Link: https://www.rcsb.org/structure/1N8Z
- Resolution: 2.52 Å — good quality, better than the 2.70 Å benchmark
Are there any other molecules in the solved structure apart from protein?
Yes. In addition to the 3 unique protein chains (light chain A, heavy chain B, and HER2 extracellular domain C), the structure contains:
| Molecule | Description | Copies |
|---|---|---|
| NAG | 2-acetamido-2-deoxy-β-D-glucopyranose (N-linked glycosylation sugar attached to HER2) | 2 |
| SO4 | Sulfate ion | 1 |
Does your protein belong to any structure classification family?
Yes. The overall complex is classified in the PDB under TRANSFERASE. The trastuzumab Fab itself belongs to the Immunoglobulin superfamily.
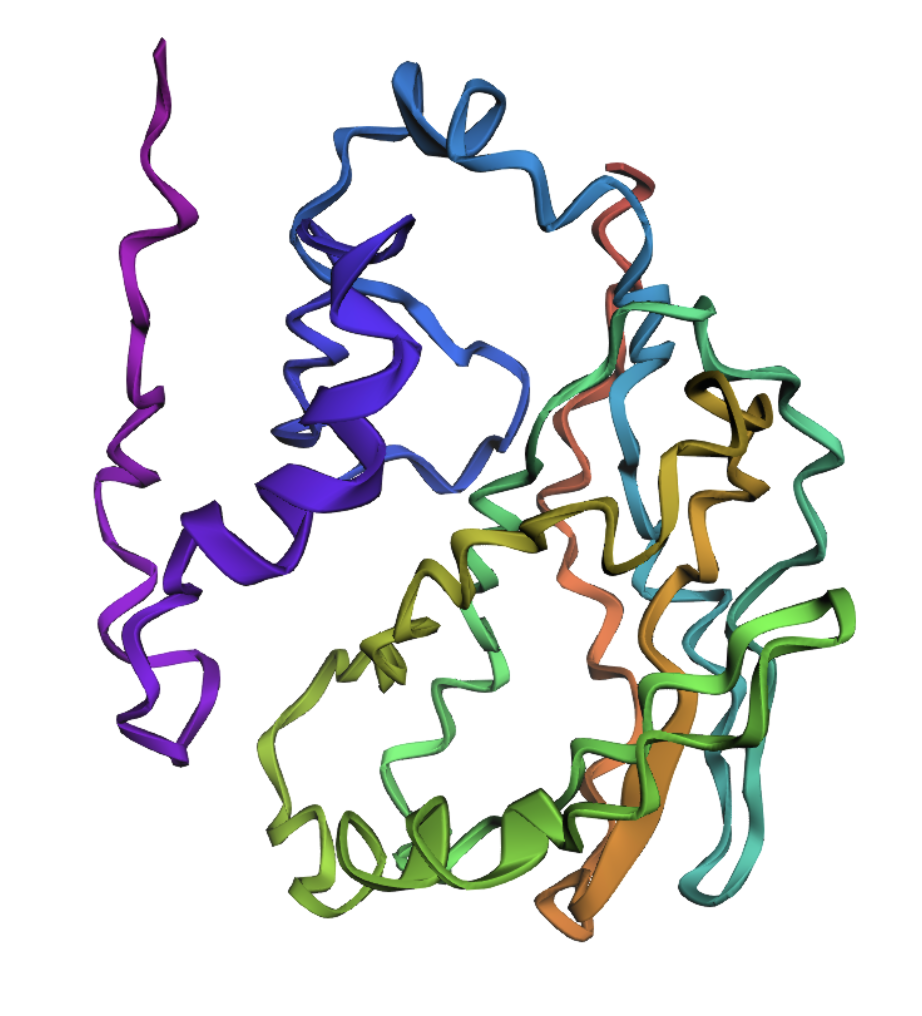
Visualize the protein as “cartoon”, “ribbon” and “ball and stick”
Cartoon
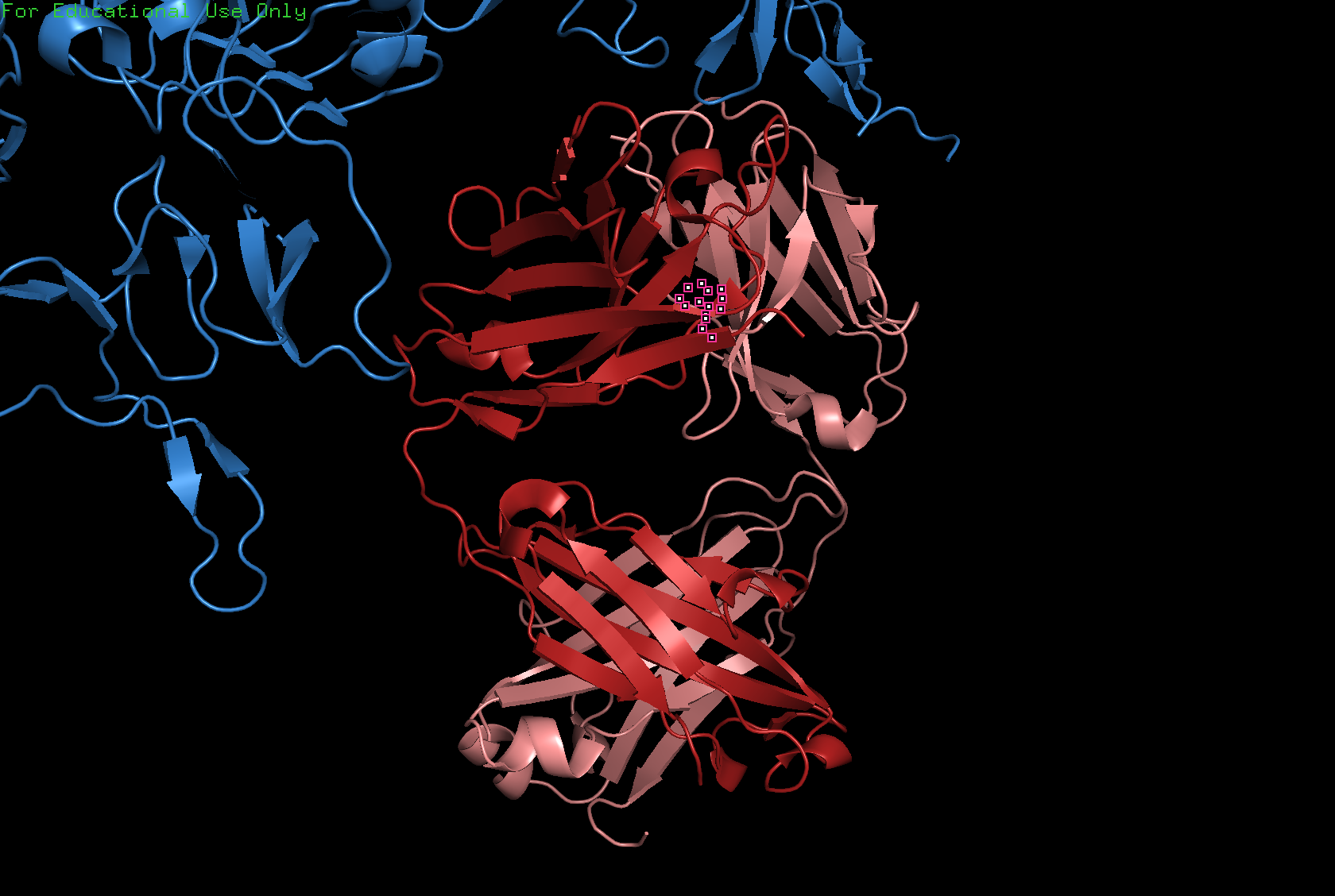

Ribbon

Ball and Stick

Color the protein by secondary structure. Does it have more helices or sheets?
The structure has more sheets than helices — specifically 215 atoms in sheets vs 30 atoms in helices.
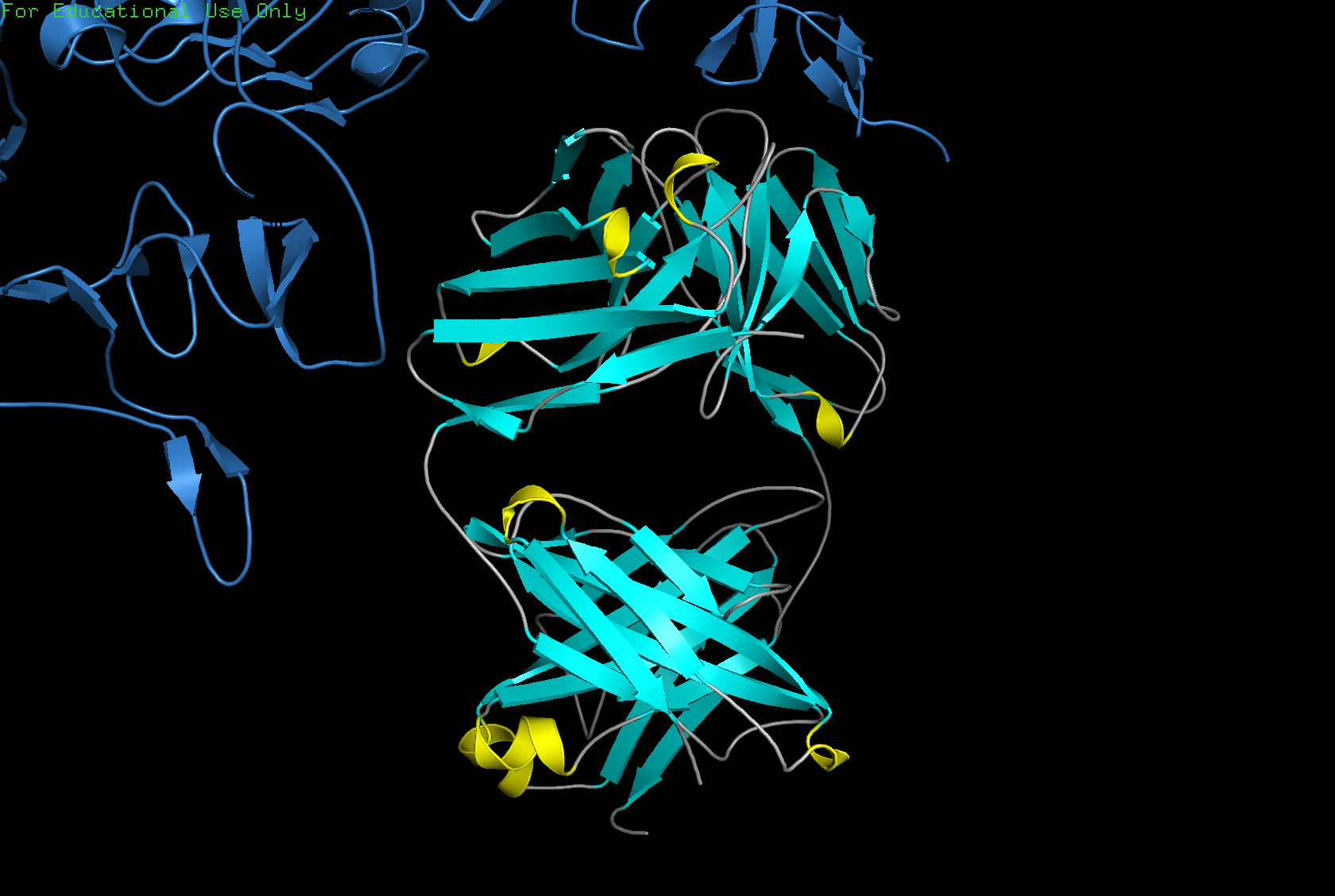

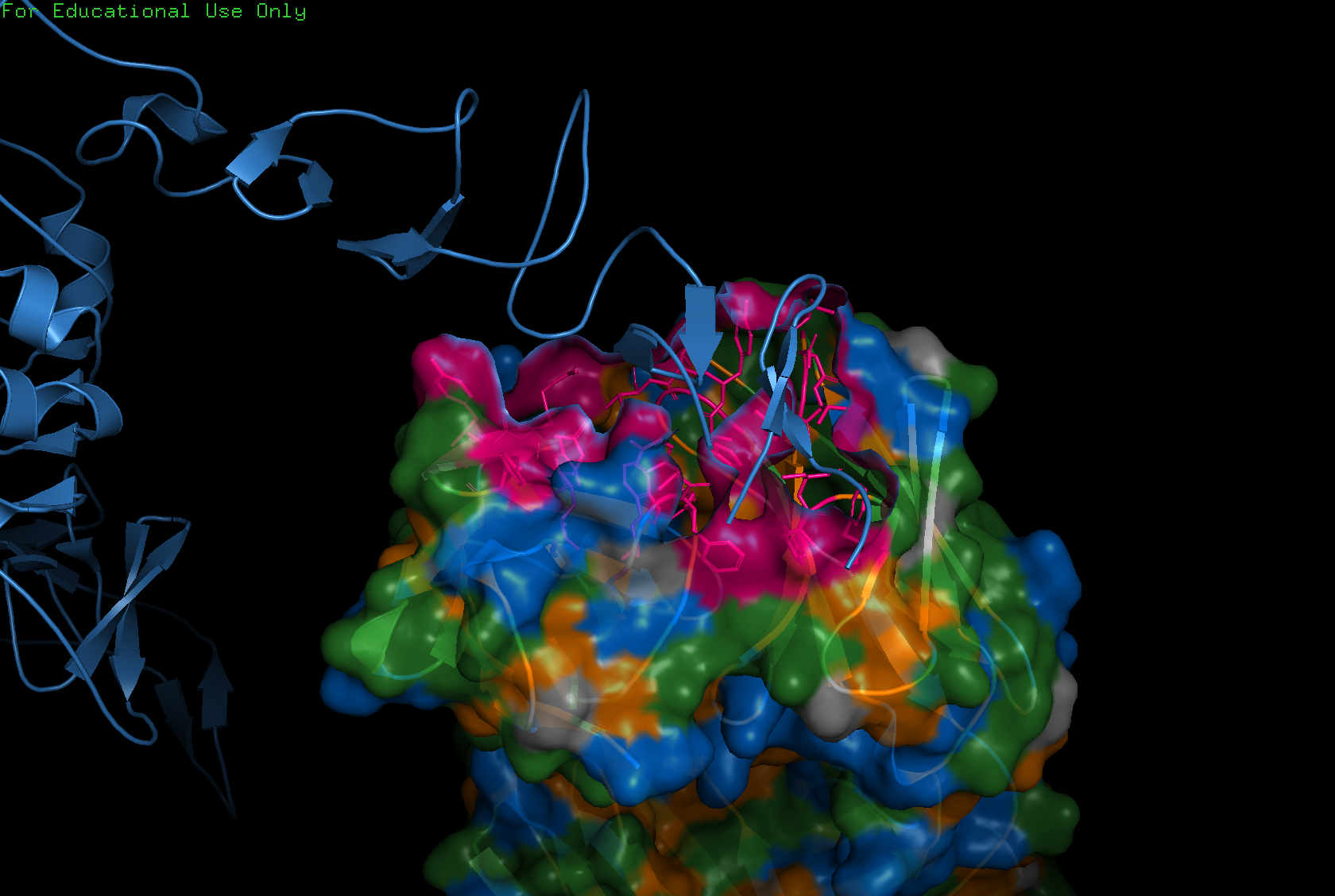
Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues?
Generally, proteins have a hydrophobic core with a hydrophilic surface, and trastuzumab follows this pattern. The immunoglobulin fold is a beta sandwich where:
- Hydrophobic residues (orange) point inward
- Hydrophilic residues (blue) point outward
(This is hard to see in the visualization because the inward and outward surfaces are not so distinct.)
However, the CDR loops — the tips that contact the target HER2 — are mixed: aromatic hydrophobics (Trp, Tyr) provide shape complementarity, while polar and charged residues form hydrogen bonds and salt bridges with the antigen.
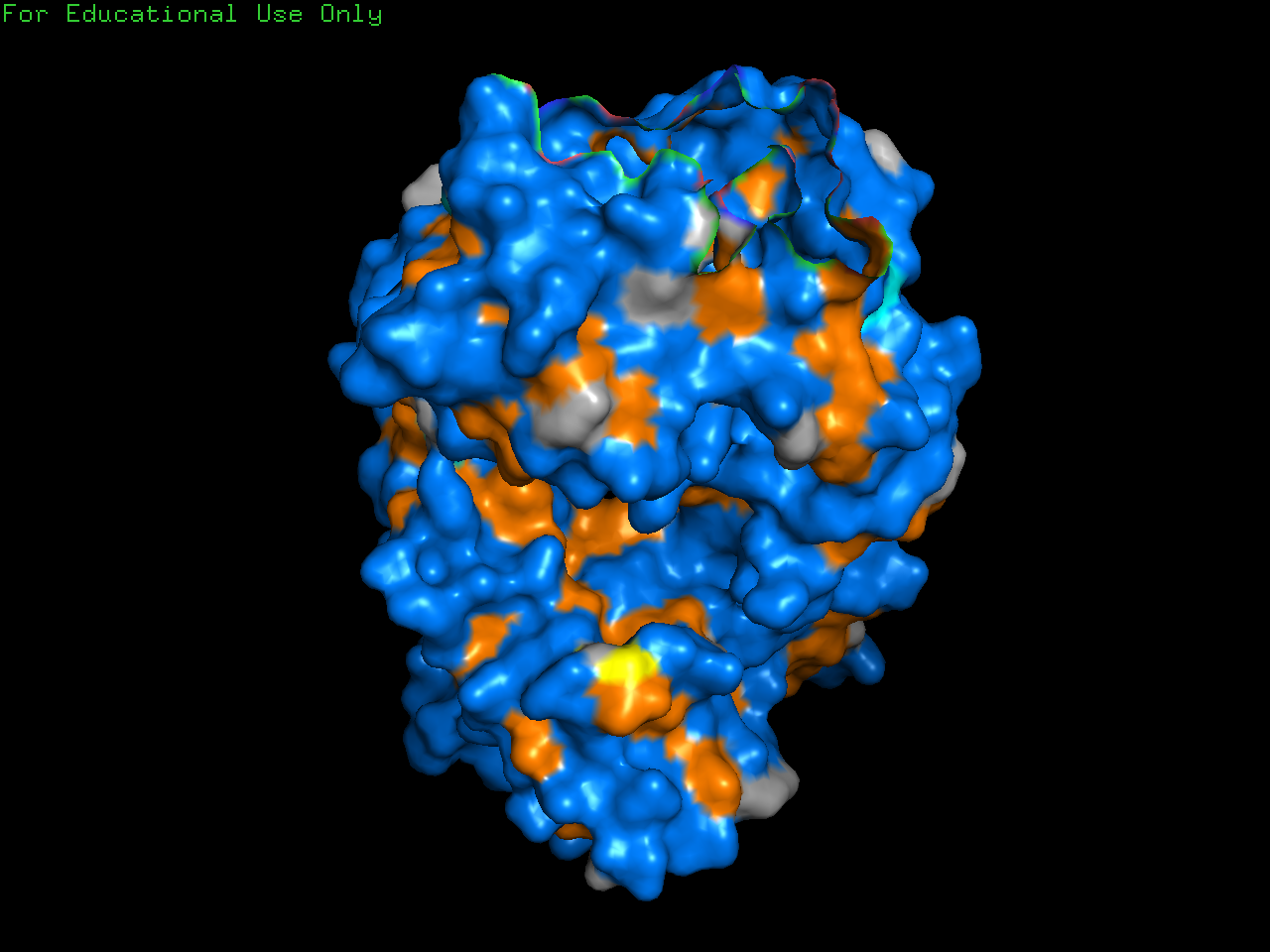

Visualize the surface of the protein. Does it have any “holes” (aka binding pockets)?
Yes, binding pockets are visible on the surface.

