Week 4 HW: Protein Design
Part A: Conceptual Questions
Why do beta-sheets tend to aggregate?
A beta-strand forms when a protein’s backbone — the repeating NH–Cα–CO chain shared by every amino acid — stretches out into a nearly flat zigzag. When two or more of these strands line up next to each other and link through hydrogen bonds (where an N–H on one strand bonds to a C=O on the neighbor), you get a beta-sheet.
The strands on the outer edges still have a full row of exposed N–H and C=O groups, allowing another strand to be added, and so on — this is why beta-sheets tend to aggregate.
What forces pull sheets together?
Hydrophobic effect — the biggest driver. In a beta-strand, side chains stick out alternately above and below the sheet. Since many side chains are hydrophobic, two sheets stack such that the greasy surfaces face inward.
Hydrogen bonding — gives the structure its regularity. Each new strand that joins the sheet edge contributes roughly one H-bond per amino acid along its length. Individually, H-bonds in water are not enormously strong (breaking one with a neighbor just lets you form one with water instead), but across a strand of ten or more residues they add up meaningfully.
Van der Waals packing — stabilizes sheets that have stacked together. These forces are much weaker and shorter-range, arising from temporary, fluctuating dipoles.
Part B: Protein Analysis and Design
Briefly describe the protein you selected and why you selected it.
I selected a monoclonal antibody for the following reasons:
- Ability to target specific proteins on cell surfaces with extreme precision, directly applicable to therapeutics
- Ability to recruit the immune system (via Fc region) to destroy tagged cells, combining specificity with immune effector functions
- Can be engineered with ML and computational methods for improved binding affinity and reduced immunogenicity
- Compared to small molecule drugs, highly specific to their target with fewer off-target effects
For this exercise, I chose trastuzumab, famous for revolutionizing the treatment of HER2-positive breast cancer. It is a humanized IgG1 monoclonal antibody that binds to the extracellular domain IV of HER2 (human epidermal growth factor receptor 2), blocking receptor dimerization and downstream signaling that drives tumor growth.
How long is it? What is the most frequent amino acid?
The full trastuzumab IgG has 2 heavy chains (449 aa each) and 2 light chains (214 aa each), for a total of ~1,326 amino acids and ~148 kDa.
However, the crystal structure (PDB: 1N8Z) contains only the Fab fragment (the antigen-binding portion), which includes:
Heavy chain Fab (chain B, 220 aa):
Light chain (chain A, 214 aa):
- Combined Fab length: 434 amino acids
- Most common amino acid: S (Serine), appearing 60 times
How many protein sequence homologs are there for your protein?
Because trastuzumab is a humanized antibody with conserved IgG1 framework regions, BLAST returns a very large number of homologs — antibodies share ~70–90% identity in their framework regions. A BLAST search of the heavy chain Fab against UniProt returns over 250 homologs. The variable CDR (complementarity-determining region) loops are what make trastuzumab unique in its HER2 specificity.
When was the structure solved? Is it a good quality structure?
Good quality = good resolution. Smaller is better (benchmark: 2.70 Å).
- Deposited: 2002-11-21
- Released: 2003-02-18
- Published: Cho et al., Nature (2003) 421: 756–760
- Link: https://www.rcsb.org/structure/1N8Z
- Resolution: 2.52 Å — good quality, better than the 2.70 Å benchmark
Are there any other molecules in the solved structure apart from protein?
Yes. In addition to the 3 unique protein chains (light chain A, heavy chain B, and HER2 extracellular domain C), the structure contains:
| Molecule | Description | Copies |
|---|---|---|
| NAG | 2-acetamido-2-deoxy-β-D-glucopyranose (N-linked glycosylation sugar attached to HER2) | 2 |
| SO4 | Sulfate ion | 1 |
Does your protein belong to any structure classification family?
Yes. The overall complex is classified in the PDB under TRANSFERASE. The trastuzumab Fab itself belongs to the Immunoglobulin superfamily.
