Week 2 HW: DNA Read, Write, & Edit
Gel Electrophoresis Designs
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks
I have created an image of mount fuji with clouds in the sky. I have inverted the image so it is easier to visualize.
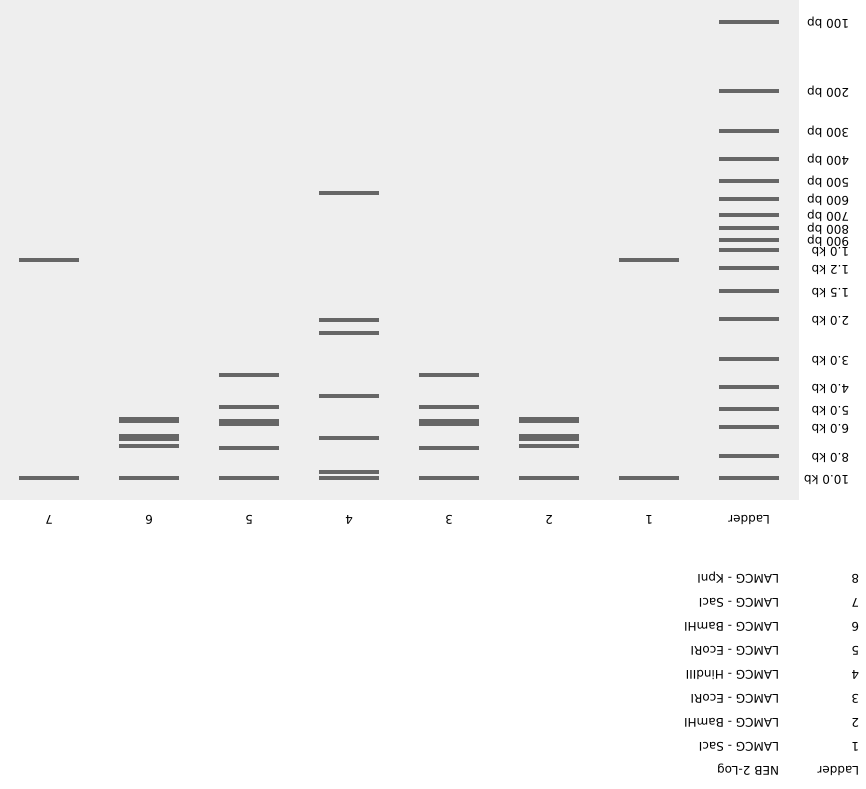

Note: Since we worked in groups during lab this week, we created a different design than the one shown above for the lab activity.
DNA Design Challenge
Choose your protein.
RES-701-3 is a tiny natural protein made by soil bacteria (Streptomyces). It belongs to a family called lasso peptides, named because their structure looks like a lasso or slipknot. The tail of the protein threads through a loop, creating a knot that is extremely hard to unravel.
This knotted shape makes lasso peptides unusually tough. They resist being broken down by digestive enzymes, heat, and harsh chemical environments. These are properties that most proteins lack, and that make them attractive as potential drugs.
RES-701-3 blocks a receptor on the surface of blood vessel cells called the endothelin type B receptor (ETB). The endothelin system controls blood vessel tightening and relaxation, and becomes dysregulated with age, contributing to high blood pressure and vascular disease. RES-701-3 acts as an inverse agonist, meaning it blocks the receptor and pushes toward a less active state than its resting baseline.
In nature, the bacteria makes this peptide in two parts:
- Leader section:
MSDITLTPMDLLDLDELAAGGGRSTARE - Core peptide sequence:
GNWHEPEIDGWNPHGW
The core is removed from the leader with an enzyme, which makes it active.
Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The nucleotide sequence of the leader and the core is shown respectively.
- Leader:
ATGAGCGATATTACCCTGACCCCGATGGATCTGCTGGATCTGGATGAACTGGCTGCTGGTGGTGGTCGTAGCACCGCTCGTGAA - Core:
GGTAACTGGCATGAACCGGAAATTGATGGTTGGAACCCGCATGGTTGGTAA
Codon optimization.