Part C: ML-Based Protein Design Tools
For this exercise, I chose the 6M0J SARS-CoV-2 Spike Receptor Binding Domain.
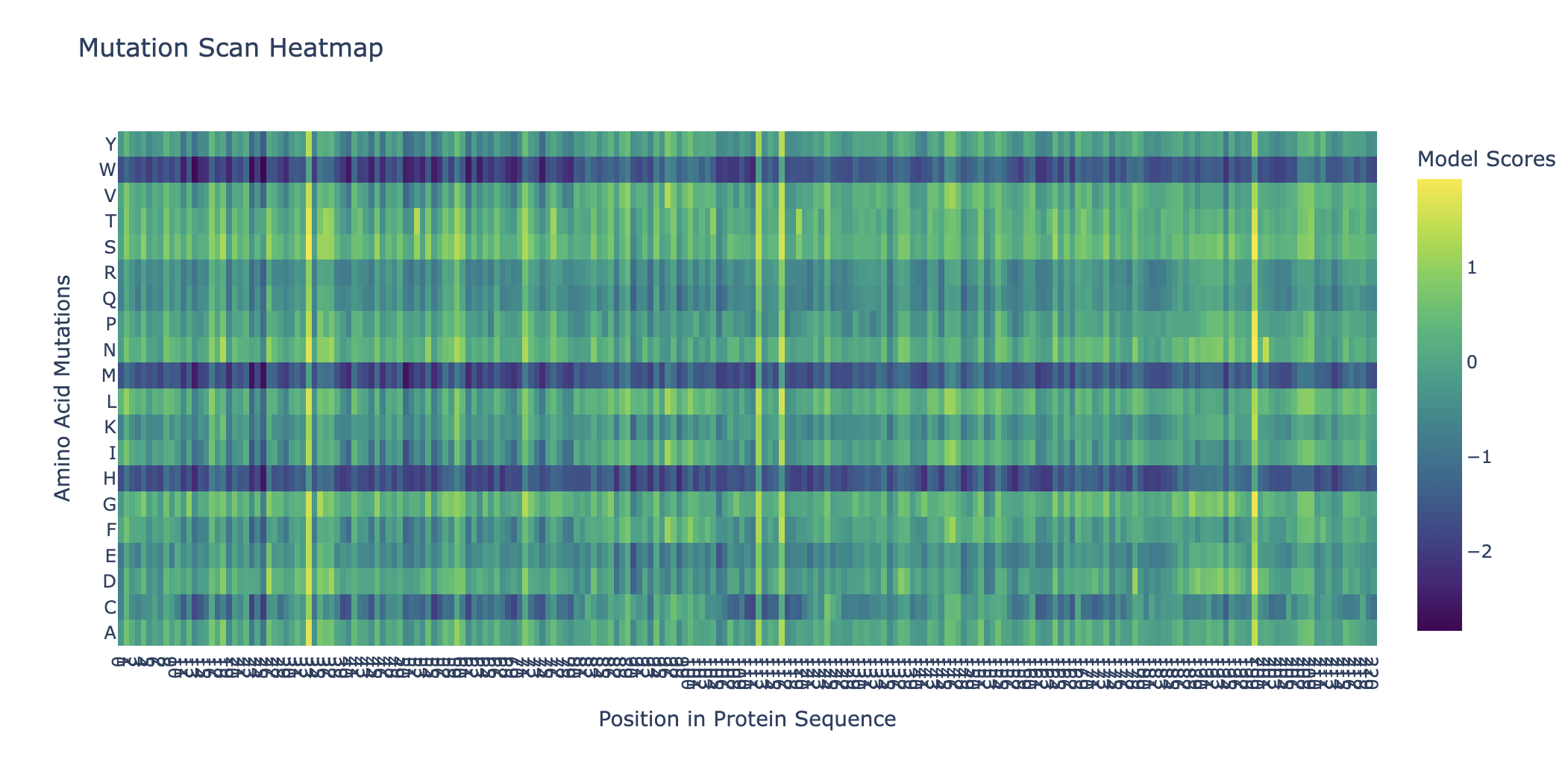
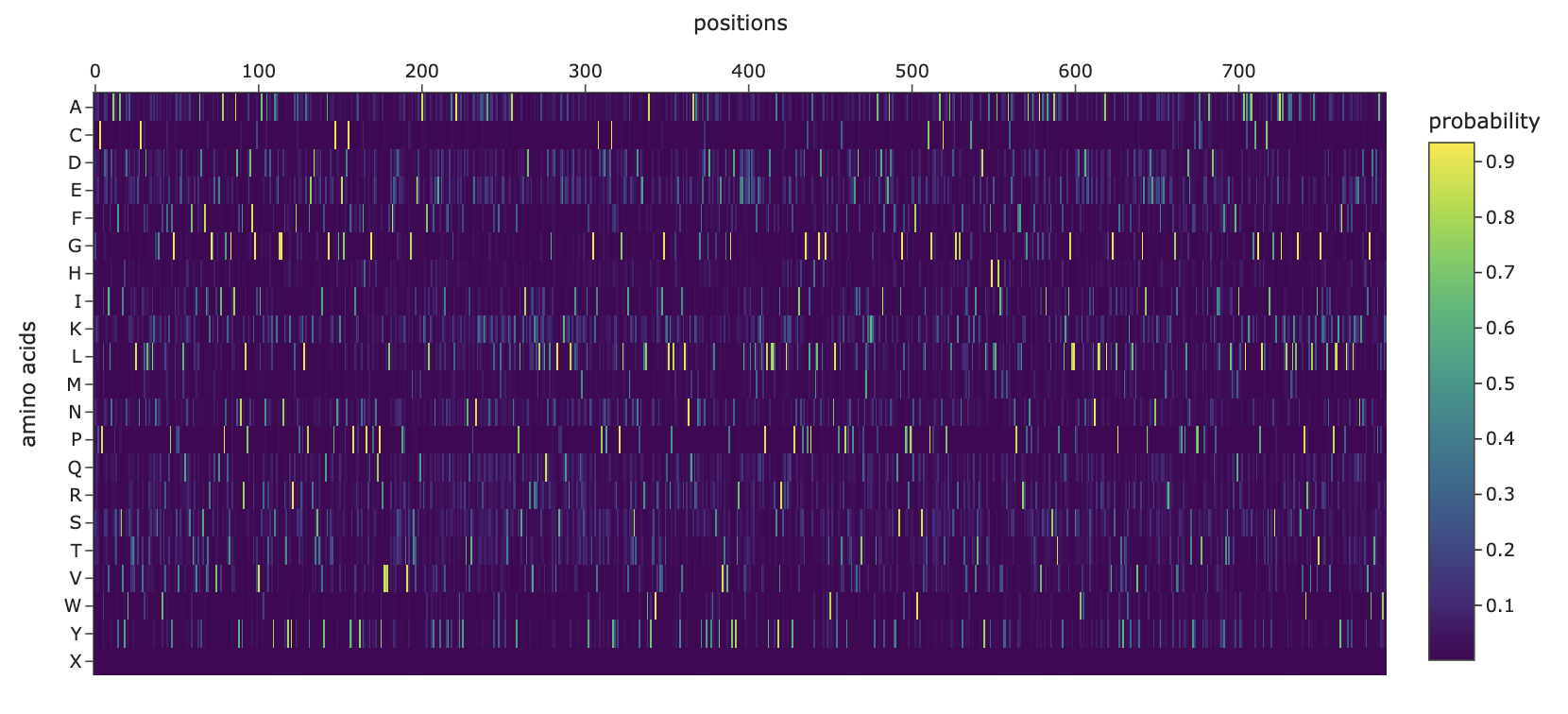
Deep Mutational Scans
Can you explain any particular pattern?
Horizontal patterns (rows): The rows for tryptophan (W), histidine (H), and methionine (M) are consistently darker across nearly all positions. These are large, bulky, or chemically complex amino acids that are difficult to accommodate at arbitrary positions without disrupting the protein’s fold. In contrast, small, simple amino acids like alanine or serine are more easily tolerated as substitutions, which is why their rows appear lighter overall.
Vertical patterns (columns): The most striking pattern is the dark purple vertical stripes at specific positions. These correspond to cysteine residues, which form disulfide bonds that hold the shape together so it can bind the human ACE2 receptor. Because ESM2 learned from millions of protein sequences that these cysteines are almost never substituted in nature, it heavily penalizes any mutation at those positions. The darkest scores appear when cysteine is mutated to something like tryptophan or proline, since these would not only break the disulfide bond but also create additional structural problems.

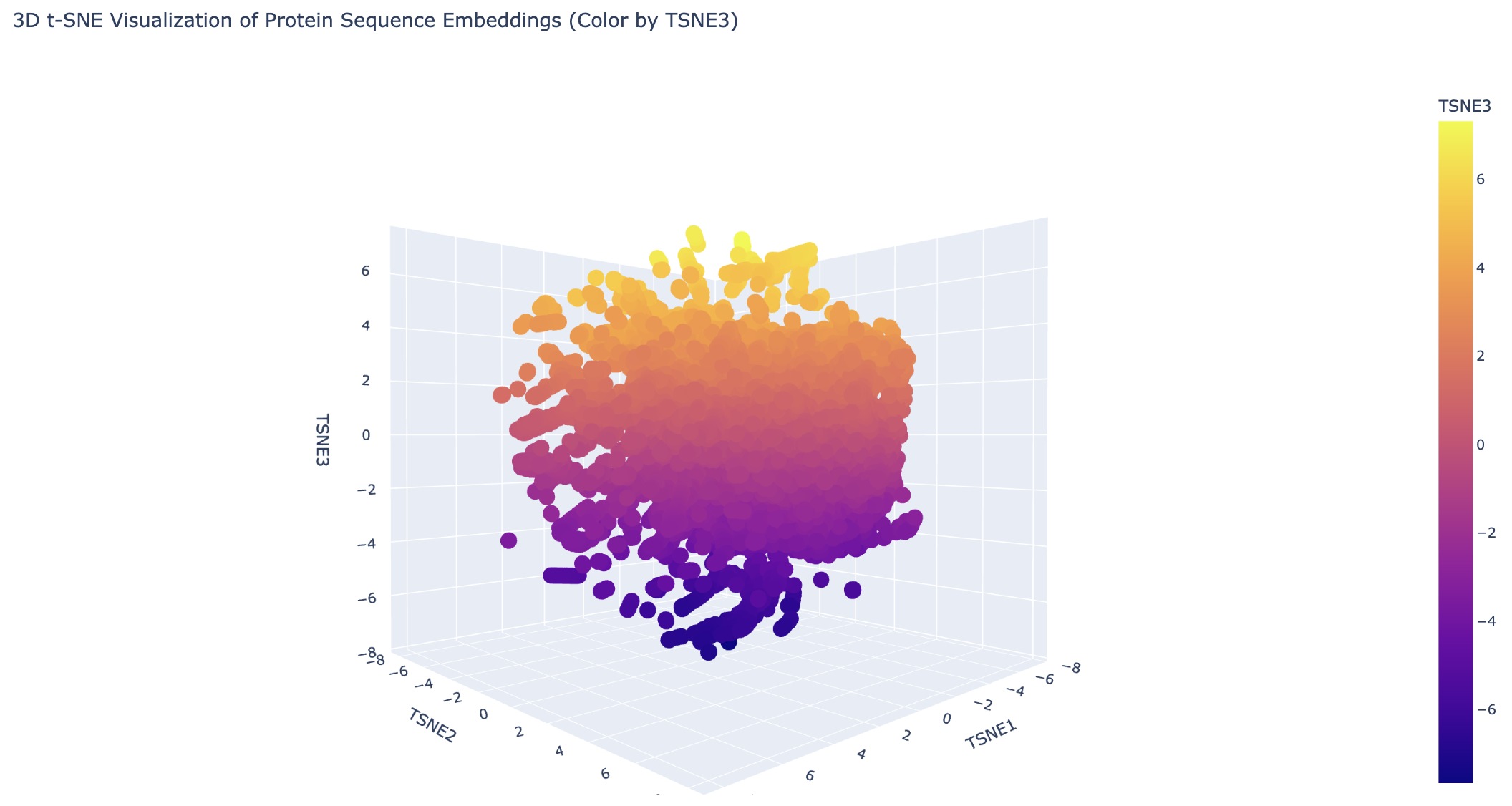
Latent Space Analysis
Analyze the different formed neighborhoods: do they approximate similar proteins?
Generally, the proteins are clustered tightly. There are some distinct clusters on the edges, which likely share a common evolutionary ancestor.

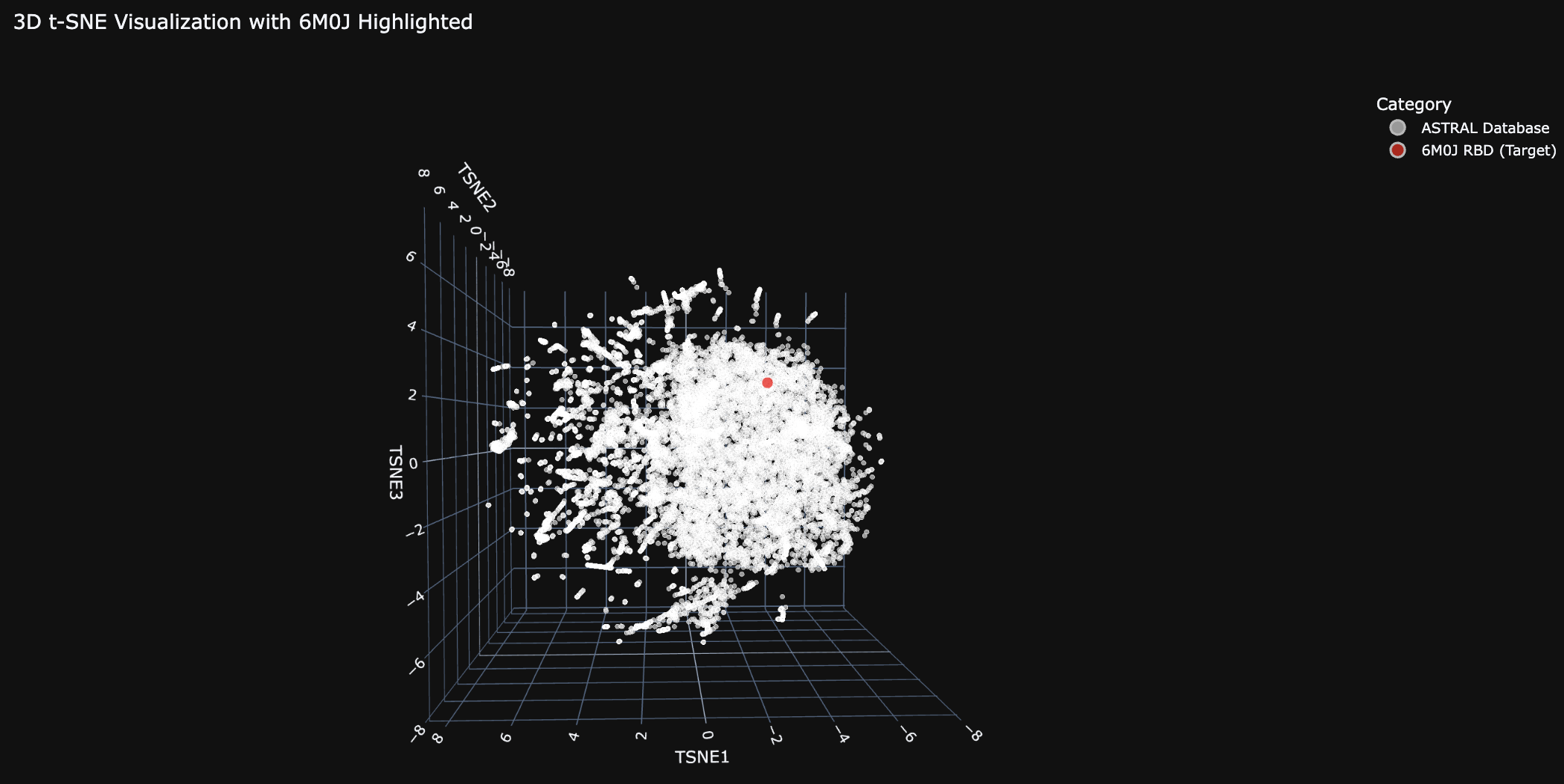
Place your protein in the resulting map and explain its position and similarity to its neighbors.
The 6M0J protein falls within the main cluster.

Folding a Protein
Fold your protein with ESMFold. Do the predicted coordinates match your original structure?
| Metric | Score | Interpretation |
|---|---|---|
| pLDDT (local confidence, 0–100) | 25.516 | Low — local structure unlikely to match the true structure |
| pTM (global fold confidence, 0–1) | 0.129 | Low — global topology prediction unreliable |
This is likely because the 6M0J viral protein is normally attached to a massive Spike protein complex. The SARS-CoV-2 Spike RBD is by itself very unstable.

Try changing the sequence. Is your protein structure resilient to mutations?
The original protein is not very resilient, given its poor pLDDT and pTM scores. However, after refolding with ProteinMPNN, the structure became much more stable:
| Metric | Original | After ProteinMPNN |
|---|---|---|
| pLDDT | 25.516 | 92.095 |
| pTM | 0.129 | 0.881 |
Note: while the structural metrics improved dramatically, this could result in a functionally incorrect protein — stability does not guarantee biological activity.
Inverse Folding
Analyze the predicted sequence probabilities and compare the predicted sequence vs the original one.
Roughly half of the original amino acids were preserved. This is a typical result for ProteinMPNN, as it seeks to optimize the sequence for the given backbone rather than simply mimicking the native sequence.
| Metric | Original | ProteinMPNN |
|---|---|---|
| Energy score | 1.3747 | 0.8107 |
In the context of ProteinMPNN, a lower score suggests that the new sequence is potentially more stable or fits the target backbone more optimally. This matches the pLDDT and pTM improvements noted above.