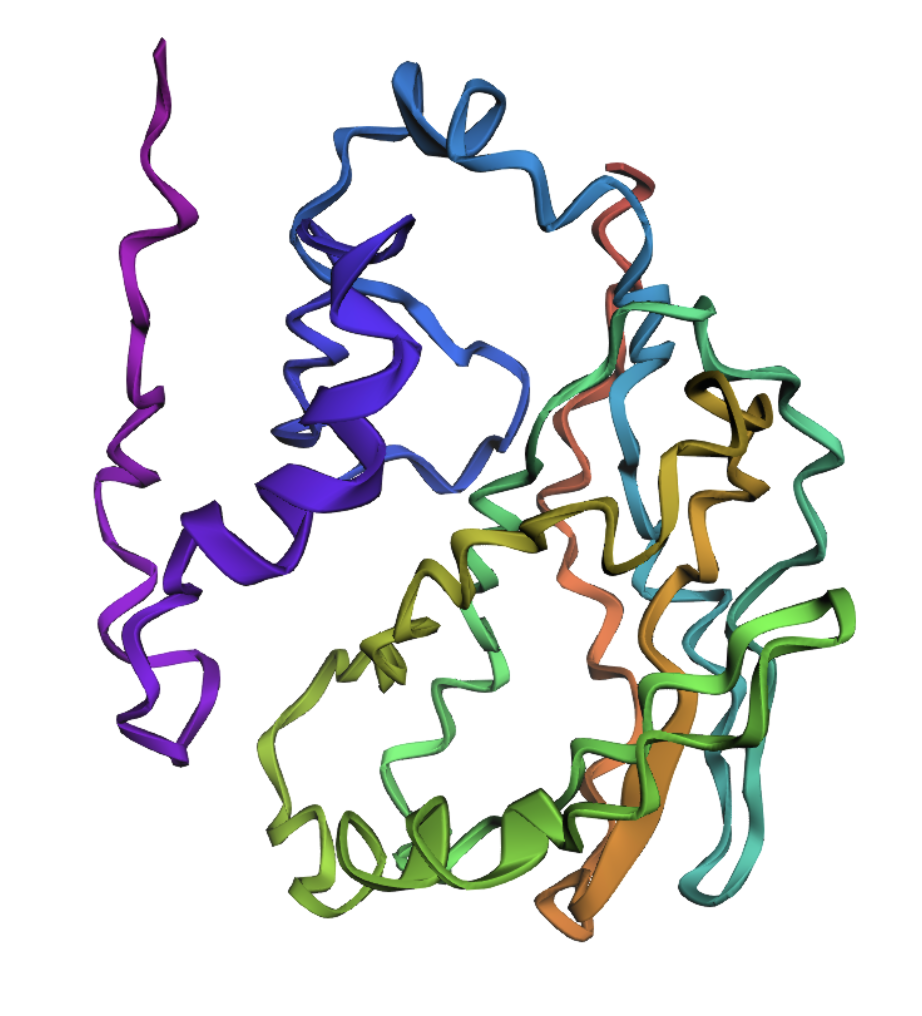
Visualize the protein as “cartoon”, “ribbon” and “ball and stick”