Due to evolution, different species have different codons it uses frequently and has abundant matching transfer RNAs for, and codons it rarely uses and has few tRNAs for. RES-701-3 comes from Streptomyces and strongly prefers codons loaded with G and C. Twist has a Streptomyces coelicolor for codon optimization.
However, it’s worth mentioning that in a 2025 paper by Shihoya et al. paper, they used Streptomyces venezuelae as organism and achieved the highest reported yields. If I was in a real drug development setting, I might go with this.
Here is the codon optimized variant for both leader and core together:
You have a sequence! Now what?
I have listed the Promoter, RBS, Start Codon, Coding Sequence, His Tag, Stop Codon, Terminator as well as the reagents needed below.
Promoter: The **ermE*p promoter is supposed to be the most widely used for gene expression in Streptomyces.
Ribosome Binding Site: We’re using Shine Dalgarno (SD) sequence, AAGGAG, which is supposed to be a good RBS for streptomyces with leaders. It is supposed to be positioned 6 to 10 nucleotides upstream of the start codon, so we will use 7 nucleotides. We’re going to put two spacers before and after the SD sequence, CGACG and ACAC.
Start Codon: This is just going to be the usual ATG.
Coding Sequence: We are going to put both of our leader and core peptide sequence together here.
His tag: This is a short string of six histidine amino acids added to the protein so you can fish it out of a mixture using a nickel column. The histidines stick to nickel, letting you pull your protein out of everything else the cell makes. However, in practice, apparently this is not actually good to put on for RES-701-3 because it would interfere with binding the ETB receptor.
Stop Codon: TGA tells the ribosome to stop building the protein here. TGA is the preferred stop codon in Streptomyces because it is relatively speaking, GC-rich, matching the organism’s DNA preferences as discussed before. For example, typical stop codon is TAA.
Terminator: Tells the cell’s RNA-copying machinery to stop making mRNA. Without it, the cell would keep reading past your gene into random neighboring DNA. We’re using the fd terminator from a bacteriophage which is commonly used in Streptomyces expression vectors.