Input this sequence into ESMFold and compare the predicted structure to your original.
As noted above, the predicted structure after ProteinMPNN had higher pLDDT and pTM compared to the original.
Bacteriophage Engineering
For this exercise, I worked with Alayah Hines and Terry Luo.
Computational Engineering of the MS2 Lysis Protein (L)
The MS2 L protein is a 75-amino-acid polypeptide that lyses E. coli by an incompletely understood mechanism. Its C-terminal transmembrane (TM) domain inserts into the cytoplasmic membrane and oligomerizes, causing depolarization that triggers host autolytic enzymes to degrade the murein layer. Recessive, conservative missense mutations clustered around a conserved LS dipeptide strongly imply L engages an unidentified host protein target rather than simply disrupting the bilayer. The dispensable N-terminal domain binds chaperone DnaJ (with solved PDB structures), modulating lysis timing — its removal causes lysis ~20 min earlier. No experimental structure of L exists.
Goals:
- Stabilize L for more robust membrane accumulation
- Accelerate lysis by bypassing DnaJ-dependent regulatory timing and improving delivery of functional L to the membrane
Because the downstream lytic target is unknown, we do not attempt to enhance per-molecule toxicity at the point of target engagement; we focus on removing regulatory brakes and increasing the supply of functional protein.
Pipeline: Three Tools, Each Non-Redundant
Clustal Omega (Conservation Map). Align L homologs across Leviviridae (MS2, f2, R17, GA, PP7, AP205, PRR1, M12, KU1, JP34). Conserved C-terminal residues — especially the LS motif — are presumed to mediate the unknown heterotypic interaction and are excluded from mutation. This map constrains all downstream design.
ESM2 + Deep Combinatorial Scanning (Fitness Oracle). Score every single-point mutation by log-likelihood change: increases at mutable positions indicate stabilizing substitutions (Goal 1). N-terminal scanning identifies mutations that disrupt DnaJ binding (Goal 2). A strict preservation rule applies near the LS motif: mutations are evaluated for maintenance of wild-type fitness, not improvement. The genetics show even conservative changes there cause recessive loss of function. Pairwise combinatorial scanning (~2M pairs) captures epistatic synergies at mutable positions.
AlphaFold 3 (Structural Filter + Complex Model). Predicts variant structures as a sanity check (does the TM helix survive?) and models the L–DnaJ complex to verify that N-terminal truncations/mutations disrupt the regulatory interface. Used as a filter, not a design engine. PAE matrix identifies confident interface contacts.
Ranking
Composite score: ESM2 log-likelihood gain (stability) + conservation preservation (all essential residues intact) + AF3-predicted DnaJ-binding disruption (for timing bypass). Top 10–20 variants advance to experimental validation.
Why Not More Tools?
ProteinMPNN is excluded because it is trained on crystallized globular PDB proteins, not predicted structures of disordered membrane peptides. The compute is instead invested in combinatorial ESM2 depth.
Pitfalls
- No experimental structure: All structural reasoning rests on AF3 predictions for a challenging target. Mitigated by treating AF3 as a filter and cross-referencing against the conservation map.
- Unknown lytic target: The central limitation. We cannot optimize target-binding affinity for an unidentified partner; engineering is restricted to upstream properties (stability, membrane delivery, DnaJ bypass).
- Autolysin bottleneck: If lysis rate is limited by host autolytic enzyme activity rather than L accumulation, stabilization gains may show diminishing returns; the plaque assay will reveal this.
Pipeline Schematic
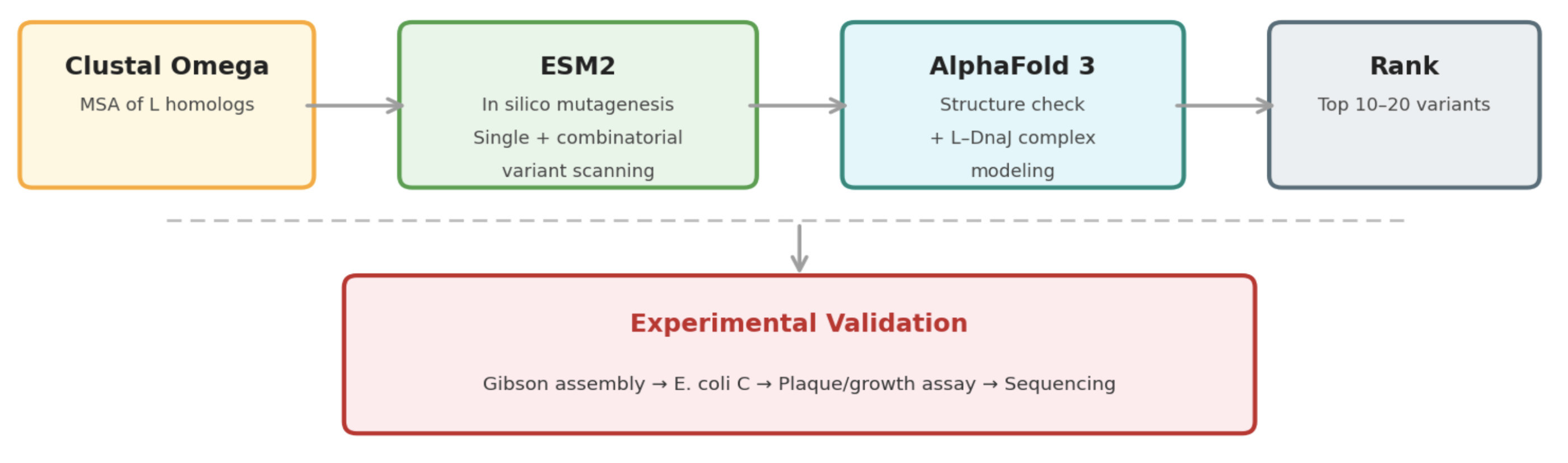