Reagents
In order to produce these proteins we also need to use some enzymes to be used as reagents, namely, LasB1, LasB2 and LasC. For this lasso peptide, LasB1 binds the leader, delivers the whole precursor to LasB2 which cuts the leader off, and then LasC closes the ring on the core. It doesn’t seem easy to order the reagents so it seems like this peptide wouldn’t be a great choice for the class. In addition, the yield is optimized by using Streptomyces venezuelae, which is also not too common.
Prepare a Twist DNA Synthesis Order
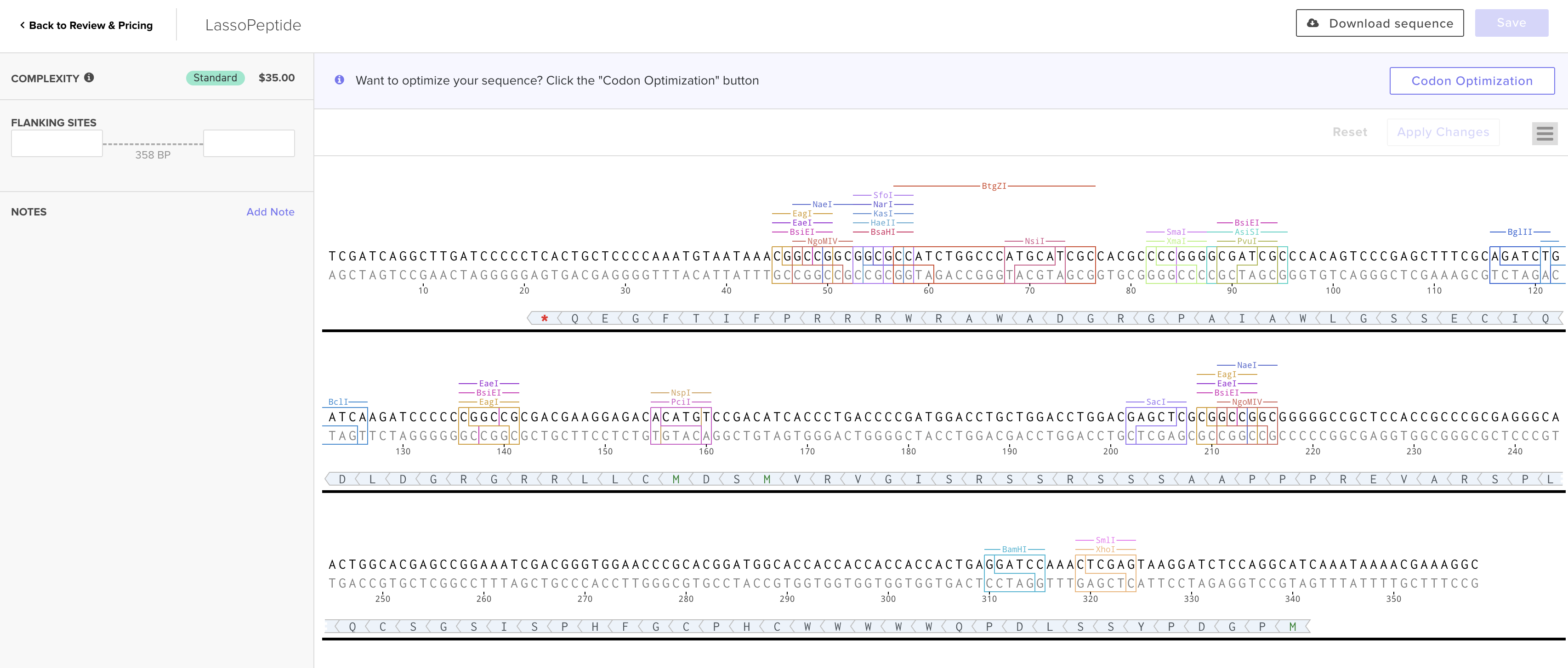
I prepared the lasso peptide order. Here is a picture of the expression cassette below in benchling.
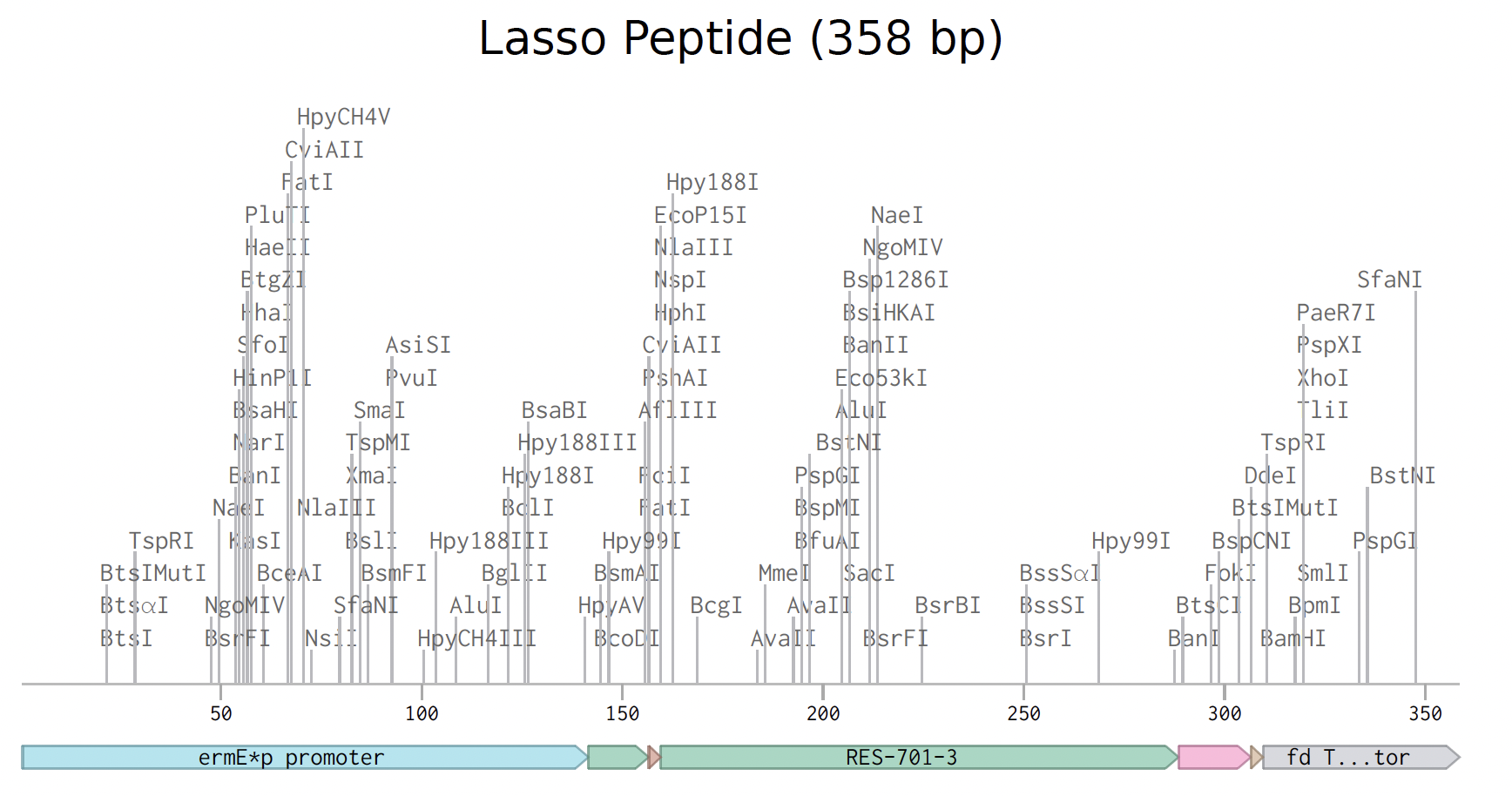

Instead of a clonal gene, I used gene fragments because they work better Streptomyces as an organism rather than e coli, which are the standard cloning vectors.

DNA Read/Write/Edit
5.1 DNA Read
What DNA would you want to sequence (e.g., read) and why?
I would want to sequence the whole genomes of all ~6,000 mammalian species. The largest current collection of mammalian genomes is the Zoonomia project, which contains around 250 whole genomes along with known maximum lifespan data for most of these species. However, expanding this to cover all mammals—paired with their maximum lifespan records—would allow us to train computational models that identify DNA patterns predicting how long a species can live. In short, more genomes means better predictions about which parts of DNA are linked to longevity.
In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Illumina short-read sequencing (second-generation): This produces highly accurate short reads (~150–300 base pairs) and is great for spotting small genetic differences between species.
Is your method first-, second-, or third-generation?
I am using both second-generation Illumina. First-generation refers to older Sanger sequencing, which reads one fragment at a time and is too slow and expensive for whole genomes. Second-generation sequences millions of short fragments in parallel, making it fast and cheap.