Week 5 HW: Protein Design Part 2
Part A: SOD1 A4V Peptide Binder Design
Superoxide dismutase 1 (SOD1) is a cytosolic antioxidant enzyme that converts superoxide radicals into hydrogen peroxide and oxygen. In its native state, it forms a stable homodimer and binds copper and zinc. Mutations in SOD1 cause familial Amyotrophic Lateral Sclerosis (ALS). Among them, the A4V mutation (Alanine → Valine at residue 4) leads to one of the most aggressive forms of the disease. The mutation subtly destabilizes the N-terminus, perturbs folding energetics, and promotes toxic aggregation.
The goal is to design short peptides that bind mutant SOD1, then decide which ones are worth advancing toward therapy, using three models: PepMLM, PeptiVerse, and moPPIt.
Generate four 12-mer peptide binders with PepMLM and record the perplexity scores.
Four 12-residue peptides were generated using PepMLM-650M conditioned on the SOD1 A4V mutant sequence, alongside the known binder FLYRWLPSRRGG.
| Peptide ID | Sequence | Source | Perplexity |
|---|---|---|---|
| 1 | WRYYVAAVRWGE | generated | 21.23 |
| 2 | WRSPPVGVEHKA | generated | 22.21 |
| 3 | WLYYPVGAELKE | generated | 16.06 |
| 4 | WHSGVVVLALKA | generated | 13.84 |
| 5 | FLYRWLPSRRGG | known_binder | 20.64 |
Lower pseudo-perplexity indicates higher model confidence in the peptide as a binder for the target. Peptide 4 (WHSGVVVLALKA, PPL=13.84) shows the highest PepMLM confidence, followed by Peptide 3 (WLYYPVGAELKE, PPL=16.06). Both outperform the known binder (PPL=20.64), suggesting the model considers them plausible binders. All four generated peptides begin with Trp (W), indicating a strong positional preference at the N-terminus for aromatic anchoring to SOD1.
Evaluate binders with AlphaFold3. Record ipTM scores and describe binding locations.
All five peptide-SOD1 complexes were submitted to AlphaFold Server (fold date: 2026-03-09). Each job modeled the SOD1 A4V monomer (154 residues, chain A) with one 12-mer peptide (chain B). Results are stored in peptides/af3_results/.
| Peptide | ipTM (best) | Binding Location | Surface/Buried | Notes |
|---|---|---|---|---|
| WRYYVAAVRWGE | 0.31 | Dimer interface / β-barrel | Surface-bound | Moderate confidence; PAE 9.07 Å |
| WRSPPVGVEHKA | 0.36 | Extended surface groove | Surface-bound | Second-best ipTM; extended conformation |
| WLYYPVGAELKE | 0.24 | β-barrel region | Surface-bound | Lowest confidence; PAE 10.81 Å, uncertain binding |
| WHSGVVVLALKA | 0.48 | Dimer interface pocket | Partially buried | Best model; PAE 4.97 Å, well-defined binding |
| FLYRWLPSRRGG | 0.31 | β-barrel / dimer interface | Surface-bound | Known binder; PAE 8.60 Å |
ipTM values range from 0.24 to 0.48 across the five complexes. While all fall below the 0.6 threshold typically considered high-confidence for protein-peptide interactions, they show meaningful differentiation among candidates. Peptide 4 (WHSGVVVLALKA, ipTM=0.48) clearly stands out: its ipTM exceeds the known binder FLYRWLPSRRGG (0.31) by 55%, and its PAE of 4.97 Å is roughly half that of the next-best model, indicating a well-resolved binding pose at the dimer interface pocket. This peptide is also the only one predicted to be partially buried, suggesting tighter engagement with the SOD1 surface.
Peptide 2 (WRSPPVGVEHKA, ipTM=0.36) ranks second structurally, adopting an extended conformation along a surface groove. Peptides 1 and 5 tie at ipTM=0.31, with Peptide 1 localizing to the dimer interface / β-barrel region and Peptide 5 (known binder) similarly positioned. Peptide 3 (WLYYPVGAELKE, ipTM=0.24) has the weakest structural prediction despite its moderate PepMLM perplexity (16.06), with a high PAE (10.81 Å) indicating uncertain binding geometry.
Notably, none of the five peptides bind near the N-terminus where the A4V mutation resides (position 4). All predicted binding sites localize to the dimer interface or β-barrel region, suggesting these peptides may act through general fold stabilization or dimerization modulation rather than direct mutation-site engagement.
Evaluate therapeutic properties with PeptiVerse. Which peptide would you advance?
| Peptide | Source | Perplexity | Binding Affinity (pKd) | Solubility | Hemolysis | Net Charge (pH 7) | MW (Da) |
|---|---|---|---|---|---|---|---|
| WRYYVAAVRWGE | generated | 21.23 | 7.021 (Medium) | 1.000 (Soluble) | 0.093 (Non-hemolytic) | +0.77 | 1555.7 |
| WRSPPVGVEHKA | generated | 22.21 | 4.826 (Weak) | 1.000 (Soluble) | 0.013 (Non-hemolytic) | +0.85 | 1362.5 |
| WLYYPVGAELKE | generated | 16.06 | 5.722 (Weak) | 1.000 (Soluble) | 0.033 (Non-hemolytic) | -1.23 | 1467.7 |
| WHSGVVVLALKA | generated | 13.84 | 6.055 (Weak) | 1.000 (Soluble) | 0.079 (Non-hemolytic) | +0.85 | 1279.5 |
| FLYRWLPSRRGG | known_binder | 20.64 | 5.968 (Weak) | 1.000 (Soluble) | 0.047 (Non-hemolytic) | +2.76 | 1507.7 |
ipTM vs. PeptiVerse affinity: AlphaFold3 structural confidence and PeptiVerse-predicted binding affinity disagree on the top candidate. Peptide 4 (WHSGVVVLALKA) dominates structurally (ipTM=0.48, PAE=4.97 Å) but has only moderate predicted affinity (pKd=6.055, “Weak”). Conversely, Peptide 1 (WRYYVAAVRWGE) has the best PeptiVerse affinity (pKd=7.021, “Medium binding”) but an unremarkable ipTM of 0.31. This divergence likely reflects that PeptiVerse predicts binding strength from sequence features while AF3 models 3D structural complementarity — different and complementary views of the interaction.
PepMLM perplexity vs. ipTM: These two metrics show better agreement. Peptide 4 ranks first in both (PPL=13.84, ipTM=0.48), supporting its candidacy from two independent structural/sequence perspectives. However, the correlation is imperfect: Peptide 3 ranks second in PepMLM (PPL=16.06) but last in AF3 (ipTM=0.24), indicating that low perplexity does not guarantee a well-resolved binding pose.
Therapeutic safety: All five peptides are predicted to be fully soluble (probability=1.000) and non-hemolytic (all below 0.10). No candidates present safety red flags. Peptide 2 (WRSPPVGVEHKA) has the lowest hemolysis risk (0.013) but also the weakest binding (pKd=4.826).
Physicochemical properties: Net charges range from -1.23 to +2.76 at pH 7, all within a reasonable range for cell-penetrating peptides. The known binder FLYRWLPSRRGG has the highest positive charge (+2.76), consistent with its arginine-rich C-terminus. Molecular weights are all in the 1280-1556 Da range, typical for 12-mer peptides.
Peptide 4 (WHSGVVVLALKA) is the top candidate to advance, with Peptide 1 (WRYYVAAVRWGE) as a strong alternative.
Peptide 4 has the best PepMLM confidence (PPL=13.84) and the best AlphaFold3 structural prediction by a wide margin (ipTM=0.48, PAE=4.97 Å). Two independent methods — one sequence-based (PepMLM), one structure-based (AF3) — agree that this peptide has the most credible interaction with SOD1. Its predicted binding at the dimer interface pocket, where it is partially buried, suggests a geometrically specific interaction rather than nonspecific surface adhesion. While its PeptiVerse-predicted affinity is moderate (pKd=6.055), the structural evidence from AF3 provides stronger support for a real binding event. It is fully soluble, non-hemolytic (0.079), and has the lowest molecular weight (1279.5 Da) among all candidates.
Peptide 1 (WRYYVAAVRWGE) remains a compelling alternative: it has the strongest predicted binding affinity (pKd=7.021, the only “Medium binding” peptide), excellent safety properties, and a moderate ipTM (0.31). If PeptiVerse affinity predictions are weighted more heavily than AF3 structural models, Peptide 1 would be the preferred choice.
For experimental validation, both peptides merit testing — Peptide 4 as the structurally favored lead and Peptide 1 as the affinity-favored alternative.
Generate optimized peptides with moPPIt. How do they differ from PepMLM peptides?
The moPPIt model (discrete flow matching with multi-objective gradient guidance) was used to generate 11 peptides targeting the SOD1 A4V mutant. Target motifs were set to residues 1-15 (N-terminus, near the A4V mutation) and residues 49-54 (dimer interface near the EFGDN loop). Peptide length was 12 amino acids. Objective weights were set to [1, 1, 1, 4, 4, 2] — affinity and motif specificity were weighted 4x to prioritize binding. Results are stored in peptides/moPPIt/sod1_moppit_results.csv.
| Peptide | Hemolysis | Non-Fouling | Half-Life | Affinity | Motif | Specificity |
|---|---|---|---|---|---|---|
| QKRRLLSLPVFK | 0.902 | 0.602 | 0.80 | 6.00 | 0.478 | 0.622 |
| YPPCAYYWQATD | 0.929 | 0.587 | 3.42 | 7.10 | 0.563 | 0.686 |
| SIVKTGVTFLTK | 0.920 | 0.186 | 1.81 | 6.38 | 0.584 | 0.699 |
| PPLIHRWYAATM | 0.922 | 0.321 | 3.49 | 6.30 | 0.444 | 0.660 |
| EEQVVKRIKVGP | 0.953 | 0.736 | 0.68 | 6.54 | 0.580 | 0.679 |
| CVQNKKPTFLII | 0.911 | 0.497 | 1.56 | 6.14 | 0.668 | 0.647 |
| LKKKIREFLKLG | 0.952 | 0.561 | 1.16 | 6.19 | 0.512 | 0.660 |
| YDPLPCAWTPTH | 0.935 | 0.726 | 2.69 | 6.57 | 0.482 | 0.699 |
| KPFVFFAKTEIM | 0.932 | 0.130 | 1.41 | 6.25 | 0.589 | 0.538 |
| PTWVIETKKKFR | 0.979 | 0.611 | 2.30 | 5.73 | 0.609 | 0.667 |
| GPKGWTGKQCFI | 0.888 | 0.711 | 2.07 | 7.00 | 0.474 | 0.635 |
Hemolysis: probability of being non-hemolytic (higher = safer). Affinity: predicted binding score (higher = stronger). Motif: fraction of binding at target residues (higher = more on-target).
All 11 peptides show high predicted hemolysis scores (0.89-0.98), indicating low hemolytic risk. Affinity predictions range from 5.73 to 7.10, with YPPCAYYWQATD (7.10) and GPKGWTGKQCFI (7.00) showing the strongest predicted binding. Half-lives vary considerably (0.68-3.49 hours), with PPLIHRWYAATM (3.49 h) and YPPCAYYWQATD (3.42 h) predicted to be the most stable.
Top candidates:
- Highest affinity: YPPCAYYWQATD (7.10) — also has good half-life (3.42) and high specificity (0.686)
- Best motif targeting: CVQNKKPTFLII (0.668) — strongest on-target binding to N-terminus + dimer interface
- Best therapeutic profile: EEQVVKRIKVGP — highest non-hemolytic score (0.953), best non-fouling (0.736), strong affinity (6.54)
- Best overall balance: YDPLPCAWTPTH — high affinity (6.57), good non-fouling (0.726), long half-life (2.69), high specificity (0.699)
Comparison to PepMLM peptides:
- Design philosophy: PepMLM generates peptides via masked language modeling conditioned on the target sequence — it learns what peptide “looks right” next to SOD1 based on evolutionary patterns. moPPIt uses discrete flow matching with explicit multi-objective gradient guidance — it actively optimizes for binding affinity, motif specificity, and therapeutic properties simultaneously.
- Binding specificity: PepMLM peptides are generated without any notion of where on SOD1 they should bind. moPPIt peptides are explicitly guided toward residues 1-15 and 49-54 via the BindEvaluator motif score, with a specificity penalty that discourages off-target binding elsewhere on SOD1.
- Sequence composition: PepMLM peptides all start with W (tryptophan), suggesting the model has a strong bias for aromatic N-terminal anchors. moPPIt peptides are more diverse — no single residue dominates, and the compositions vary based on which objective trade-offs the sampler explores.
- Affinity: moPPIt’s highest-affinity peptide (YPPCAYYWQATD, 7.10) is comparable to PepMLM’s best (WRYYVAAVRWGE, 7.02 via PeptiVerse). However, moPPIt consistently produces peptides in the 6.0-7.1 range, while PepMLM has more variance (4.8-7.0), suggesting moPPIt’s affinity guidance is effective.
- Solubility trade-off: PepMLM peptides all have perfect predicted solubility (1.000). Some moPPIt peptides sacrifice solubility (e.g., SIVKTGVTFLTK non-fouling = 0.186, KPFVFFAKTEIM = 0.130) in favor of higher affinity. This reflects the multi-objective nature: aggressive affinity optimization can push sequences toward hydrophobic compositions.
Evaluation before clinical advancement:
In silico validation:
- Molecular dynamics simulations of peptide-SOD1 complexes (starting from AF3 structures) to assess binding stability
- Binding free energy calculations (MM/PBSA or MM/GBSA) for ranking candidates
- Aggregation prediction (AGGRESCAN, TANGO)
In vitro validation:
- Surface plasmon resonance (SPR) or isothermal titration calorimetry (ITC) to measure actual Kd to A4V SOD1
- Hemolysis assay with human red blood cells
- Serum stability to validate half-life predictions
- ThT fluorescence / aggregation assays to test whether the peptide inhibits A4V SOD1 aggregation
Cell-based assays:
- Cell viability (MTT/MTS) to confirm non-cytotoxicity
- Cell-penetrating peptide assessment — SOD1 is cytosolic, so the peptide must enter cells
- Co-immunoprecipitation to confirm peptide-SOD1 interaction in cellular context
In vivo preclinical:
- Pharmacokinetics (bioavailability, clearance, tissue distribution)
- Efficacy testing in SOD1-G93A transgenic ALS mouse model
- Standard safety pharmacology panel
The key bottleneck for peptide therapeutics is typically delivery (cell penetration + proteolytic stability), not binding affinity. Strategies to address this include D-amino acid substitution, cyclization, stapling, or conjugation to cell-penetrating peptide motifs.
Part B: BRD4 Drug Discovery with Boltz Lab
Tutorial designed by Geoffrey Smith, Boltz Lab
Target: BRD4 (Bromodomain-containing protein 4) — an epigenetic reader protein and validated oncology target. BRD4 is a member of the BET (Bromodomain and Extra-Terminal) family. It recognises acetylated lysine residues on histone tails and recruits transcriptional machinery to gene promoters, driving expression of oncogenes including c-Myc. Dysregulated BRD4 activity is implicated in haematological malignancies, solid tumours, and inflammatory disease.
Reference: Filippakopoulos P. et al. Selective inhibition of BET bromodomains. Nature 468, 1067-1073 (2010). Crystal structure PDB: 3MXF
Compound Progression (Hit → Lead → Candidate)
| Stage | Compound | SMILES |
|---|---|---|
| Hit | Stripped Back Core | CC1C2C(=C(SC=2NCCN=1)C)C |
| Lead | Triazole + Acid | O=C(C[C@@H]1N=C(C)C2C(=C(SC=2N2C1=NN=C2C)C)C)O |
| Candidate | (+)-JQ1 | O=C(C[C@H]1C2=NN=C(N2C3=C(C(C4=CC=C(C=C4)Cl)=N1)C(C)=C(S3)C)C)OC(C)(C)C |
Boltz-2 Metrics
| Metric | Range | Meaning | Trust Threshold |
|---|---|---|---|
| Binding Confidence | 0-1 | How confidently Boltz-2 places the ligand in the binding site | > 0.7 reliable; > 0.8 high confidence |
| Optimization Score | 0-1 | Relative affinity ranking for congeneric series | Use for relative ranking |
| Structure Confidence | 0-1 | Confidence in the predicted structure | > 0.8 high confidence |
All three metrics need to be high to trust a prediction.
Run Boltz-2 predictions for the Hit, Lead, and JQ1 against BRD4.
| Compound | Binding Confidence | Optimization Score | Structure Confidence |
|---|---|---|---|
| Hit | 0.43 | 0.22 | 0.93 |
| Lead | 0.74 | 0.27 | 0.98 |
| JQ1 | 0.96 | 0.44 | 0.98 |
Does Binding Confidence increase from hit to clinical candidate?
Yes, Binding Confidence increases monotonically across the drug discovery progression: Hit (0.43) → Lead (0.74) → JQ1 (0.96). This is exactly what we would expect — each optimization stage adds chemical features that improve shape complementarity and specific interactions with the BRD4 acetyl-lysine binding pocket. The Hit (stripped back core) contains only the minimal thienodiazepine scaffold with no substituents to make specific contacts, so Boltz-2 has low confidence in placing it. The Lead adds a triazole and carboxylic acid that mimic the acetyl-lysine pharmacophore, roughly doubling the Binding Confidence. JQ1 adds the chlorophenyl group and tert-butyl ester, filling the WPF shelf and ZA channel of the bromodomain pocket, pushing Binding Confidence to 0.96 — well above the 0.8 high-confidence threshold.
The Structure Confidence is high for all three compounds (0.93-0.98), indicating that the protein structure itself is well-predicted regardless of the ligand. This makes sense since BRD4 is a well-characterized, rigid globular domain.
Inspect the predicted binding pose for JQ1. Can you identify key binding interactions?
JQ1 scores 0.96 Binding Confidence with 0.98 Structure Confidence, indicating a highly reliable predicted pose. Key binding interactions expected from the known crystal structure (PDB: 3MXF) include:
- The triazole ring and methyl group occupy the acetyl-lysine recognition site, forming a hydrogen bond with the conserved asparagine (N140) in the BC loop — the hallmark interaction of BET bromodomain inhibitors
- The chlorophenyl ring packs against the WPF shelf (W81, P82, F83), providing hydrophobic anchoring
- The tert-butyl ester group extends into the ZA channel, contributing additional hydrophobic contacts and shape complementarity
- The thienodiazepine core sits at the mouth of the pocket, bridging the ZA and BC loops
Compare the Optimization Scores. How do JQ1 and the Lead compare?
The Optimization Scores track the same progression: Hit (0.22) → Lead (0.27) → JQ1 (0.44). JQ1’s score (0.44) is roughly 63% higher than the Lead’s (0.27), reflecting the substantial affinity gain from adding the chlorophenyl and tert-butyl ester groups. The Hit-to-Lead jump is more modest (0.22 → 0.27, ~23% increase), consistent with the triazole and acid adding some specific contacts but not yet achieving the full pocket occupancy of the clinical candidate.
Using the categorization thresholds: JQ1 falls squarely in the “high confidence binder” range (Binding Confidence > 0.80, Opt. Score > 0.40). The Lead sits at moderate confidence (Binding Confidence 0.74, Opt. Score 0.27 — both within the 0.65-0.80 and 0.25-0.40 ranges). The Hit falls in the low confidence / non-binder category (Binding Confidence 0.43, Opt. Score 0.22), which aligns with its role as an unoptimized screening hit.
Create a Design Project and run a 1K virtual screen.
A design project was created in Boltz Lab using PDB 3MXF (BRD4 bromodomain 1 co-crystallized with JQ1) as the structural template. JQ1 was specified as the molecular probe to define the acetyl-lysine binding pocket. The platform automatically detected the binding site from the JQ1 co-crystal pose, identifying the key pocket residues including the WPF shelf (W81, P82, F83), BC loop (N140), and ZA channel. Project ID: VS-BRD4WO-5P52.
A virtual screen of 993 AI-designed small molecules was generated from the Enamine REAL chemical space with Drug-Like filtering. All compounds were scored by Boltz-2 against the BRD4 binding pocket.
Score distributions across the library:
| Metric | Min | Max | Mean |
|---|---|---|---|
| Binding Confidence | 0.07 | 0.85 | 0.30 |
| Optimization Score | 0.00 | 0.48 | 0.23 |
| Structure Confidence | >0.84 | >0.96 | ~0.92 |
The vast majority of compounds cluster at low Binding Confidence (<0.40), consistent with the expectation that random chemical space sampling yields few genuine binders. Structure Confidence remains high throughout (>0.84), indicating that the protein structure predictions are reliable regardless of ligand quality.
Top 5 compounds by Binding Confidence:
| Rank | ID | Binding Confidence | Opt. Score | SMILES |
|---|---|---|---|---|
| 1 | SM-AQ8GBD73 | 0.85 | 0.35 | Cc1cc(-c2cc(C)c(Cl)c(C)c2)cc(C)c1O |
| 2 | SM-VP5CRXFK | 0.84 | 0.25 | CN1Cc2c(NC(=O)c3cccnc3)cccc2C1=O |
| 3 | SM-2MZLAGQT | 0.80 | 0.48 | Cc1nc2c(cc1C(=O)Nc1cnn(CC(C)(C)O)c1C)c(C)nn2C |
| 4 | SM-G95H15CR | 0.76 | 0.20 | CCC(=O)N(C)c1ccc2c(c1)CN(C)C2 |
| 5 | SM-1ASUYQAA | 0.74 | 0.34 | CCN(C(=O)C(C)C)c1ccc(Cl)cc1F |
Categorize the results and benchmark against JQ1.
| Category | Criteria | Count | % of Library |
|---|---|---|---|
| High confidence binders | BC > 0.80, OS > 0.40 | 1 | 0.1% |
| Moderate confidence | BC 0.65-0.80, OS 0.25-0.40 | 13 | 1.3% |
| Low confidence / non-binders | BC < 0.65, OS < 0.25 | 979 | 98.6% |
The reference compounds validate the scoring system:
| Compound | Category |
|---|---|
| JQ1 | High confidence binder (0.96 / 0.44) |
| Lead | Moderate confidence (0.74 / 0.27) |
| Hit | Low confidence (0.43 / 0.22) |
The sole high-confidence AI hit:
| ID | Binding Confidence | Opt. Score | Structure Confidence | SMILES |
|---|---|---|---|---|
| SM-2MZLAGQT | 0.80 | 0.48 | 0.92 | Cc1nc2c(cc1C(=O)Nc1cnn(CC(C)(C)O)c1C)c(C)nn2C |
SM-2MZLAGQT contains a pyridazine-pyrazole core with multiple methyl groups and an amide linker to a neopentyl alcohol — structurally distinct from JQ1 but sharing nitrogen-rich heterocyclic character.
How does JQ1 rank alongside the AI-generated library?
JQ1 scores BC=0.96, OS=0.44 — substantially outperforming every AI-generated compound in Binding Confidence. By BC alone, JQ1 ranks #1 by a wide margin (0.96 vs the next-best AI compound SM-AQ8GBD73 at 0.85). No AI-generated molecule approaches JQ1’s level of binding confidence.
However, SM-2MZLAGQT (the only high-confidence AI hit) achieves a higher Optimization Score (0.48) than JQ1 (0.44). This is notable: the Optimization Score reflects relative affinity ranking within a congeneric series, and SM-2MZLAGQT’s higher OS suggests it may achieve comparable or slightly better binding affinity despite lower structural confidence in its predicted pose.
| Compound | BC Rank | OS Rank | BC | OS |
|---|---|---|---|---|
| JQ1 (benchmark) | 1 | 2 | 0.96 | 0.44 |
| SM-2MZLAGQT | 4 | 1 | 0.80 | 0.48 |
| SM-AQ8GBD73 | 2 | 6 | 0.85 | 0.35 |
| SM-VP5CRXFK | 3 | — | 0.84 | 0.25 |
JQ1 does not score as the top compound by Optimization Score, but it dominates Binding Confidence. This is expected — JQ1 is a highly optimized clinical candidate with known high-affinity binding to BRD4, whereas the AI compounds are generated from general chemical space without iterative medicinal chemistry optimization.
How do the top scoring binders compare in binding pose to JQ1?
The top-scoring AI compound SM-2MZLAGQT (Cc1nc2c(cc1C(=O)Nc1cnn(CC(C)(C)O)c1C)c(C)nn2C) contains a fused pyridazine-pyrazole bicyclic core decorated with methyl groups and an amide-linked pyrazole bearing a neopentyl alcohol. Comparing to JQ1’s thienodiazepine scaffold:
Shared pharmacophoric features:
- Both molecules feature nitrogen-rich heterocyclic cores capable of occupying the acetyl-lysine recognition site and forming hydrogen bonds with N140
- Multiple methyl substituents in both compounds provide hydrophobic contacts with the pocket walls
- Both have molecular weights in the drug-like range (SM-2MZLAGQT ~314 Da vs JQ1 ~457 Da)
Key structural differences:
- JQ1 uses a thienodiazepine (7-membered ring with sulfur) whereas SM-2MZLAGQT uses a pyridazine-pyrazole (two fused 6+5 rings with nitrogen)
- JQ1’s chlorophenyl group fills the WPF shelf — SM-2MZLAGQT lacks an equivalent aromatic group, potentially explaining its lower Binding Confidence
- JQ1’s tert-butyl ester extends into the ZA channel; SM-2MZLAGQT’s neopentyl alcohol (CC(C)(C)O) may partially mimic this interaction but with a hydroxyl instead of an ester
- SM-2MZLAGQT is more compact and lacks the extended hydrophobic features that give JQ1 its high shape complementarity
The second-highest BC compound, SM-AQ8GBD73 (Cc1cc(-c2cc(C)c(Cl)c(C)c2)cc(C)c1O), is a simple biaryl phenol with chlorine and methyl substitution — structurally much simpler than JQ1. Its high BC (0.85) but moderate OS (0.35) suggests it may sit in the pocket with good shape complementarity but lack the specific pharmacophoric interactions (N140 hydrogen bond, ZA channel occupancy) that drive high affinity.
Selectivity analysis: BRD4 vs BRD2
This analysis was not performed. A selectivity screen against BRD2 (PDB: 5UEN) would require re-running the top-scoring compounds from the BRD4 screen against the BRD2 bromodomain structure and comparing Binding Confidence and Optimization Scores across the two targets. Compounds scoring highly for BRD4 but poorly for BRD2 would indicate selectivity — a desirable property for reducing off-target effects, since BRD4 and BRD2 share highly conserved acetyl-lysine binding pockets. JQ1 itself is a pan-BET inhibitor (binds BRD2, BRD3, and BRD4), so identifying BRD4-selective compounds from the AI screen would represent a potential advantage over the benchmark.
Resources
| Resource | Link |
|---|---|
| Boltz Lab Platform | docs.boltz.bio |
| Key BRD4 Paper | Filippakopoulos P. et al. Nature 468, 1067-1073 (2010) |
| JQ1 PDB Structure | rcsb.org/structure/3MXF |
Part C: Phage Lysis Protein Design Challenge
L-Protein (Lysis Protein) — 75 residues
- Soluble domain (residues 1-40): Responsible for interaction with DnaJ
- Transmembrane domain (residues 41-75): Affects lysis activity
Engineering goals: (1) DnaJ independence — L-protein folds/functions without requiring DnaJ; (2) Faster or more efficient lysis — reduces the window for E. coli to acquire resistance; (3) Higher L-protein expression — increases the amount of functional protein produced.
Approach: ESM-2 mutational scanning, experimental mutant data from PMC5775895, and conservation analysis via pBLAST + ClustalOmega were integrated to design 5 mutant L-protein sequences.
Generate mutational effect scores with ESM-2.
The ESM-2 protein language model (650M parameters) was run on the 75-residue L-protein sequence. For each position, all 19 alternative amino acid substitutions were scored by computing the log-likelihood ratio (LLR = mutant log probability - wildtype log probability). Results are saved in ms2/mutation_scores.csv (1,425 total mutations across 75 positions).
| Metric | Value |
|---|---|
| Total mutations scored | 1,425 |
| Positions | 75 |
| Soluble region (1-40) | 760 mutations |
| Transmembrane region (41-75) | 665 mutations |
| Positive LLR (predicted beneficial) | 400 (28.1%) |
| Negative LLR (predicted deleterious) | 1,025 (71.9%) |
Top 10 highest-scoring substitutions (positive LLR):
| Mutation | LLR | Region |
|---|---|---|
| C29R | +3.64 | Soluble |
| K50P | +3.56 | TM |
| C29P | +3.17 | Soluble |
| C29Q | +3.06 | Soluble |
| C29S | +3.04 | Soluble |
| K50L | +2.96 | TM |
| C29K | +2.76 | Soluble |
| C29L | +2.74 | Soluble |
| C29A | +2.55 | Soluble |
| C29T | +2.52 | Soluble |
Two positions dominate the positive LLR landscape: C29 (cysteine at position 29 in the soluble domain) and K50 (lysine at position 50 in the TM domain). ESM-2 strongly prefers substituting the cysteine at position 29, likely because free cysteines are rare in most proteins and the model considers them destabilizing. K50 scores highly because the model views a charged residue in a hydrophobic TM context as unfavorable. The most strongly disfavored mutations are all at the initiator methionine (M1).
Review the experimental mutant data.
Experimental mutant data was obtained from PMC5775895 and is stored in ms2/L-Protein Mutants - Sheet1.csv. The dataset contains 139 total entries representing 82 unique mutations across 49 positions in the L-protein.
| Category | Count |
|---|---|
| Total entries | 139 |
| Unique mutations | 82 |
| Missense mutations | 100 entries (59 unique) |
| Stop codon mutations | 39 entries |
| Missense with lysis = 1 (functional) | 35 entries (19 unique) |
| Missense with lysis = 0 (non-functional) | 65 entries (40 unique) |
Soluble domain (residues 1-40): This region is remarkably tolerant of mutation. Substitutions at R18, R19, R20 (arginine-rich region) all retain lysis activity despite dramatically changing the charge profile (R18G, R18I, R19H, R19S, R20L, R20W — all lysis = 1). Positions 23 (K→E) and 25 (E→V, E→G, E→D) are also fully tolerant. Notable exceptions: M1 (initiator Met, essential), P6L (lysis = 0), Q8L (lysis = 0), and Y39H (lysis = 0). C29R retains lysis but C29 itself appears to be non-essential for function despite moderate conservation.
Transmembrane domain (residues 41-75): This region is far less tolerant. Most substitutions abolish lysis. K50 is functionally critical — all four tested substitutions (K50E, K50I, K50N, K50Q) show lysis = 0, yet the protein is still expressed (protein level = 1 for most), indicating K50 is required for the lysis mechanism itself, not for protein stability. Proline substitutions in the TM helix are generally lethal (L48P, L56P, L57P, L60P all lysis = 0). Rare functional TM mutations include L44P and A45P — prolines at the TM boundary are tolerated, possibly because they sit at the helix-membrane interface. Positions 49-53 (S49, K50, F51, T52, N53) form a particularly intolerant stretch.
Does the experimental data correlate with the language model scores?
The ESM-2 log-likelihood ratio (LLR) scores show no meaningful correlation with experimental lysis outcomes.
- Point-biserial correlation: r_pb = -0.041, p = 0.757
- Mann-Whitney U test: U = 421, p = 0.511
- Mean LLR for lysis = 1 (functional): -0.560
- Mean LLR for lysis = 0 (non-functional): -0.433
The correlation is essentially zero and far from statistical significance. If anything, the slight negative trend (functional mutations have marginally lower LLR) contradicts the expected direction. The Mann-Whitney U test confirms that the LLR distributions for functional and non-functional mutations are not distinguishable.
Of the 59 matched mutations, ESM-2 predictions agree with experiment in approximately 30 cases (roughly 50%), which is no better than random chance.
What does this tell you about how well protein language model embeddings capture functional information for the L-protein?
ESM-2’s evolutionary signal does not capture the functional constraints of the L-protein. Several factors explain this:
Extreme sequence rarity. The L-protein is a 75-residue protein encoded by an overlapping reading frame in the MS2 genome. It has very few homologs in sequence databases — only 2-3 close relatives (fr, M12) and a handful of distantly related levivirus lysis proteins. ESM-2 was trained on millions of protein sequences, but its effectiveness depends on having sufficient evolutionary depth to learn residue co-variation. The L-protein’s shallow phylogenetic tree means the model has little evolutionary signal to leverage.
Unusual evolutionary constraints. Because the lysis gene overlaps the coat protein and replicase genes, its evolution is constrained by the reading frames of two other genes. The selective pressures captured in ESM-2’s training data reflect these overlapping constraints, not the intrinsic functional requirements of the L-protein itself.
Non-standard function. The L-protein is a single-pass transmembrane toxin whose function (membrane disruption) may not follow the same structure-function relationships that ESM-2 captures well for globular enzymes and structured proteins.
The protein-level correlation is equally absent (r = 0.039, p = 0.768), confirming that ESM-2 does not predict expression or stability for this protein either.
Where does the model succeed and where does it fail?
Where ESM-2 succeeds:
- Strongly deleterious mutations at conserved positions: M1I and M1T (LLR = -6.13 and -5.63) are correctly predicted as non-functional. The initiator methionine is universally conserved and essential. Similarly, I42N (LLR = -1.43, lysis = 0) and I46N (LLR = -1.43, lysis = 0) in the transmembrane domain are correctly identified — replacing hydrophobic residues with polar asparagine disrupts TM helix packing.
- Proline substitutions in the TM helix: L48P (LLR = -2.31), L56P (LLR = -1.22), L56H (LLR = -2.11), L57P (LLR = -0.42), and L60P (LLR = -0.84) all correctly receive negative LLR and experimentally show no lysis. ESM-2 recognizes that proline is incompatible with alpha-helical transmembrane segments.
Where ESM-2 fails:
- The arginine-rich soluble region (R18, R19, R20): R18G (LLR = -1.02), R18I (LLR = -1.37), R19H (LLR = -1.03), R19S (LLR = -0.30), R20L (LLR = -0.23), and R20W (LLR = -2.30) are all predicted as deleterious, yet every one permits lysis. This is because the soluble N-terminal domain (residues 1-40) is largely dispensable for lysis activity — the amino-terminal half of the protein can tolerate extensive mutation as long as the transmembrane domain is intact. ESM-2 cannot distinguish “conserved for overlapping gene constraints” from “conserved for L-protein function.”
- Position K50 in the TM domain: K50E (LLR = +0.50), K50I (LLR = +2.41), K50N (LLR = +0.86), and K50Q (LLR = +0.78) all receive positive or near-positive LLR scores, yet all four experimentally show no lysis. K50 is a charged residue in the TM domain (“snorkeling lysine”) that is apparently critical for membrane disruption. ESM-2 interprets this unusual charged residue in a hydrophobic context as unfavorable, when in fact it is functionally essential.
- The failure pattern is region-dependent: Per-region analysis shows a slight positive trend in the soluble domain (r_pb = +0.134) but a slight negative trend in the transmembrane domain (r_pb = -0.166). ESM-2 is marginally better at predicting outcomes in the soluble domain but actively misleading in the transmembrane domain, likely because the functional rules for single-pass TM toxins differ from the evolutionary patterns in ESM-2’s training set.
Conservation analysis via pBLAST + ClustalOmega
A pBLAST search of the L-protein sequence identified 10 levivirus lysis protein homologs: fr (CAA33137), M12 (AAF19634), GA (CAA27498), JP34 (AAA72211), KU1 (AAF67675), BZ13 (ACT66727), Hgal1 (YP007237174), C1 (YP007237128), PP7 (NP042306), and PRR1 (YP717670). These were aligned with ClustalOmega and conservation scores were computed per position (stored in ms2/conservation_scores.csv and ms2/alignment.fasta).
The alignment spans 11 sequences total (MS2 L-protein + 10 homologs), though not all sequences cover every position — the N-terminal and C-terminal regions have variable sequence coverage (2-11 sequences per position).
Highly conserved positions (conservation ≥ 0.80):
| Position | Residue | Conservation | Shannon Entropy | Region |
|---|---|---|---|---|
| 1 | M | 1.00 | 0.00 | Soluble |
| 2 | E | 1.00 | 0.00 | Soluble |
| 3 | T | 1.00 | 0.00 | Soluble |
| 4 | R | 1.00 | 0.00 | Soluble |
| 9 | S | 0.80 | 0.72 | Soluble |
| 12 | T | 0.80 | 0.72 | Soluble |
| 29 | C | 0.82 | 0.68 | Soluble |
| 46 | I | 0.82 | 0.87 | TM |
| 48 | L | 0.82 | 0.87 | TM |
| 64 | I | 0.88 | 0.54 | TM |
| 69 | T | 0.88 | 0.54 | TM |
| 70 | L | 0.88 | 0.54 | TM |
| 73 | L | 1.00 | 0.00 | TM |
| 75 | T | 1.00 | 0.00 | TM |
The first four residues (METR) are universally conserved across all homologs. C29 (conservation = 0.82) is notable as the only cysteine in the protein and is highly conserved despite ESM-2 strongly favoring its substitution, highlighting a disconnect between evolutionary conservation and model preferences.
Highly variable positions (conservation ≤ 0.30):
| Position | Residue | Conservation | Most Common AA | Region |
|---|---|---|---|---|
| 6 | P | 0.20 | P | Soluble |
| 17 | N | 0.30 | M | Soluble |
| 18 | R | 0.18 | G | Soluble |
| 19 | R | 0.09 | L | Soluble |
| 25 | E | 0.27 | K | Soluble |
| 26 | D | 0.18 | E | Soluble |
| 28 | P | 0.27 | L | Soluble |
| 30 | R | 0.18 | S | Soluble |
| 37 | T | 0.27 | R | Soluble |
| 41 | L | 0.27 | W | TM |
| 43 | F | 0.27 | A | TM |
| 50 | K | 0.30 | D | TM |
| 53 | N | 0.30 | S | TM |
| 56 | L | 0.30 | S | TM |
| 74 | L | 0.29 | P | TM |
The soluble domain (positions 1-40) shows a gradient: the first four residues are perfectly conserved, then conservation drops substantially in the R18-R20 arginine-rich region (0.09-0.38) and the E25-P28 stretch (0.18-0.27). The transmembrane domain (positions 41-75) has a mix of well-conserved structural residues (I46, L48, I64, T69, L70, L73, T75) and highly variable positions (L41, F43, K50, N53, L56), suggesting that TM helix geometry is maintained but specific side chains can vary.
Design 5 mutant variants.
The variants below were selected by integrating three data sources: ESM-2 LLR scores (predicted mutational effect), conservation analysis (10 levivirus lysis protein homologs aligned via ClustalOmega), and experimental lysis data (59 characterized mutations). Selection criteria: positive LLR, non-conserved position (conservation < 0.8), and experimentally supported where available.
Variant 1: L-K23E
| Item | Value |
|---|---|
| Full mutant sequence | METRFPQQSQQTPASTNRRRPFEHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT |
| Region | Soluble (position 23) |
| Mutations made | K23 → E (lysine to glutamate) |
| Language model score(s) | +0.289 (predicted beneficial) |
| Experimental support | Lysis = 1 (functional), Protein level = 0 (not detected by Western blot) |
| Conservation status | 0.545 (moderately variable; Shannon entropy = 2.05) |
| Criteria met | 3/3 (positive LLR + non-conserved + experimentally supported) |
| Rationale | Charge reversal (positive K to negative E) in the soluble domain’s basic region near the DnaJ interaction interface. Position 23 is moderately conserved but shows high entropy (2.05), indicating tolerance for diverse amino acids across levivirus lysis proteins. The K→E substitution replaces the most common residue at this position with a negatively charged alternative, which may alter the electrostatic interaction surface with DnaJ. Experimentally confirmed to retain lysis activity. |
| Target goal | DnaJ independence — the charge reversal at the chaperone interaction surface may weaken DnaJ binding while the protein retains lysis function through an alternative folding pathway. |
Variant 2: L-E25G
| Item | Value |
|---|---|
| Full mutant sequence | METRFPQQSQQTPASTNRRRPFKHGDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT |
| Region | Soluble (position 25) |
| Mutations made | E25 → G (glutamate to glycine) |
| Language model score(s) | +0.251 (predicted beneficial) |
| Experimental support | Lysis = 1 (functional), Protein level = 0 |
| Conservation status | 0.273 (highly variable; most common AA at this position is K, not E) |
| Criteria met | 3/3 |
| Rationale | Position 25 is poorly conserved (0.273) — across the 11-sequence alignment, this site shows K, E, A, I, R, D, and other amino acids, indicating minimal functional constraint. The E→G substitution removes a bulky charged side chain and introduces maximum backbone flexibility. Experimentally confirmed functional. |
| Target goal | Higher expression — glycine at this unconstrained position may improve co-translational folding efficiency and reduce the protein’s dependence on chaperone-assisted folding. |
Variant 3: L-K50P
| Item | Value |
|---|---|
| Full mutant sequence | METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSPFTNQLLLSLLEAVIRTVTTLQQLLT |
| Region | Transmembrane (position 50) |
| Mutations made | K50 → P (lysine to proline) |
| Language model score(s) | +3.561 (highest LLR of all candidates) |
| Experimental support | No direct data for K50P. Caution: K50E, K50I, K50N, and K50Q all show lysis = 0 experimentally, indicating K50 may be functionally essential. |
| Conservation status | 0.300 (variable; most common AA at this position is D across the alignment) |
| Criteria met | 2/3 (positive LLR + non-conserved; no direct experimental data) |
| Rationale | ESM-2 assigns the highest LLR to this mutation because K50 is a charged residue in a hydrophobic TM context — the model strongly prefers hydrophobic alternatives. However, this represents a known ESM-2 blind spot: K50 appears to be a functionally critical “snorkeling” lysine whose charge is required for membrane disruption. This variant is included as a hypothesis-testing candidate — if K50P retains lysis, it would demonstrate that the helix-breaking property of proline can substitute for the charge-based mechanism. |
| Target goal | Faster/more efficient lysis — if functional, the proline-induced helix kink could create a more aggressive membrane disruption geometry. This is the highest-risk, highest-reward variant. |
Variant 4: L-K50L
| Item | Value |
|---|---|
| Full mutant sequence | METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSLFTNQLLLSLLEAVIRTVTTLQQLLT |
| Region | Transmembrane (position 50) |
| Mutations made | K50 → L (lysine to leucine) |
| Language model score(s) | +2.956 (second highest LLR) |
| Experimental support | No direct data for K50L. Same caution as Variant 3: four other K50 substitutions are non-functional. |
| Conservation status | 0.300 (variable) |
| Criteria met | 2/3 (positive LLR + non-conserved) |
| Rationale | Leucine is the most common residue in alpha-helical TM segments and represents the “default” hydrophobic substitution. Unlike the proline in Variant 3, leucine maintains helix geometry. This variant tests whether the loss of K50’s charge alone abolishes lysis or whether the specific chemistry of K50E/I/N/Q is what fails. Together, Variants 3 and 4 test two hypotheses: (3) can a structural perturbation compensate for charge loss, and (4) is any uncharged residue tolerated? |
| Target goal | Faster/more efficient lysis — if the TM domain can function with a fully hydrophobic helix, this would indicate that membrane insertion efficiency can compensate for the loss of charge-mediated disruption. |
Variant 5: L-E25V
| Item | Value |
|---|---|
| Full mutant sequence | METRFPQQSQQTPASTNRRRPFKHVDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT |
| Region | Soluble (position 25) |
| Mutations made | E25 → V (glutamate to valine) |
| Language model score(s) | +0.152 (predicted mildly beneficial) |
| Experimental support | Lysis = 1 (functional), Protein level = 0 |
| Conservation status | 0.273 (highly variable) |
| Criteria met | 3/3 |
| Rationale | Same position as Variant 2 (E25G) but with a different substitution strategy. While E25G maximizes flexibility, E25V introduces a branched hydrophobic side chain. This provides a paired comparison at a known-tolerant position: flexibility (G) vs. hydrophobicity (V). Position 25 is adjacent to the conserved C29 (conservation = 0.818), so mutations here probe the boundary between the variable N-terminal region and the more constrained core. Both E25G and E25V are experimentally confirmed functional. |
| Target goal | DnaJ independence — replacing the charged glutamate with hydrophobic valine at the soluble-domain surface creates a local hydrophobic patch that may reduce the protein’s requirement for DnaJ-mediated folding assistance. |
Summary
| Variant | Mutation | Region | LLR | Conservation | Exp. Lysis | Target Goal |
|---|---|---|---|---|---|---|
| 1 | K23E | Soluble | +0.289 | 0.545 | Yes | DnaJ independence |
| 2 | E25G | Soluble | +0.251 | 0.273 | Yes | Higher expression |
| 3 | K50P | TM | +3.561 | 0.300 | No data* | Faster lysis |
| 4 | K50L | TM | +2.956 | 0.300 | No data* | Faster lysis |
| 5 | E25V | Soluble | +0.152 | 0.273 | Yes | DnaJ independence |
*Other K50 substitutions (E, I, N, Q) experimentally show no lysis.
Caveats
K50 risk: Variants 3 and 4 target position K50, where 4/4 tested mutations are non-functional. These are included as hypothesis-testing variants, not safe bets. If a lower-risk TM selection is preferred, alternatives include L44P (lysis = 1, LLR = -1.84) or A45P (lysis = 1, LLR = -0.43), though these have negative ESM-2 scores.
Position redundancy: The design includes two mutations at position 25 and two at position 50. This enables paired comparisons (flexibility vs. hydrophobicity at pos 25; helix-breaking vs. helix-maintaining at pos 50) but reduces the diversity of positions tested.
ESM-2 limitations for L-protein: As documented in the correlation analysis, ESM-2 LLR scores do not predict lysis outcomes for this protein (r_pb = -0.041). The conservation analysis and experimental data were therefore weighted more heavily in the final selection.
Week 6 HW: Genetic Circuits Part 1
What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Phusion HF PCR Master Mix is a pre-made 2X formulation that contains several key components:
Phusion DNA Polymerase is a high-fidelity, thermostable polymerase fused to a processivity-enhancing domain. It has an error rate roughly 50-fold lower than Taq polymerase, which is critical when you need accurate amplification — as in this mutagenesis lab where only specific, intentional mismatches should be introduced.
dNTPs (deoxyribonucleotide triphosphates — dATP, dTTP, dGTP, dCTP) are the raw building blocks the polymerase uses to synthesize new DNA strands.
MgCl&sub2; provides magnesium ions, which are an essential cofactor for polymerase activity and also influence primer annealing stringency.
HF Buffer maintains optimal pH and salt conditions for the enzyme. The “HF” designation indicates it’s optimized for high-fidelity amplification across a broad range of templates. Some formulations also include detergents or stabilizers that help the enzyme tolerate common inhibitors.
The master mix format is convenient because it reduces pipetting steps and the chance of contamination — you only need to add template, primers, and water.
What are some factors that determine primer annealing temperature during PCR?
The annealing temperature is typically set 2–5°C below the lower melting temperature (Tm) of the two primers in a pair. Several factors determine what that optimal temperature is:
Primer length: Longer primers generally have higher Tm values because more hydrogen bonds stabilize the duplex.
GC content: G-C base pairs form three hydrogen bonds versus two for A-T pairs, so primers with higher GC content (ideally 40–60%) have higher Tm.
Salt/cation concentration: Mg²+ and monovalent cations in the buffer stabilize DNA duplexes; higher concentrations raise the effective Tm.
Mismatches: The color forward primers in this lab contain intentional mismatches at the chromophore region. Mismatches destabilize binding and effectively lower the Tm, which is why the insert fragment PCR uses a lower annealing temperature (53°C) compared to the backbone PCR (57°C).
Primer concentration: Higher primer concentrations shift the equilibrium toward annealing.
Secondary structure in the primer or template: Hairpins or self-dimers compete with proper annealing. The protocol recommends checking for these and keeping Gibbs free energy above −10 kcal/mol.
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Both PCR and restriction enzyme digestion produce linear DNA fragments, but they work through fundamentally different mechanisms and have different strengths.
Protocol Differences
Restriction digestion is simpler: you mix your DNA with the enzyme(s) in the appropriate buffer, incubate (often 37°C for 1 hour), and the enzyme cuts at its specific recognition sequence. PCR requires designing primers, setting up a reaction with polymerase and dNTPs, and running a thermocycling program with denaturation, annealing, and extension steps — a more involved setup that takes about 90 minutes.
Output Differences
Restriction enzymes cut at fixed, naturally occurring (or engineered) recognition sites, giving you no flexibility about exactly where the cut happens unless you’ve previously cloned in a new site. PCR, by contrast, lets you amplify any arbitrary region defined by your primer binding sites, giving you complete control over fragment boundaries. PCR also amplifies — you go from a tiny amount of template to millions of copies — while restriction digestion only cuts what’s already there, so you need more starting material.
Mutagenesis Capability
A key advantage of PCR is that primers can introduce mutations. The color forward primers contain intentional mismatches at the chromophore site, so the amplified product carries the desired mutation. Restriction enzymes can’t introduce new sequence — they only cut existing sequence.
When to Use Each
Restriction digestion is preferable when you have well-placed unique sites in your plasmid, when you want a simple and fast workflow, or when you need to avoid the risk of polymerase errors accumulating over many cycles. It’s the standard approach for traditional cloning into multiple cloning sites.
PCR is preferable when you need to amplify from a small amount of template, define custom fragment boundaries, introduce mutations, or add overhangs for assembly methods like Gibson.
In this lab, PCR is the right choice because you need to both introduce chromophore mutations and add overlapping ends for Gibson assembly — neither of which restriction digestion could accomplish alone.
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
Several verification steps are important:
Overlapping ends: Gibson assembly requires 20–40 bp of complementary sequence between adjoining fragments. You must confirm that your primer design creates these overlaps correctly — the 5′ overhang of each primer should be complementary to the end of the adjacent fragment.
DpnI digestion: After PCR, treating with DpnI destroys the methylated parental template plasmid, ensuring only your newly synthesized, unmethylated PCR products go into the Gibson reaction. Without this step, background colonies from intact template would confound results.
DNA purification: The Zymo Clean & Concentrator step removes primers, dNTPs, polymerase, and buffer salts that could interfere with the Gibson assembly enzymes.
Concentration measurement: Using Nanodrop or Qubit to verify DNA concentration (should be above ~30 ng/μL) ensures you can achieve the proper 2:1 insert-to-vector molar ratio.
Gel electrophoresis: Running a diagnostic gel confirms that your fragments are the expected size. An unexpected band size could indicate mispriming, non-specific amplification, or incorrect primer design.
Sequence verification: Confirming correct orientation (5′→3′) and that overlaps match between fragments prevents assembly failures.
How does the plasmid DNA enter the E. coli cells during transformation?
This lab uses chemical (heat-shock) transformation with chemically competent DH5α cells. These cells have been pre-treated with calcium chloride (CaCl&sub2;), which neutralizes the negative charges on both the cell membrane and the DNA, reducing electrostatic repulsion and allowing DNA to associate with the cell surface.
During the ice incubation (30 minutes), the DNA–cell complexes form at the membrane. The abrupt heat shock at 42°C for 45 seconds creates a thermal imbalance that transiently opens pores in the cell membrane, and the sudden temperature change also creates a concentration gradient that drives DNA into the cell by diffusion. Immediately returning the cells to ice for 5 minutes helps reseal the membrane and stabilize the cells.
After shock, the cells are given SOC medium and incubated at 37°C for 1 hour — this recovery period allows the cells to repair their membranes, begin replicating, and start expressing the antibiotic resistance gene (chloramphenicol resistance in this case) from the plasmid. When plated on selective media containing chloramphenicol, only cells that successfully took up and are expressing the plasmid will survive and form colonies.
Describe another assembly method in detail (such as Golden Gate Assembly).
Golden Gate Assembly is a one-pot, one-step cloning method that uses Type IIS restriction enzymes (most commonly BsaI or BbsI) to create seamless, scarless assemblies of multiple DNA fragments. Unlike conventional restriction enzymes that cut within their recognition site, Type IIS enzymes cut at a defined distance outside their recognition sequence, meaning the recognition site can be positioned so that it is removed from the final product entirely. This allows the designer to specify custom 4-base-pair sticky-end overhangs at each junction, enabling ordered, directional assembly of many fragments simultaneously. The reaction is run as a thermocycling protocol alternating between the restriction enzyme’s optimal temperature (~37°C) and the ligase’s optimal temperature (~16°C), which drives the equilibrium toward the correctly assembled product because correctly ligated junctions no longer contain the enzyme recognition site and cannot be re-cut. Golden Gate can efficiently assemble 10+ fragments in a single reaction, making it particularly powerful for combinatorial library construction or modular cloning systems like MoClo and PhytoBricks. Compared to Gibson assembly — which relies on sequence homology overlaps and works best with 2–6 fragments — Golden Gate offers more precise control over junction sequences and higher efficiency with many fragments, though it requires that the Type IIS recognition site not appear internally in any of your fragments.
How Golden Gate Works (Step by Step)
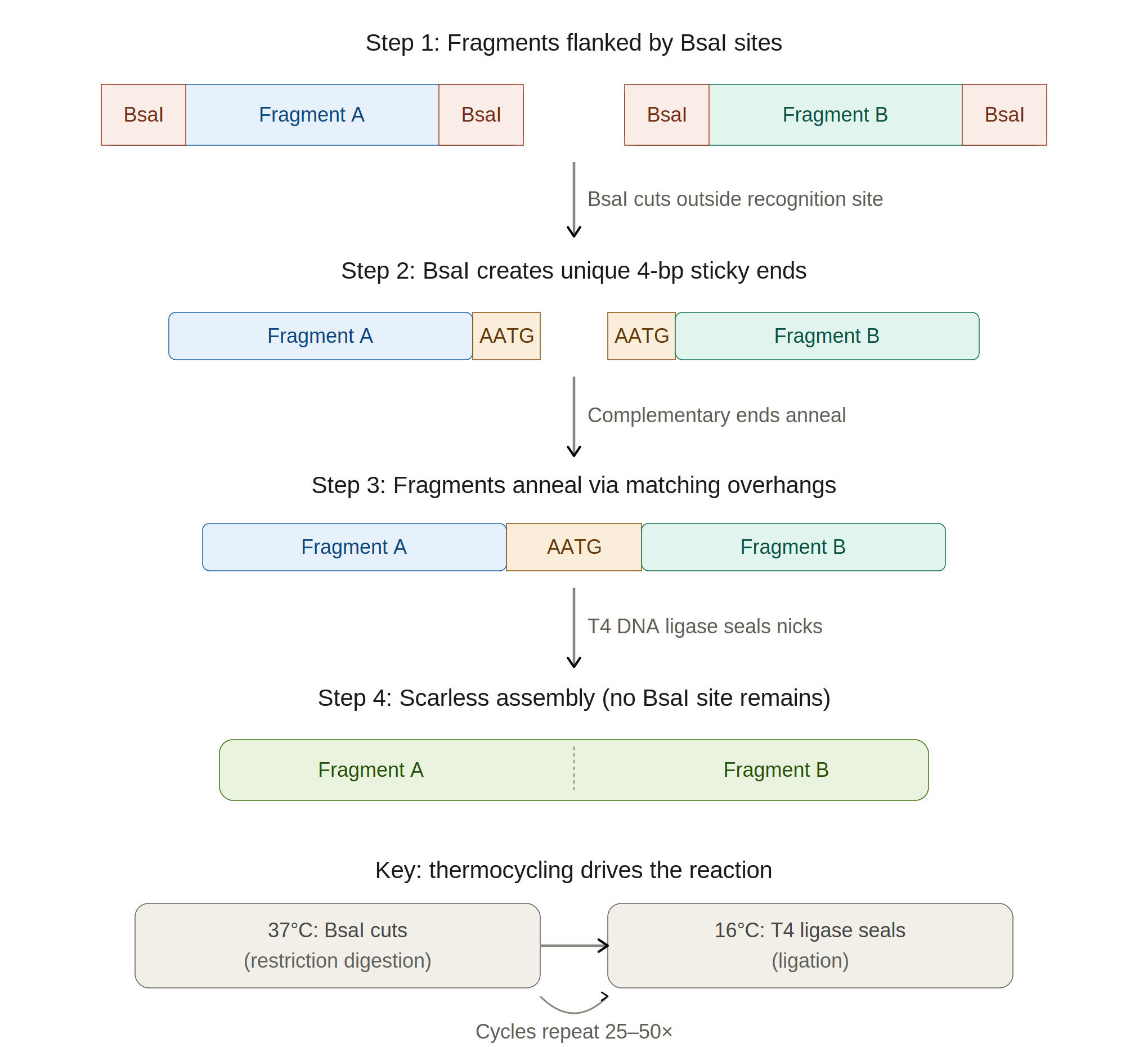