What is your input? How do you prepare your input?
The input is genomic DNA extracted from tissue or blood samples of each mammalian species. The essential preparation steps are:
- DNA extraction: Isolate high-quality DNA from the biological sample.
- Fragmentation: Break the DNA into smaller pieces.
- Adapter ligation: Attach short known DNA sequences adapters to the ends of each fragment so the sequencing machine can recognize and handle them.
- PCR amplification (Illumina): Make many copies of each fragment to boost the signal.
- Quality check: Verify the library is the right size and concentration before loading it onto the sequencer.
What are the essential steps of your chosen sequencing technology? How does it decode bases (base calling)?
Fragmented DNA is attached to a glass surface flow cell, amplified into clusters, and then sequenced one base at a time. In each cycle, a fluorescently labeled nucleotide is added, a camera captures which color lights up at each cluster where each of the four bases has a different color, and the machine records the base. This process repeats hundreds of times to read out each fragment.
What is the output?
The output is digital sequence files, typically in FASTQ format, containing millions of reads—short or long strings of A, T, C, and G letters—along with quality scores indicating how confident the machine is about each base call. These reads are then assembled and aligned computationally to reconstruct each species’ complete genome.
5.2 DNA Write
What DNA would you want to synthesize (e.g., write) and why?
Based on the sequencing data above, I would use trained computational models to predict specific DNA sequences associated with high maximum lifespan. I would then synthesize these predicted longevity-linked sequences—for example, specific gene variants or regulatory elements found in long-lived species like bowhead whales or naked mole-rats—so they can be tested in cell cultures or animal models. The goal is to move from computational prediction to experimental validation: do these DNA sequences actually promote cellular health and longevity?