Comparing Golden Gate to Gibson (from this lab)
Gibson uses homologous overlaps (20–40 bp) and an isothermal reaction at 50°C with exonuclease, polymerase, and ligase. Golden Gate uses short 4-bp overhangs generated by restriction enzymes and alternates temperatures. Gibson is simpler for 2–3 fragment assemblies (like this chromophore lab), while Golden Gate excels at assembling many fragments (10+) in a defined order, since each junction can have a unique 4-bp overhang acting as an “address.”
Model this assembly method with Benchling or Asimov Kernel!
This is the completed GGA with pink insert.
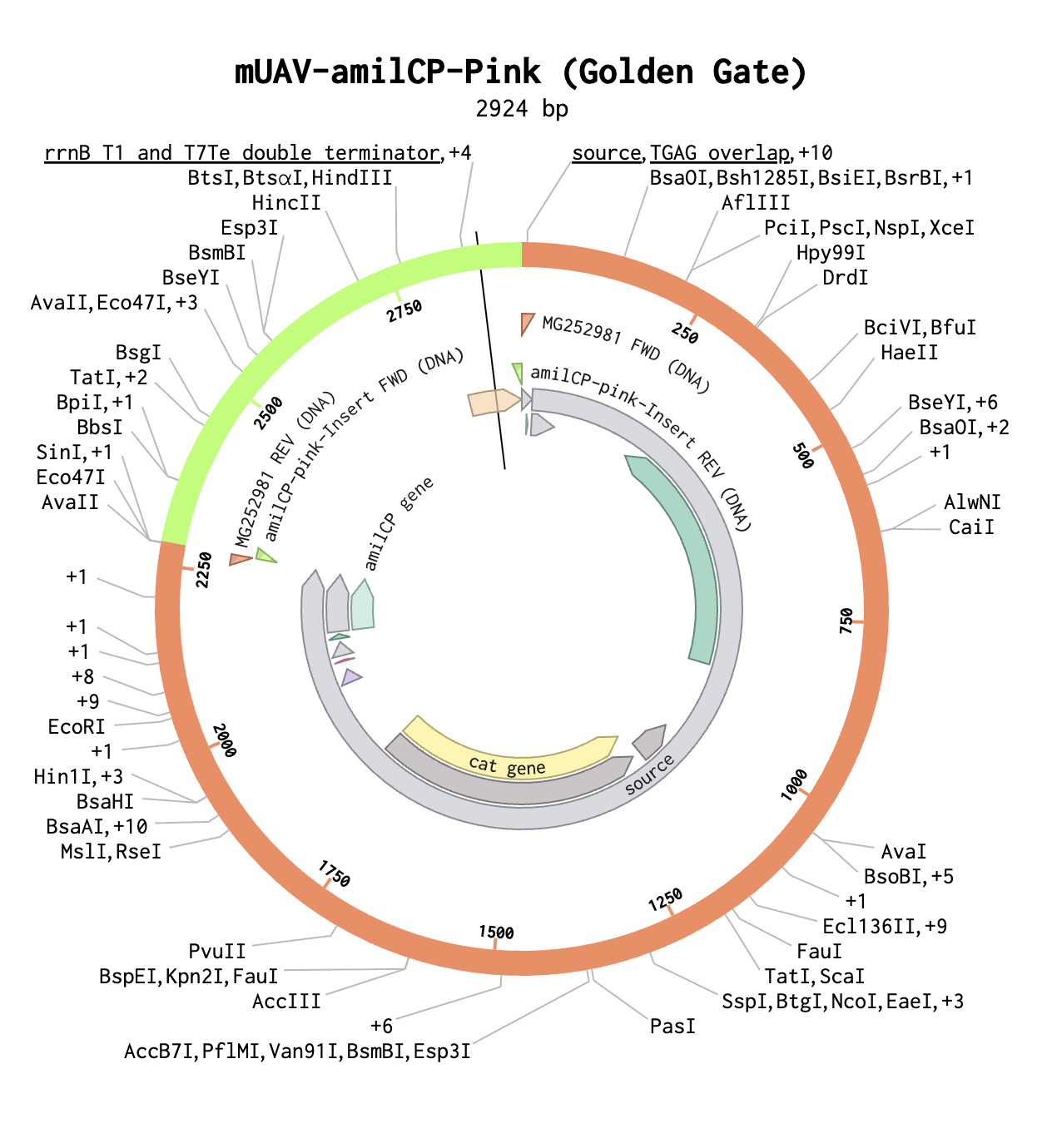