What technology or technologies would you use to perform this DNA synthesis and why?
- Oligonucleotide synthesis from Twist Bioscience: For building short to medium DNA fragments (up to a few thousand base pairs). These companies use chemical synthesis on microchips to build many sequences in parallel, making it fast and affordable.
- Gibson Assembly or Golden Gate Assembly: For stitching shorter synthesized fragments together into larger constructs. These are molecular cloning methods that use enzymes to join DNA pieces seamlessly.
What are the essential steps of your chosen synthesis method?
- Sequence design: Use computational models to design the target DNA sequences, optimizing codon usage for the target organism and avoiding problematic features (e.g., long repeats, extreme GC content).
- Oligonucleotide synthesis: Short single-stranded DNA pieces (oligos, ~50–200 bases) are built base by base using chemical reactions on a solid support. Each cycle adds one nucleotide at a time.
- Assembly: Overlapping oligos are combined and joined enzymatically into longer double-stranded fragments (a few hundred to a few thousand base pairs).
- Cloning: The assembled fragments are inserted into a circular DNA carrier (plasmid vector) and introduced into bacteria, which copy the DNA as they grow.
- Verification: The final constructs are sequenced to confirm the correct sequence was built.
- Large construct assembly: Multiple verified fragments are stitched together using Gibson Assembly or Golden Gate Assembly to create larger genetic constructs.
What are the limitations of your synthesis method in terms of speed, accuracy, and scalability?
- Speed: Synthesizing and assembling long constructs (>10,000 base pairs) can take weeks, since each fragment must be built, verified, and then joined together step by step.
- Accuracy: Chemical synthesis introduces errors at a rate of roughly 1 in 200 bases per oligo. While these errors are corrected through screening and verification, it adds time and cost.
- Scalability: Very long or repetitive sequences are difficult to synthesize because the oligos may misassemble or fold in unwanted ways. Sequences with extreme GC content are also harder to build reliably.
5.3 DNA Edit
What DNA would you want to edit and why?
I would want to edit specific genes in model organisms (such as mice) to replace their native sequences with the longevity-associated sequences identified from the analysis above. For example, if the computational model predicts that a certain variant of a DNA repair gene is linked to longer lifespan in mammals, I would edit a mouse’s genome to carry that variant. This would let us test whether swapping in these predicted “long-life” DNA variants actually extends lifespan or improves age-related health outcomes like cancer resistance or cellular repair.
What technology or technologies would you use to perform these DNA edits and why?
I would use CRISPR-Cas9 gene editing, because it is the most precise, versatile, and widely used genome editing tool available. It can make targeted changes at specific locations in the genome of living cells and organisms, and it works well in mammalian systems including mice.
How does your technology edit DNA? What are the essential steps?
- Target selection: Identify the exact location in the genome you want to edit.
- Guide RNA design: Design a short RNA sequence that matches the target DNA site.
- Cutting: The Cas9 protein, guided by the RNA, binds to the matching DNA site and makes a double-strand break.
- Repair: The cell’s natural repair machinery fixes the break. If a DNA template with the desired new sequence is provided alongside the CRISPR components, the cell can use it as a blueprint to incorporate the new sequence, called homology-directed repair.
- Screening: Edited cells are sequenced to confirm the desired change was made correctly.
What preparation do you need to do, and what is the input?
- Design inputs: The target DNA sequence, a custom guide RNA matching that sequence, and a DNA donor template carrying the desired new sequence flanked by regions that match the area around the cut site.
- Molecular inputs: Cas9 protein or mRNA, synthesized guide RNA, donor template DNA, and delivery reagents.
- Biological inputs: Target mouse cell.
What are the limitations of your editing method in terms of efficiency or precision?
- Off-target edits: The guide RNA can sometimes bind to similar sites elsewhere in the genome, causing unintended cuts and mutations.
- Low HDR efficiency: Only a fraction of edited cells may carry the precise desired change, requiring extensive screening.
- Delivery challenges: Getting CRISPR components into every target cell efficiently, especially in living animals, remains difficult. Some tissues are harder to reach than others.