Asimov Kernel
Create a Repository and Notebook
Explore the Bacterial Demos Repo
Explore the devices in the Bacterial Demos Repo to understand how the parts work together by running the Simulator on various examples, following the instructions for the simulator found in the “Info” panel (click the “i” icon on the right to open the Info panel).
Recreate the Repressilator
Create a blank Construct and save it to your Repository. Recreate the Repressilator in that empty Construct by using parts from the Characterized Bacterial Parts repository — search the parts using the Search function in the right menu and drag and drop them into the Construct. Confirm it works as expected by running the Simulator (“play” button) and compare your results with the Repressilator Construct found in the Bacterial Demos repository.
Build Three Custom Constructs
Build three of your own Constructs using the parts in the Characterized Bacterial Parts Repo. For each construct, explain how you think it should function, run the simulator, and share the results. If the results don’t match your expectations, speculate on why and see if you can adjust the simulator settings to get the expected outcome.
Construct 1:
Construct 2:
Construct 3:
Week 7 HW: Genetic Circuits Part 2
Part 1: Intracellular Artificial Neural Networks (IANNs)
What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
IANNs offer three advantages over Boolean genetic circuits. They operate on graded, continuous intracellular signals rather than discrete ON/OFF states, enabling weighted summation, nonlinear activation, and universal function approximation. Weiss-coauthored neuromorphic circuits demonstrated these capabilities through analog computation, soft majority voting, and ternary switching in living cells. IANNs also permit tunable decision boundaries without topological redesign because effective weights and biases can be adjusted by modifying stoichiometry, promoter strength, or recognition-site placement. The PERSIST endoRNase system illustrates this: the same RNase acts as a repressor or activator depending on 5′-UTR versus 3′-UTR target-site positioning. Finally, multilayer IANNs have greater expressive power per circuit, representing smooth classifiers and nonlinearly separable response surfaces that Boolean truth tables cannot efficiently encode.
Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
An autonomous cell-state classifier for stem-cell differentiation would be a strong use case. The IANN would integrate sensors for an endothelial-intermediate RNA signature (x₁), residual pluripotency (x₂), and off-target lineage markers (x₃), computing a weighted sum z = w₁x₁ − w₂x₂ − w₃x₃ + b passed through a nonlinear output node driving a fluorescent reporter or differentiation factor. Weiss and colleagues used endoRNase-mediated miRNA sensors in a similar fashion to monitor cell-state transitions and guide multistep hiPSC differentiation toward a hematopoietic lineage. Limitations include resource loading in mammalian cells (Weiss’s 2020 work showed competing modules can reduce unregulated gene expression by up to 70%), RNase saturation and cross-cleavage at high enzyme ratios as observed in PERSIST cascades, stochastic weight variation across cells from poly-transfection, and the 650 ng total-DNA constraint imposed by the class protocol, which the supplied two-layer design already saturates.
Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.
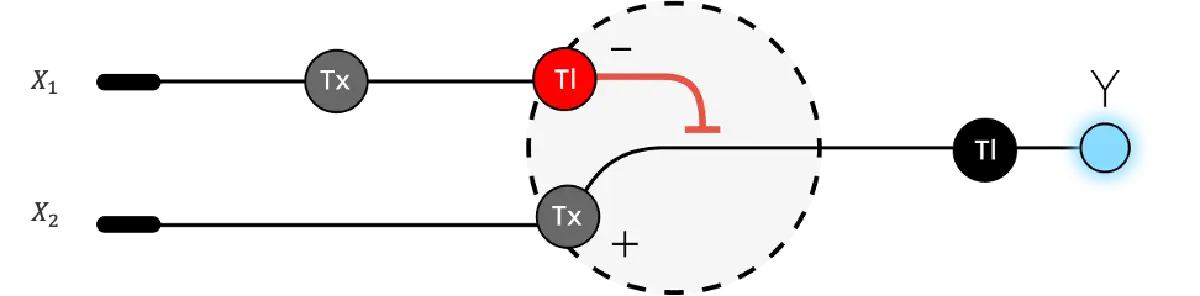

Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
The diagram below shows a two-layer intracellular perceptron built from the supplied parts. In Layer 1, input DNA X1 encodes Csy4, which is transcribed and translated into Csy4 protein that cleaves the Csy4 recognition site on the hidden-layer transcript (Csy4_rec_CasE), repressing it and producing the hidden-node output H = CasE. In Layer 2, CasE protein acts on the CasE recognition site in the output transcript (CasE_rec_mNeonGreen), repressing it to produce the fluorescent output Y = mNeonGreen. Both RNase links are drawn as repression to match the supplied single-layer example. In PERSIST-style designs, the sign of each edge can be inverted by repositioning the recognition site from a 5′-UTR OFF configuration to a 3′-UTR ON configuration. In the provided spreadsheet, this design corresponds to X1 = Csy4 + mKO2, X2 = Csy4_rec_CasE + eBFP2, and Bias = CasE_rec_mNeonGreen, and it consumes the full 650 ng class DNA limit.
Part 2: Fungal Materials
What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Most fungal materials are mycelium-based. Mycelium packaging (used by Dell as a Styrofoam replacement) is made by inoculating agricultural waste with fungal spores and molding it into custom shapes. Mycelium leather from MycoWorks and Bolt Threads (Mylo) uses roughly 70% less water and emits 68% fewer greenhouse gas emissions than cattle leather. Other products include fire-resistant construction and insulation panels with favorable thermal conductivity and sound absorption, compostable foams (Ecovative), and fungal protein food products (Mycorena). The common advantages over traditional counterparts are biodegradability, use of waste feedstocks, and reduced environmental impact. The common disadvantages are limited mechanical performance (mycelium compressive strength around 0.1 to 0.2 MPa versus 17 to 28 MPa for concrete), moisture susceptibility, batch-to-batch variability, scaling difficulties, and in the case of leather substitutes, affordability problems that have forced some manufacturers to shut down.
What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Engineering targets in fungi include modifying cell-wall biosynthetic genes (chitin synthase, alpha-glucan synthase, acetyltransferases) to tune material properties at the genome level, activating silent secondary-metabolite gene clusters through synthetic transcription factors or heterologous expression in hosts like Aspergillus oryzae, producing non-native compounds (cannabinoids, biofuels, therapeutic proteins), and embedding synthetic gene circuits into mycelium to create stimulus-responsive living materials. Fungi have several advantages over bacteria for synthetic biology. As eukaryotes, they perform post-translational modifications (glycosylation, disulfide bonds, proteolytic processing) that are needed for functional human therapeutic proteins. Filamentous fungi secrete 10 to 1,000 times more protein than bacterial hosts. They harbor secondary-metabolite pathways with large intron-containing gene clusters that bacterial systems cannot properly express. Mycelium grows into three-dimensional networks that can be used directly as structural materials, which no bacterial system offers. They also thrive on lignocellulosic waste streams that most bacteria cannot degrade. The tradeoffs are slower growth rates, less well-characterized genetics, and a synthetic biology toolkit that remains less mature than what is available for E. coli, though recent efforts like the Fungal Modular Cloning Toolkit (96 standardized parts for filamentous fungi) are narrowing the gap.
Part 3: First DNA Twist Order
All draft Round 0 constructs have been deposited in Benchling at https://benchling.com/seanmurp/f_/KopGo3fSDI-htgaa_final_project/, organized into sub-folders for Round 0 constructs (by reporter: sfGFP, mCherry, and NanoLuc) and controls.
The library comprises 80 unique 20-nt T7 promoter-spacer variants (+1 to +20) each paired with three reporters (sfGFP, mCherry, NanoLuc), yielding 240 test constructs plus 9 controls (dead-promoter negatives, no-RBS negatives, and synonymous codon-variant sfGFP controls). Spacers are drawn from five design categories: published reference variants (WT T7, T7Max, T7c62, T7#4), systematic ITS mutagenesis at positions +1 to +6, RBS/translation efficiency variants, context-interaction variants designed to produce reporter-dependent expression differences, and random space-filling variants for unbiased landscape coverage.
All constructs are designed as linear DNA fragments. The construct architecture is: 5’-[59 bp buffer]-[T7 consensus promoter]-[20 nt variable spacer]-[reporter CDS]-[T7 terminator]-[59 bp buffer]-3’, with total lengths ranging from ~720 bp (NanoLuc) to ~920 bp (sfGFP). The 59 bp flanking buffers provide protection against residual RecBCD exonuclease activity in the BL21 Star lysate, per Ginkgo’s recommendation of 50–80 bp padding for linear DNA templates in their cell-free system. Constructs will be synthesized as linear gene fragments (e.g., via Twist Bioscience) and used directly as CFPS templates at 15–20 nM without plasmid cloning.
Week 9 HW: Cell Free Systems
Part 1: Cell-Free Protein Synthesis
Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
Cell-free protein synthesis (CFPS) removes the constraint of keeping a living cell alive. In a normal in vivo expression experiment, every design choice has to be compatible with growth, metabolism, membrane integrity, and host viability. In CFPS, the transcription and translation machinery is retained, but the cell itself is gone, so the reaction becomes an open biochemical system that can be directly tuned. DNA concentration, magnesium and potassium levels, redox state, chaperones, cofactors, detergents, lipids, noncanonical amino acids, and energy substrates can all be adjusted without worrying about whether the host will survive.
That open format gives two major advantages. First, CFPS is much more flexible for rapid prototyping: I can test many DNA templates, promoter/RBS designs, or reaction conditions in parallel in a few hours instead of building and transforming strains. Second, it gives tighter experimental control because every important variable is directly set by the user rather than indirectly filtered through cellular regulation. If translation drops, I can alter Mg2+ or template concentration immediately; if a membrane protein aggregates, I can add nanodiscs or detergent directly to the reaction.
Cell-free expression is especially beneficial in at least two cases:
- Toxic proteins: pore-forming toxins, nucleases, or strong metabolic enzymes often kill or stress living hosts, but can still be expressed in CFPS because there is no cell viability to protect.
- Membrane proteins: these are difficult to express in vivo because they misfold, aggregate, or overload the membrane insertion machinery; in CFPS, membrane mimics such as liposomes, nanodiscs, or mild detergents can be added directly.
- Rapid circuit prototyping: gene circuits, biosensors, and promoter libraries can be screened much faster without cloning into cells and waiting for growth.
- Proteins with noncanonical chemistry: CFPS is well suited for adding isotope labels, unnatural amino acids, or unusual cofactors that may be hard for living cells to tolerate or import.
Describe the main components of a cell-free expression system and explain the role of each component.
A typical cell-free expression reaction has several core parts:
- Cell extract or purified Tx/Tl machinery: This is the engine of the system. Crude lysates from E. coli, wheat germ, insect, or mammalian cells contain ribosomes, tRNAs, aminoacyl-tRNA synthetases, translation factors, and many metabolic enzymes. In PURE systems, these are supplied as purified components rather than as a crude extract.
- DNA or mRNA template: This encodes the protein of interest. If DNA is used, it must include promoter, ribosome binding site or Kozak sequence, coding sequence, and terminator/polyadenylation features appropriate to the system.
- Amino acids: These are the building blocks used by ribosomes to make the protein.
- Nucleotides (ATP, GTP, CTP, UTP): These are required for transcription and for many steps of translation and energy transfer.
- Energy source and regeneration system: Protein synthesis consumes large amounts of ATP and GTP, so the reaction needs both an initial energy pool and a way to recycle it.
- Salts and cofactors: Magnesium and potassium are especially important because they control ribosome function, RNA folding, and enzyme activity. Other cofactors such as spermidine, folate derivatives, or reducing agents may also be needed.
- Buffer system: This maintains the pH and ionic environment so the enzymes remain active throughout the reaction.
- Accessory additives: Chaperones, disulfide bond isomerases, detergents, nanodiscs, liposomes, RNase inhibitors, protease inhibitors, or crowding agents can be added depending on the target protein.
In short, CFPS works by reconstituting the minimum biochemical environment needed for transcription and translation, then tuning that environment for the protein or circuit of interest.
Why is energy regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Energy regeneration is critical because protein synthesis is extremely energy-intensive. ATP is required for tRNA charging and many upstream metabolic steps, while GTP is consumed during translation initiation, elongation, and translocation. In a closed reaction, the energy pool is depleted quickly, and inhibitory byproducts such as inorganic phosphate can accumulate. If ATP collapses, transcription slows, translation stalls, and yield drops sharply even if all other components are present.
One practical way to maintain ATP is to use a phosphoenolpyruvate (PEP) plus pyruvate kinase regeneration system. In this setup, ATP is consumed during the reaction and converted to ADP. Pyruvate kinase then transfers the high-energy phosphate from PEP back onto ADP, regenerating ATP continuously. This method is common because it is simple, fast, and effective for short to medium CFPS reactions.
In my experiment, I would pair PEP regeneration with optimization of magnesium and phosphate balance, because even a good energy donor can fail if phosphate buildup poisons the reaction. For longer reactions, I would also consider slower-burning substrates such as 3-phosphoglycerate, glucose, or maltodextrin, which often improve reaction longevity by releasing energy more gradually.
Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
| Feature | Prokaryotic CFPS | Eukaryotic CFPS |
|---|---|---|
| Common source | E. coli lysate or PURE | Wheat germ, insect, rabbit reticulocyte, or mammalian lysate |
| Speed and cost | Fast and inexpensive | Slower and more expensive |
| Yield | Often very high for simple proteins | Lower to moderate, but better for complex proteins |
| PTMs | Limited | Better support for folding, disulfides, and some post-translational modifications |
| Best use cases | Enzymes, reporters, circuit prototyping, bacterial proteins | Secreted proteins, receptors, antibodies, and other eukaryotic targets |
A prokaryotic system is best when the goal is speed, low cost, and high yield for proteins that do not require elaborate post-translational processing. If I wanted to produce sfGFP, I would choose an E. coli CFPS system because sfGFP folds well in bacterial conditions, does not need glycosylation, and can be produced quickly at high yield. It is also an ideal reporter for reaction optimization because fluorescence gives a direct readout of productive expression.
A eukaryotic system is preferable when the protein requires a eukaryotic folding environment, disulfide bond formation, microsomal insertion, or other processing steps. If I wanted to produce human erythropoietin (EPO), I would choose a mammalian or insect-derived cell-free system because EPO is a secreted human glycoprotein whose activity and stability depend strongly on proper eukaryotic folding and post-translational processing. An E. coli lysate could make the polypeptide, but it would be much less likely to produce a properly folded, functional therapeutic-like product.
How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
To optimize a membrane protein, I would design the experiment around co-translational insertion into a membrane mimic rather than expressing the protein into free solution and hoping it folds afterward. As a concrete example, I would use an E. coli CFPS system to express the bacterial potassium channel KcsA with a C-terminal GFP tag for rapid screening. The reaction would include preassembled nanodiscs made from MSP1D1 scaffold protein and a POPC:POPG lipid mixture, because KcsA is far more likely to remain soluble and native-like if it inserts into a bilayer during translation.
The main challenges are:
- Aggregation: hydrophobic transmembrane segments tend to precipitate in aqueous solution.
- Misfolding: even if the protein is made, it may not adopt the correct conformation or oligomeric state.
- Poor membrane insertion: the reaction may produce full-length protein that never enters a lipid environment.
- Reaction inhibition: detergents, excess DNA, or incorrect salt balance can reduce overall translation efficiency.
To address these issues, I would screen a matrix of conditions:
- nanodiscs versus small liposomes versus mild detergents such as DDM or LMNG
- low versus moderate DNA concentration
- 25, 30, and 37 degrees C reaction temperatures
- magnesium concentration and potassium glutamate concentration
- optional chaperone supplementation such as DnaK/DnaJ/GrpE
I would measure three outputs: total protein made, soluble or membrane-associated fraction, and functional activity after reconstitution. Total yield could be checked by SDS-PAGE or in-gel GFP fluorescence. Membrane insertion could be assessed by co-migration with nanodisc fractions or flotation assays. Function could be tested with a potassium flux assay after purification or direct reconstitution. The best condition would not simply be the one with the most protein, but the one that gives the highest amount of correctly inserted and functional channel.
Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
Three common causes of low CFPS yield are:
Poor template design or poor template quality.
If the promoter is weak, the RBS is poorly matched, the DNA is degraded, or the coding sequence has problematic secondary structure, transcription and translation can both suffer. I would troubleshoot by checking DNA quality, comparing plasmid versus linear template, redesigning the 5’ untranslated region, and testing a stronger promoter or codon-optimized construct.Incorrect reaction chemistry.
CFPS depends sensitively on magnesium, potassium, pH, and energy balance. A reaction that is slightly off can collapse even when all components are present. I would run a small design-of-experiments screen varying Mg2+, K+, DNA concentration, and energy substrate, while using a known positive-control template such as sfGFP to determine whether the problem is the reaction mixture or the target itself.Protein instability, aggregation, or degradation.
Some proteins fold poorly, are protease-sensitive, or precipitate as they are made. I would troubleshoot by lowering reaction temperature, shortening reaction time, adding chaperones, adding protease inhibitors, or including membrane mimics or redox helpers if the target is a membrane protein or disulfide-rich protein.
Low yield is usually not caused by one single factor. In practice, I would troubleshoot in the order of template quality, reaction chemistry, and protein-specific folding issues, because that sequence separates general reaction failure from target-specific failure.
Part 2: Design of a Useful Synthetic Minimal Cell
1. Pick a function and describe it.
I would design a synthetic minimal cell (SMC) that senses theophylline and, in response, activates a nearby engineered probiotic bacterium. The idea is to convert a small molecule that the bacterium does not naturally monitor into a standard bacterial induction signal.
- Function: user-controlled activation of a probiotic gene program
- Input: theophylline
- Output of the synthetic minimal cell: IPTG release
- Output of the whole hybrid system: sfGFP in a proof-of-principle strain of E. coli Nissle 1917, or a therapeutic payload in a future version
This function could not be realized by cell-free Tx/Tl alone without encapsulation. If IPTG were simply mixed into a bulk cell-free reaction, it would diffuse directly to the bacteria and there would be no gated actuator step. The membrane compartment is what lets the SMC store the output signal until the input molecule triggers pore formation.
This function could be realized by a genetically modified natural cell, but that would require engineering a living probiotic to directly sense theophylline and carry the entire logic internally. The synthetic-cell version is more modular: the same probiotic responder could be paired with many different SMC sensors just by swapping the sensing module.
The desired outcome is that the probiotic turns on only when theophylline is present, giving an external chemical control knob over bacterial behavior without permanently hard-wiring the sensing logic into the living cell.
2. Design all components that would need to be part of the synthetic cell.
| Component | Design Choice | Why |
|---|---|---|
| Membrane | POPC:cholesterol vesicle, optionally stabilized with a small amount of DSPE-PEG2000 | Stable phospholipid compartment that can hold small molecules and support pore insertion |
| Tx/Tl source | E. coli cell-free expression system | Fast, inexpensive, and compatible with bacterial riboswitch control |
| Input sensing module | Theophylline-responsive riboswitch upstream of pore gene | Theophylline is membrane permeable and the riboswitch can directly control translation |
| Output release module | Alpha-hemolysin pore | Allows stored IPTG to exit only after the sensor is activated |
| Encapsulated cargo | IPTG, amino acids, nucleotides, salts, energy substrate, cell-free enzymes | IPTG is the communication signal; the rest are required for expression of the pore |
| Receiver cell | E. coli Nissle carrying a LacI-regulated reporter plasmid | Converts released IPTG into an easily measured bacterial response |
I would use a bacterial Tx/Tl system, not a mammalian one, because the key regulatory element here is a small-molecule riboswitch and the output is just pore formation and inducer release. No mammalian glycosylation or nuclear machinery is needed.
The SMC would communicate with the environment in two steps. First, theophylline diffuses across the vesicle membrane and binds the riboswitch, turning on pore synthesis. Second, alpha-hemolysin inserts into the vesicle membrane and releases encapsulated IPTG, which then diffuses to the surrounding probiotic cells and activates their lac-regulated gene circuit.
3. Experimental details
Lipids and genes
- Lipids: POPC, cholesterol, DSPE-PEG2000
- Tx/Tl system: E. coli S30 extract or PURE system
- Energy system: 3-phosphoglycerate or PEP-based ATP regeneration
- Synthetic-cell gene: Staphylococcus aureus
hlaencoding alpha-hemolysin, controlled by a theophylline riboswitch - Encapsulated small-molecule cargo: IPTG
- Responder-cell genes: constitutive
lacIplussfGFPunderPlacUV5orPtacin E. coli Nissle 1917
Measurement strategy
I would measure function primarily through the GFP output of the responder bacteria. In the presence of theophylline, the SMC should synthesize alpha-hemolysin, release IPTG, and induce bacterial GFP. The cleanest readout would be flow cytometry or plate-reader fluorescence of the E. coli Nissle reporter strain.
Key controls would include:
- no theophylline
- no
hlaDNA - SMCs without encapsulated IPTG
- responder bacteria without the lac-regulated reporter
If needed, IPTG release could also be confirmed indirectly by comparing fluorescence kinetics or directly by chemical assay of the supernatant.
Part 3: Freeze-Dried Cell-Free Systems in Materials
One-sentence pitch
I propose a soft-robotic skin with embedded freeze-dried cell-free microcapsules that detect damage, generate a visible warning signal, and locally produce a crosslinking enzyme to help seal small tears.
How will the idea work?
The robotic skin would contain patterned microcapsules loaded with freeze-dried cell-free reactions, a DNA template for a visible chromoprotein, and a DNA template for microbial transglutaminase. These capsules would be embedded inside an elastomer layer that also contains a thin repair hydrogel rich in crosslinkable residues. When the skin is punctured or torn, a built-in water reservoir or ambient moisture would rehydrate the damaged region and activate the local cell-free reactions. The chromoprotein would mark the damaged area for easy inspection, while transglutaminase would crosslink the repair layer and help slow crack growth or fluid leakage long enough for replacement.
What societal challenge or market need will this address?
Soft robots are increasingly used in medical devices, warehouse automation, and search-and-rescue environments, but their compliant materials are vulnerable to small tears, abrasion, and puncture. Today, many failures are only discovered after performance drops or a leak becomes severe. A self-reporting, partially self-sealing skin would reduce downtime, improve safety, and make soft robots more practical in environments where immediate maintenance is difficult.
How do you envision addressing the limitation of cell-free reactions?
I would address activation and stability by storing the reactions in trehalose-stabilized, vacuum-sealed microcapsules laminated inside the material until damage occurs. Water-triggering is actually useful here, because damage can be coupled to capsule rupture or exposure to a local hydration layer. The one-time-use limitation can be handled by making the sensing-and-repair elements modular and replaceable, like sacrificial patches in high-strain regions. For long shelf life, the material would use oxygen and moisture barrier films so the cell-free modules stay dormant until needed.
Part 4: Mock Genes in Space Proposal
1. Background information
Long-duration missions may depend on dried DNA templates for on-demand production of medicines, enzymes, and diagnostics. Space radiation and temperature cycling could damage these templates and reduce the reliability of cell-free manufacturing. I want to test how well lightweight shielding preserves the functional expression capacity of stored DNA. This matters for humanity because future crews will need compact, stable biotechnology systems far from Earth, and it is scientifically interesting because it connects the space environment directly to the survival of usable genetic information.
2. Molecular or genetic target
Plasmid DNA encoding sfGFP under a T7 promoter, plus the T7-promoter-to-sfGFP junction as a PCR integrity marker.
3. How the target relates to the challenge
If spaceflight damages the promoter or coding sequence, BioBits should produce less GFP even when the same amount of DNA is added. Measuring fluorescence therefore converts DNA integrity into a simple functional readout. By comparing shielded and unshielded templates, I can test whether stored genetic instructions remain usable for future in-space biomanufacturing and biosensing.
4. Hypothesis or research goal
My hypothesis is that DNA stored behind lightweight, hydrogen-rich shielding will retain higher functional expression capacity than unshielded DNA after space exposure. The goal is to compare practical storage strategies for preserving genetic templates that could later be used in cell-free systems aboard spacecraft. This hypothesis is based on the fact that ionizing radiation causes strand breaks and base damage, while hydrogen-rich materials can reduce secondary particle damage more effectively than many denser materials. A functional BioBits readout is especially useful because a template may still be amplifiable by PCR yet perform poorly in transcription or translation.
5. Experimental plan
I would test freeze-dried plasmid aliquots stored in three conditions: unshielded, polyethylene-shielded, and aluminum-shielded, with matched ground controls. After exposure, each sample would be rehydrated in BioBits and GFP output would be measured with the P51 Molecular Fluorescence Viewer at fixed time points. The miniPCR would amplify the T7-sfGFP region from the same samples as an integrity control. Fresh plasmid would serve as a positive control, and no-DNA reactions would serve as negative controls.
Week 10 HW: Advanced Imaging
Homework: Final Project
Q1. Please identify at least one (ideally many) aspect(s) of your project that you will measure.
This project has four distinct measurable outputs that span computational filtering, protein expression, antimicrobial activity, and drug synergy:
Peptide physicochemical properties (computational, pre-synthesis): During AI candidate selection, I’ll measure charge, amphipathicity, and hydrophobic moment of ~2,000 AMP-Diffusion candidates, as well as CLIP binding scores for PepPrCLIP candidates against E. coli FtsZ and LpxC targets. PeptiVerse provides predicted hemolysis probability, solubility, and toxicity scores for all final candidates.
Bacterial growth inhibition ($\text{OD}_{600}$): This is the core experimental measurement. After expressing each peptide via cell-free protein synthesis, I’ll read optical density at 600 nm on both E. coli ATCC 25922 and B. subtilis ATCC 6633 plates after overnight incubation. Each peptide’s $\text{OD}_{600}$ is compared to the scrambled-peptide negative control to calculate percent growth inhibition, producing a 2D activity matrix (peptide $\times$ organism).
Fractional Inhibitory Concentration Index (FICI) for synergy: For the top 5–6 active peptides, I’ll measure $\text{OD}_{600}$ of co-expressed pairs (both DNA templates at half-dose in one CFPS reaction) vs. each peptide expressed alone at half-dose. FICI classifies each pair as synergistic ($\leq 0.5$), additive ($0.5$–$1.0$), or indifferent/antagonistic ($> 1.0$). This is the measurement that directly answers the central hypothesis about whether cross-method AMP pairs are more synergistic than within-method pairs.
Gram-selectivity profiles: Running every peptide against both organisms generates a selectivity ratio (% inhibition on E. coli vs. B. subtilis). This is especially important for Group C constructs; if MadSBM becomes available, the 25/50/75% interpolants between magainin-2 (gram-negative) and HNP-1 (gram-positive) should show a measurable shift in this ratio.
Q2. Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements.
Computational measurements are performed before any wet lab work. AMP-Diffusion generates ~2,000 candidate sequences at lengths 20/25/30/35 amino acids; I filter these programmatically by physicochemical properties (removing sequences with unfavorable charge, low amphipathicity, or homopolymer runs) and select the top 6 diverse candidates plus 3 fallbacks. PepPrCLIP ranks ~100K candidates per target by CLIP binding score, and I take the top 2 per target. PeptiVerse runs as a HuggingFace web app and returns developability predictions per peptide.
OD600 growth inhibition assay follows a standard broth microdilution format. I dilute overnight cultures of each organism to ~5 x 10^5 CFU/mL in Mueller-Hinton broth, dispense 100 uL per well of a 96-well flat-bottom plate, then add 5 uL of crude CFPS reaction to each well. After overnight incubation at 37 °C, I read absorbance at 600 nm using a plate reader. Three biological replicates per construct (45 reactions for 15 constructs, plus 9 control reactions) enable statistical comparison. The same CFPS reactions are split across two plates (one per organism) so expression variability is controlled between the two bacterial targets.
Synergy measurement uses the same OD600 readout but with modified CFPS input: two DNA templates at half-dose (25–50 ng each) in a single 20 uL reaction, alongside single-agent half-dose controls. I then calculate FICI from the resulting inhibition values, separately for each organism. Cross-method pairings (e.g., AMP-Diffusion generalist + PepPrCLIP targeted binder) are prioritized because they test the central synergy hypothesis most directly.
Gram-selectivity measurement is a derived metric: no separate experiment is needed. By reading the same CFPS reactions against both organisms in parallel, every peptide’s selectivity ratio drops out of the primary screen data automatically.
Q3. What are the technologies you will use? Describe in detail.
Cell-free protein synthesis (CFPS): The Ginkgo Bioworks E. coli cell-free kit (BL21 Star DE3 lysate, T7 RNA polymerase-driven) is the expression platform. Linear Twist gene fragments serve directly as templates, with no cloning required. Each construct carries a T7 promoter, strong RBS, the codon-optimized peptide ORF, and a T7 terminator. NEBExpress GamS Nuclease Inhibitor (NEB #P0774S) is added at ~0.6 $\mu\text{g}$ per 20 $\mu\text{L}$ reaction to protect linear DNA from RecBCD exonuclease degradation in the crude lysate. Reactions run at 30 $^\circ\text{C}$ for 4 hours.
Synthetic gene fragments (Twist Bioscience): 15 linear DNA constructs ($\geq 300$ bp each, padded with inert flanking sequence to meet Twist’s minimum) are ordered as gene fragments. This is DNA synthesis, not cloning; the fragments arrive ready for direct use in CFPS.
$\text{OD}_{600}$ plate reader (spectrophotometry): A standard microplate reader measuring optical density at 600 nm is the primary analytical instrument. It quantifies bacterial growth in 96-well format, enabling high-throughput comparison of all peptides and combinations across both organisms in a single read.
AI/ML peptide design tools: AMP-Diffusion (diffusion-based generative model for antimicrobial peptide sequences), PepPrCLIP (CLIP-based peptide design using the 650M-parameter ESM-2 protein language model, run on Google Colab with GPU), and potentially MadSBM (latent-space interpolation between known AMPs). These are the computational “technologies” that generate the candidate peptides before any synthesis.
Codon optimization: Selected peptide sequences are reverse-translated and codon-optimized for E. coli expression (likely using IDT or Benchling codon optimization tools) to maximize translational efficiency in the BL21-derived CFPS lysate.
Standard microbiology (Mueller-Hinton broth microdilution): This CLSI-standard antimicrobial susceptibility testing method uses the two reference strains E. coli ATCC 25922 and B. subtilis ATCC 6633, both standard quality-control organisms for susceptibility testing, in 96-well format.
Part I — Molecular Weight
Q1. Calculate the theoretical molecular weight of eGFP from the amino acid sequence using ExPASy ProtParam.
The full eGFP construct (247 amino acids, including the C-terminal LE linker and $\text{His}_6$ tag) was submitted to ExPASy ProtParam:
ProtParam reports:
| Property | Value |
|---|---|
| Number of amino acids | 247 |
| Average molecular weight | 28,006.60 Da |
| Monoisotopic molecular weight | 27,988.96 Da |
| Theoretical pI | ~6.2 |
The average molecular weight (28,006.60 Da) is the reference value used below for accuracy calculations. Note that this theoretical value does not account for eGFP chromophore maturation, which removes approximately 20 Da (one water loss + one oxidation) via autocatalytic cyclization of residues Thr65–Tyr66–Gly67. The mature chromophore mass would be closer to $28{,}006.60 - 20.03 \approx 27{,}986.57$ Da.
Q2. Determine the molecular weight from the LC-MS charge state envelope using the adjacent-charge-state method.
In electrospray ionization, a protein of mass $M$ carrying $z$ protons (each of mass $H = 1.00728$ Da) appears at:
$$\frac{m}{z} = \frac{M + z \cdot H}{z}$$
For two adjacent peaks where Peak A has charge $z$ and Peak B has charge $z - 1$:
$$z = \frac{(m/z)_B - H}{(m/z)_B - (m/z)_A}$$
and then:
$$M = z \left[\left(\frac{m}{z}\right)_A - H\right]$$
Worked example: using the peaks at $m/z$ = 903.7148 (Peak A) and 933.8044 (Peak B):
$$z_A = \frac{933.8044 - 1.00728}{933.8044 - 903.7148} = \frac{932.797}{30.090} = 30.998 \approx 31$$
$$M = 31 \times (903.7148 - 1.00728) = 31 \times 902.708 = 27{,}983.9 ;\text{Da}$$
Cross-check table (five adjacent pairs from the denatured LC-MS spectrum):
| $(m/z)_A$ | $(m/z)_B$ | $z_A$ (calc → round) | $M$ (Da) |
|---|---|---|---|
| 757.3019 | 778.2277 | 37.14 → 37 | 27,982.9 |
| 903.7148 | 933.8044 | 31.00 → 31 | 27,983.9 |
| 933.8044 | 965.9584 | 30.01 → 30 | 27,983.9 |
| 965.9584 | 1000.5021 | 28.99 → 29 | 27,983.6 |
| 1000.5021 | 1037.4423 | 28.04 → 28 | 27,985.9 |
$$M_{\text{experiment}} = \text{mean of five values} \approx \mathbf{27{,}984.0 ;\text{Da}}$$
Q2 (cont.). Calculate the accuracy of the measurement.
$$\text{Accuracy} = \frac{|M_{\text{exp}} - M_{\text{theory}}|}{M_{\text{theory}}} = \frac{|27{,}984.0 - 28{,}006.6|}{28{,}006.6} = \frac{22.6}{28{,}006.6} = 0.081% \approx \mathbf{810 ;\text{ppm}}$$
This is relative to the average theoretical mass from ProtParam. If instead we compare to the monoisotopic mass (27,988.96 Da), the error drops to $|27{,}984.0 - 27{,}988.96|/27{,}988.96 \approx 177$ ppm, and if we further account for the ~20 Da chromophore maturation ($M_\text{theory,mature} \approx 27{,}986.6$ Da), the agreement improves to roughly 90 ppm. The remaining discrepancy is well within the expected accuracy of intact-protein ESI-MS deconvolution.
Q3. Can you observe the charge state from the zoomed-in peak? If yes, what is it? If no, why not?
Whether the charge state can be read directly from a single peak depends on mass resolving power. For eGFP at $z \approx 30$, adjacent isotope peaks in the isotopic envelope are separated by:
$$\Delta(m/z) = \frac{1.003}{z} \approx \frac{1.003}{30} \approx 0.033 ;\text{Da}$$
Resolving this requires $R = m/z / \Delta \approx 1{,}000 / 0.033 \approx 30{,}000$. If the instrument (e.g., Orbitrap) achieves this resolution, the isotope peaks are resolved and the charge state can be determined by:
$$z = \frac{1.003}{\text{spacing between adjacent isotope peaks}}$$
If the zoomed-in inset shows resolved isotope peaks with spacing $\sim$0.033 Da, then $z = 1.003/0.033 \approx 30$, confirming the charge state directly.
If the instrument resolution is insufficient (e.g., a low-resolution QTOF), the isotope peaks merge into a single broad hump and the charge state cannot be determined from that peak alone, so the adjacent-charge-state method (Q2) must be used instead.
Part II — Secondary and Tertiary Structure
Q1. Explain the difference between native and denatured protein conformations as seen in mass spectrometry.
In denatured ESI-MS (Figure 2, top panel), the protein is unfolded by organic solvent and acid. The extended chain exposes many basic residues (Lys, Arg, His) to solution, each of which can accept a proton. This produces a broad charge state distribution at high charge states ($z \approx 27$–$37$ for eGFP), so the peaks appear at relatively low $m/z$ values (~750–1050). The wide, multi-peak envelope is a hallmark of a disordered, extended conformation.
In native ESI-MS (Figure 2, bottom panel), the protein is sprayed from a near-physiological buffer (typically ammonium acetate, pH ~7). The protein remains compactly folded, burying most ionizable side chains in its interior. This results in fewer, lower charge states ($z \approx 9$–$11$ for eGFP), so the peaks appear at high $m/z$ values (~2500–3100). The narrow charge state distribution (often only two or three peaks) directly reflects the compact, globular conformation.
Key insight: the charge state distribution is a proxy for protein conformation. Compact → fewer charges → higher $m/z$; unfolded → more charges → lower $m/z$.
Q2. Zooming into the native mass spectrum at ~2800 m/z (Figure 3), can you discern the charge state? What is it?
Yes. At $m/z \approx 2800$ for a protein of mass ~28,000 Da, the charge state is:
$$z = \frac{M}{m/z} \approx \frac{28{,}000}{2{,}800} = \mathbf{10}$$
This can be confirmed from the isotopic fine structure. If the inset shows resolved isotope peaks, the spacing between adjacent isotopic peaks is:
$$\Delta(m/z) = \frac{1.003}{z} = \frac{1.003}{10} = 0.1003 ;\text{Da}$$
Counting approximately 10 isotope peaks per 1 Da interval, or measuring the spacing directly and computing $z = 1.003 / \Delta$, confirms $z = 10$. Resolving this spacing requires $R = 2800 / 0.10 = 28{,}000$, which is achievable on modern Orbitrap and FT-ICR instruments.
As a consistency check: $(28{,}006.6 + 10 \times 1.007)/10 = 2{,}801.7$ $m/z$, which matches the observed peak position.
Part III — Peptide Mapping
Q1. How many Lysines (K) and Arginines (R) are in the eGFP sequence?
The eGFP construct contains 20 Lysines (K) and 6 Arginines (R), for a total of 26 tryptic cleavage sites.
Highlighted sequence (K and R in bold):
MVSKGEELFTGVVPILVELDGDVNGHKFSVSGEGEGDATYGKLTLKFICTTGKLPVPWPTL VTTLTYGVQCFSRYPDHMKQHDFFKSAMPEGYVQERTIFFKDDGNYKTRAEVKFEGDTLVN RIELKGIDFKEDGNILGHKLEYNYNSHNVYIMADKQKNGIKVNFKIRHNIEDGSVQLAD HYQQNTPIGDGPVLLPDNHYLSTQSALSKDPNEK RDHMVLLEFVTAAGITLGMDELYKLE HHHHHH
Q2. How many peptides are expected from a tryptic digest? How many have $[\text{M+H}]^+$ > 500 Da?
Using ExPASy PeptideMass with trypsin (cleaves after K and R, no missed cleavages), the 26 cleavage sites produce 27 tryptic peptides.
Of these 27, 19 peptides have a monoisotopic $[\text{M+H}]^+ > 500$ Da and are therefore likely to be detected by LC-MS. The remaining 8 are very small (1–4 residues) and typically fall below the practical detection or retention limit.
Representative predicted peptides (monoisotopic $[\text{M+H}]^+$):
| # | Residues | Sequence | $[\text{M+H}]^+$ (Da) |
|---|---|---|---|
| 1 | 1–4 | MVSK | 464.25 |
| 2 | 5–27 | GEELFTGVVPILVELDGDVNGHK | 2437.26 |
| 3 | 28–42 | FSVSGEGEGDATYGK | 1503.66 |
| 5 | 47–53 | FICTTGK | 769.39 |
| 6 | 54–74 | LPVPWPTLVTTLTYGVQCFSR | 2378.26 |
| 9 | 87–97 | SAMPEGYVQER | 1266.58 |
| 14 | 115–123 | FEGDTLVNR | 1050.52 |
| 17 | 133–141 | EDGNILGHK | 982.50 |
| 23 | 170–210 | HNIEDGSVQLADHYQQNTPIGDGPVLLPDNHYLSTQSALSK | 4472.18 |
| 26 | 217–239 | DHMVLLEFVTAAGITLGMDELYK | 2566.29 |
| 27 | 240–247 | LEHHHHHH | 1083.50 |
Q3. How many chromatographic peaks are visible in the peptide map between 0.5 and 6.0 minutes (Figure 5a)?
Counting the labeled peaks in Figure 5a with retention times between 0.5 and 6.0 minutes and relative intensity above ~10%:
0.61, 0.79, 1.20, 1.43, 1.80, 1.85, 1.93, 2.17, 2.26, 2.54, 2.78, 3.27, 3.53, 3.59, 3.70, 4.30, 4.48, 4.64, 4.87, 5.06, 5.43
Approximately 19–21 peaks, depending on the intensity threshold used and whether closely spaced doublets (e.g., 1.80/1.85 and 3.53/3.59) are counted as one or two.
Q4. Does the number of chromatographic peaks match the predicted number of tryptic peptides?
The counts roughly agree but are not identical. We predicted 19 peptides with $[\text{M+H}]^+ > 500$ Da and observe ~19–21 chromatographic peaks. The differences arise from:
- Very small peptides not detected: R (175 Da), QK (275 Da), TR (276 Da), IR (288 Da), and a few other small fragments elute in the void volume or fall below the detection limit, reducing the observed count.
- Co-elution: Some peptides with similar hydrophobicity may co-elute and appear as a single peak, further reducing the count.
- Modifications or partial cleavages: Oxidized or miscleaved forms of some peptides can produce extra peaks.
Overall, the observed ~19–21 peaks are consistent with the predicted 19 detectable tryptic peptides.
Q5. Identify the peptide in Figure 5b. What is the charge state, and what is the $[\text{M+H}]^+$ mass?
The dominant peak in Figure 5b is at $m/z = 525.76712$. A second peak is visible at $m/z = 1050.52438$.
The relationship between these two peaks reveals the charge:
$$2 \times 525.767 = 1051.534 \approx 1050.524 + 1.007$$
The 525.767 peak is the doubly charged $[\text{M+2H}]{2+}$ ion, and the 1050.524 peak is the singly charged $[\text{M+H}]{+}$ ion. Therefore $z = 2$.
$$[\text{M+H}]^+ = z \times (m/z) - (z-1) \times H = 2 \times 525.76712 - 1 \times 1.00728 = \mathbf{1050.527 ;\text{Da}}$$
This can be confirmed by the direct singly charged peak at $m/z = 1050.524$ Da.
Q6. Identify the peptide by comparison with PeptideMass, and calculate the mass accuracy in ppm.
Comparing the observed $[\text{M+H}]^+ = 1050.527$ Da against the PeptideMass output, the match is peptide FEGDTLVNR (residues 115–123), with a predicted monoisotopic $[\text{M+H}]^+ = 1050.5214$ Da.
$$\text{ppm error} = \frac{|1050.527 - 1050.5214|}{1050.5214} \times 106 = \frac{0.0056}{1050.5214} \times 106 \approx \mathbf{5.3 ;\text{ppm}}$$
Using the singly charged peak directly ($m/z = 1050.52438$):
$$\text{ppm error} = \frac{|1050.52438 - 1050.5214|}{1050.5214} \times 10^6 \approx \mathbf{2.8 ;\text{ppm}}$$
Both values represent excellent mass accuracy, typical of Orbitrap instruments (specification $\leq 5$ ppm).
Q7. What percentage of the eGFP sequence is confirmed by peptide mapping?
From Figure 6, 88% of the eGFP sequence was identified with high confidence by peptide mapping. The unconfirmed 12% corresponds primarily to the very small tryptic fragments (R, QK, TR, IR) that are too small to be retained or detected, and possibly the large 41-residue peptide (HNIEDGSVQLAD…SALSK) which may have had poor chromatographic recovery.
Part IV — KLH Oligomers by CDMS
Identify the KLH oligomeric species on the CDMS spectrum (Figure 7).
Keyhole limpet hemocyanin (KLH) is built from two subunit types: a 7-functional-unit (7FU) monomer of 340 kDa and an 8-functional-unit (8FU) monomer of 400 kDa. These assemble into decamers and higher-order multimers.
| CDMS Peak (MDa) | Assignment | Expected Mass | Calculation | Match |
|---|---|---|---|---|
| 3.4 | 7FU Decamer | 3.40 MDa | $10 \times 340;\text{kDa}$ | exact |
| 4.01 | 8FU Decamer | 4.00 MDa | $10 \times 400;\text{kDa}$ | 0.3% |
| 8.33 | 8FU Didecamer | 8.00 MDa | $20 \times 400;\text{kDa}$ | 4.1% |
| 12.67 | 8FU 3-Decamer | 12.00 MDa | $30 \times 400;\text{kDa}$ | 5.6% |
| — | 8FU 4-Decamer | 16.00 MDa | $40 \times 400;\text{kDa}$ | not visible |
The 7FU Decamer ($10 \times 340 = 3{,}400$ kDa) matches the 3.4 MDa peak precisely. The 8FU Didecamer ($20 \times 400 = 8{,}000$ kDa) corresponds to the ~8.33 MDa peak, and the 8FU 3-Decamer ($30 \times 400 = 12{,}000$ kDa) corresponds to the ~12.67 MDa peak. The slight upward mass shifts in the didecamer and 3-decamer peaks likely reflect associated solvent, salt, or lipid.
The 8FU 4-Decamer ($40 \times 400 = 16{,}000$ kDa = 16.0 MDa) is not clearly visible on the spectrum, suggesting it is either absent from this preparation, present at very low abundance, or beyond the measured mass range.
Additional peaks visible in Figure 7 at ~0.79 and ~1.52 MDa likely correspond to sub-decameric fragments (dimers and tetramers of 7FU or 8FU subunits).
Part V — Did I Make GFP?
| Property | Theoretical | Observed (Intact LC-MS) | PPM Error |
|---|---|---|---|
| Molecular weight (kDa) | 28.007 | ~27.984 | ~820 |
| Peptide mapping coverage | 100% | 88% | — |
| Peptide FEGDTLVNR $[\text{M+H}]^+$ (Da) | 1050.5214 | 1050.5270 | ~5 |
Conclusion: Yes. The intact mass agrees with the theoretical eGFP mass to within ~820 ppm (largely explained by GFP chromophore maturation, which removes ~20 Da and is not reflected in the ProtParam theoretical value). The tryptic peptide map confirms 88% of the amino acid sequence with sub-5 ppm peptide mass accuracy. Together, the intact mass and sequence-level peptide coverage provide strong orthogonal confirmation that the expressed protein is eGFP.
Week 11 HW: Bioproduction & Cloud Labs
Cell-Free Protein Synthesis Lab — Questions & Answers
Q1. Provide a 1–2 sentence description of each component’s role in the 20-hour NMP-Ribose-Glucose master mix.
E. coli Lysate
BL21 (DE3) Star Lysate — Provides the core transcription/translation machinery (ribosomes, tRNAs, aminoacyl-tRNA synthetases, initiation/elongation/release factors, and metabolic enzymes). The “Star” strain carries an RNase E mutation that stabilizes mRNA, and the (DE3) lysogen supplies T7 RNA Polymerase for high-level transcription from T7 promoters.
Salts / Buffer
- Potassium Glutamate — Supplies $\text{K}^+$ ions critical for ribosome assembly, tRNA binding, and translation fidelity; glutamate is the preferred counter-ion because $\text{Cl}^-$ inhibits many lysate enzymes.
- HEPES-KOH pH 7.5 — A zwitterionic buffer that holds the reaction near physiological pH, preventing acidification as glycolysis and ATP hydrolysis generate protons over the long incubation.
- Magnesium Glutamate — $\text{Mg}^{2+}$ is an essential cofactor for RNA polymerase, ribosomes (stabilizes rRNA tertiary structure and the small/large subunit interface), and virtually every NTP-using enzyme in the system.
- Potassium Phosphate Monobasic / Dibasic (1.6:1 ratio) — Provides inorganic phosphate ($\text{P}_\text{i}$) that feeds substrate-level phosphorylation in glycolysis to regenerate ATP from ADP, while the dibasic:monobasic ratio sets the buffering pH.
Energy / Nucleotide System
- Ribose — Phosphorylated by ribokinase to ribose-5-phosphate, which feeds the pentose phosphate pathway and serves as a precursor for nucleotide salvage / regeneration of NTPs from NMPs.
- Glucose — The primary carbon and energy source; glycolysis converts it to pyruvate, generating ATP and NADH that drive sustained energy regeneration over the 20-hour reaction.
- AMP, CMP, UMP — Nucleoside monophosphate precursors that endogenous kinases (NMP and NDP kinases) phosphorylate to ATP, CTP, and UTP for transcription; cheaper and more stable than supplying NTPs directly.
- GMP — Listed at 0 mM in this recipe; GTP is instead generated from guanine via the salvage pathway, avoiding the cost of GMP and reducing inhibitory phosphate accumulation.
- Guanine — Converted to GMP by HPT (hypoxanthine/guanine phosphoribosyltransferase) using PRPP, then phosphorylated to GTP for transcription and translation (GTP powers initiation, elongation, and release).
Translation Mix (Amino Acids)
- 17 Amino Acid Mix — Supplies 17 of the 20 proteinogenic amino acids used as substrates by aminoacyl-tRNA synthetases to charge tRNAs for protein synthesis.
- Tyrosine (pH 12) — Tyrosine is poorly soluble near neutral pH, so it is prepared in a high-pH stock and added separately to ensure it stays in solution at the correct concentration.
- Cysteine — Added separately because cysteine readily oxidizes to cystine (and forms disulfides), so it requires its own fresh stock to deliver reduced, usable amino acid into the reaction.
Additives
- Nicotinamide — Precursor for $\text{NAD}^+/\text{NADH}$ regeneration via the salvage pathway; $\text{NAD}^+$ is essential for the GAPDH step of glycolysis, which is required for ATP regeneration from glucose during the long incubation.
Backfill
- Nuclease-Free Water — Brings the reaction to final volume while ensuring no contaminating RNases or DNases degrade the DNA template, mRNA, or tRNAs during the extended incubation.
Q2. Describe the main differences between the 1-hour PEP/NTP and 20-hour NMP-Ribose-Glucose master mixes (2–3 sentences).
The PEP/NTP system is engineered for speed: it directly supplies the four high-energy NTPs (ATP, GTP, CTP, UTP) plus phosphoenolpyruvate (PEP-Mono) and maltodextrin as fast-discharging energy donors, giving an immediate burst of transcription and translation that runs out within ~1 hour. The NMP-Ribose-Glucose system instead supplies cheap low-energy precursors (NMPs, ribose, glucose, guanine) and lets the lysate’s native metabolism — glycolysis fueled by phosphate buffer and $\text{NAD}^+$ regenerated from nicotinamide — slowly regenerate NTPs over ~20 hours, trading peak rate for sustained yield, lower cost, and avoidance of inhibitory byproducts like accumulated phosphate. As a result, the 1-hour mix also relies on extra small-molecule boosters (spermidine, DMSO, cAMP, NAD, folinic acid) to maximize a short burst, while the 20-hour mix’s design philosophy is metabolic self-sufficiency for long-running, sustainable protein production.
Q3. Identify and explain at least one biophysical or functional property of each of the six fluorescent proteins that affects cell-free expression or readout (1–2 sentences each).
1. sfGFP (superfolder GFP) — Engineered specifically for rapid, robust folding (maturation ~14 min) even when fused to misfolded partners, which makes it nearly ideal for CFPS where lysate chaperone capacity is limited. Like all Aequorea-lineage GFPs, however, chromophore maturation requires molecular oxygen, so sealed/anaerobic reaction wells will cap final fluorescence regardless of how much protein is translated.
2. mRFP1 — A first-generation monomeric DsRed derivative with slow, two-step oxygen-dependent maturation (on the order of ~1 hour or more), meaning a substantial fraction of translated mRFP1 in a short cell-free run will be present but non-fluorescent. It is also moderately acid-sensitive ($\text{p}K_a \approx 4.5$) and the dimmest of the six (low quantum yield ~0.25), so pH drift and incomplete maturation both suppress readout.
3. mKO2 — A monomeric coral (Fungia) FP with reasonably fast maturation (~30–60 min) and good photostability, but it is acid-sensitive ($\text{p}K_a \approx 5.5$): as glycolysis acidifies the CFPS reaction over 36 hours, mKO2 fluorescence is progressively quenched even if protein levels keep rising. Like all Anthozoa-derived FPs, its red-shifted chromophore requires a second oxidation step that consumes $\text{O}_2$.
4. mTurquoise2 — Aequorea-lineage cyan FP with the highest quantum yield among CFPs (~0.93), fast maturation, and excellent pH stability ($\text{p}K_a \approx 3.1$), so per-molecule readout is very strong and largely insensitive to reaction acidification. Folding is efficient in E. coli lysate, making it one of the most “forgiving” reporters for cell-free conditions.
5. mScarlet-I — A synthetic-template monomeric red FP whose “I” variant trades a small drop in quantum yield for dramatically faster maturation (~36 min vs. ~174 min for mScarlet), which is critical in CFPS where you want signal accumulation to track translation rather than lag behind it. It is still $\text{O}_2$-dependent (two-step Anthozoa-type chromophore) and benefits from sustained energy regeneration over long incubations.
6. Electra2 — A 2022 blue FP derived from mRuby3 (Anthozoa/eqFP611 lineage), engineered via dual bacterial+mammalian screening for high intracellular brightness and efficient folding in the E. coli cytoplasm — directly relevant to lysate-based CFPS. It inherits the two-step oxygen-dependent maturation of its Anthozoa parent, so $\text{O}_2$ availability and incubation time both gate final readout.
Q4. Create a hypothesis for how adjusting one or more reagents in the cell-free master mix could improve a specific biophysical or functional property identified above, over a 36-hour reaction.
Protein: mRFP1 Reagent change: Increase HEPES-KOH (pH 7.5) from 45 mM to ~80 mM (matching the 1-hr PEP/NTP mix), and slightly raise Magnesium Glutamate from 7.0 mM toward ~8–9 mM.
Rationale / expected effect: mRFP1’s two limiting properties in CFPS are slow oxygen-dependent maturation and moderate acid sensitivity. Over 36 hours, glycolysis of the supplied glucose/ribose accumulates pyruvate, lactate, and inorganic phosphate, dropping the reaction pH — which both quenches the existing mRFP1 chromophore (acid $\text{p}K_a \approx 4.5$) and slows the late oxidation step of chromophore maturation, which proceeds best near neutral pH. Raising HEPES nearly doubles buffering capacity so the reaction stays close to pH 7.5 deep into the incubation, preserving fluorescence of already-matured mRFP1 and giving the slow-maturing fraction the neutral-pH window it needs to finish oxidizing. The small $\text{Mg}{2+}$ bump compensates for additional $\text{Mg}{2+}$ chelation by the higher buffer/phosphate load and keeps ribosomes and NMP/NDP kinases active, so translation continues feeding new mRFP1 molecules into that maturation pipeline through the full 36 hours rather than stalling at hour ~10–15.
Proposed control: Test the elevated-HEPES condition against mTurquoise2 (pH-stable, fast-maturing) in parallel — if the hypothesis is correct, the buffer boost should help mRFP1 substantially more than mTurquoise2, isolating the pH/maturation effect from a generic translation-yield effect.