Cartoon
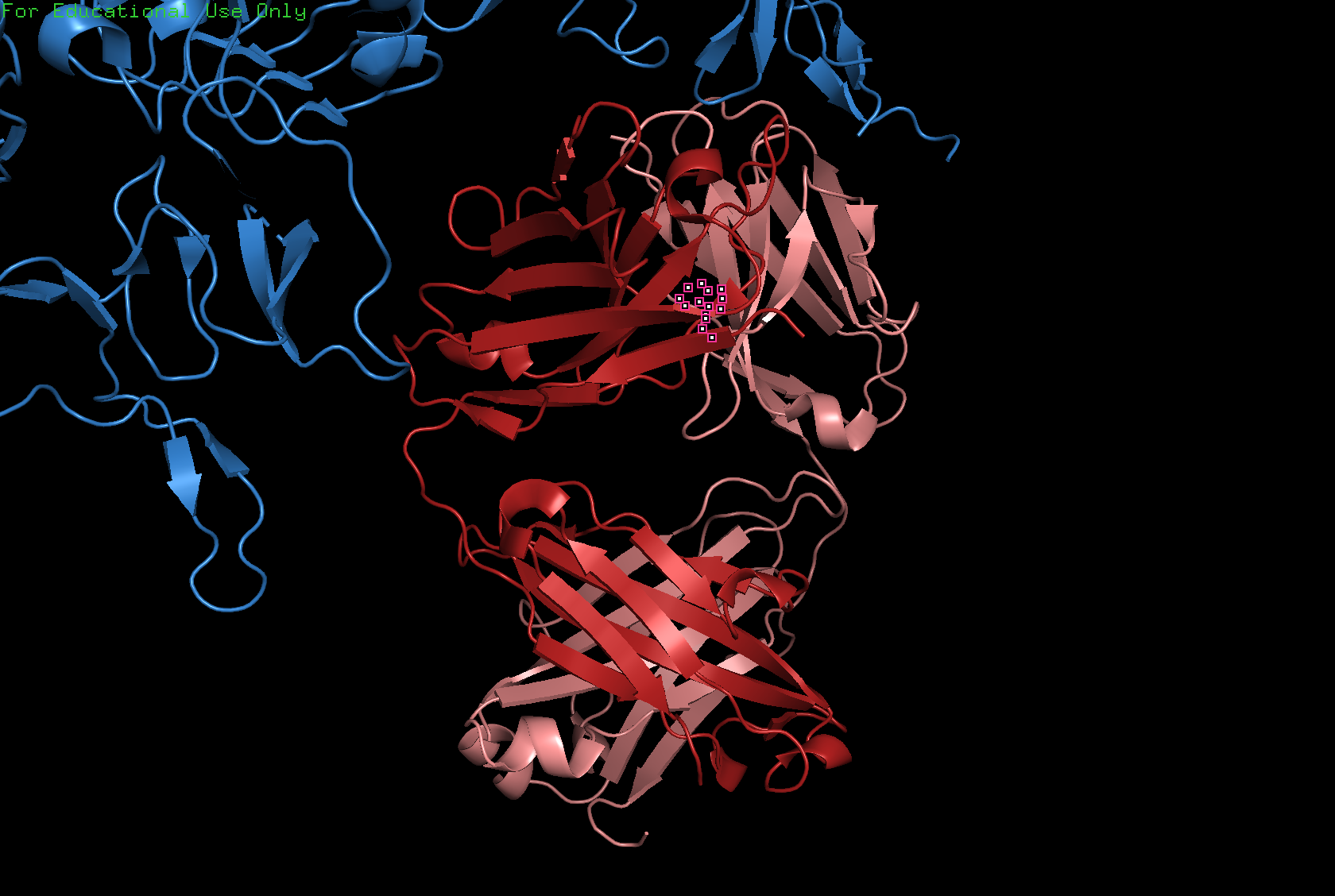

Ribbon
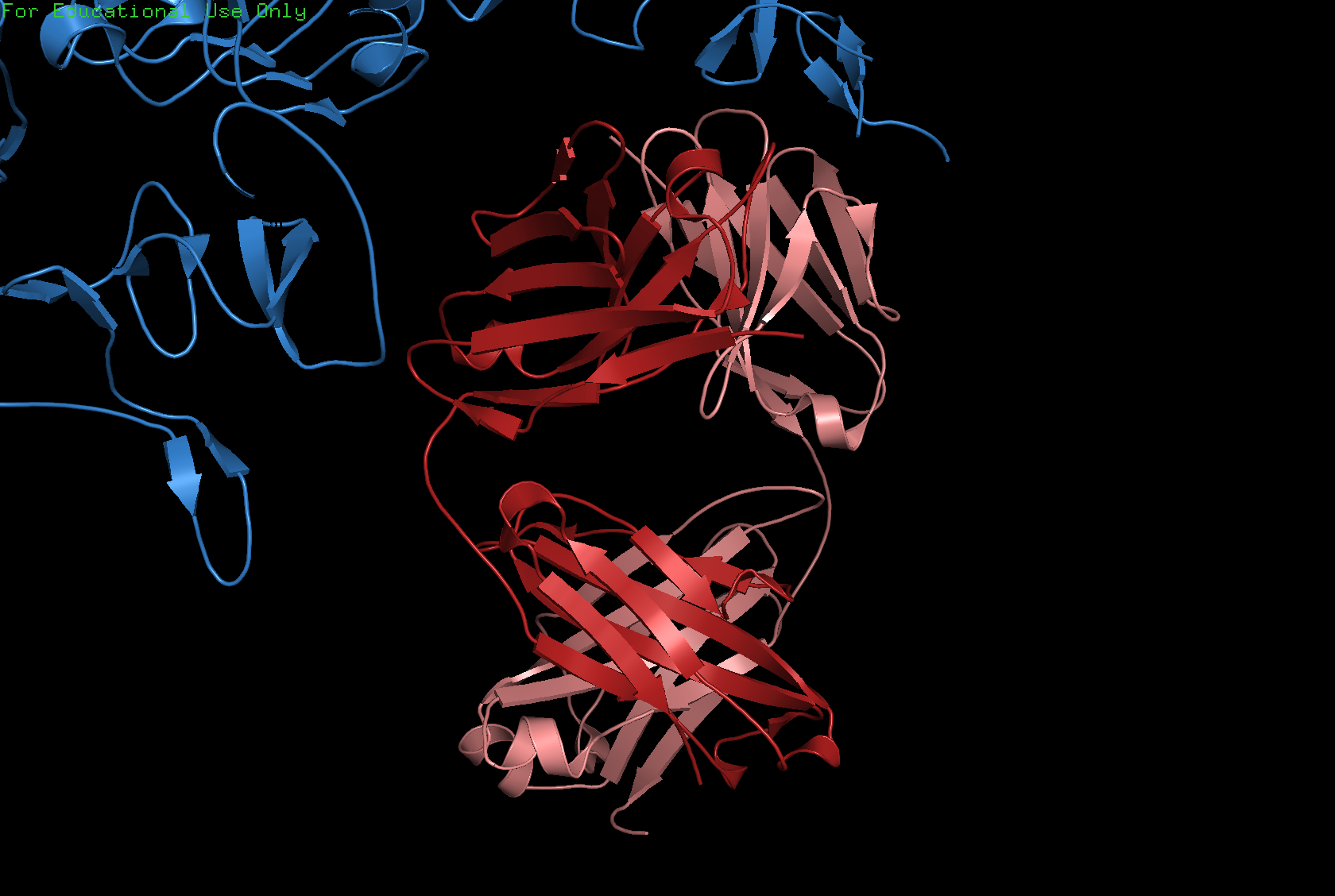

Ball and Stick
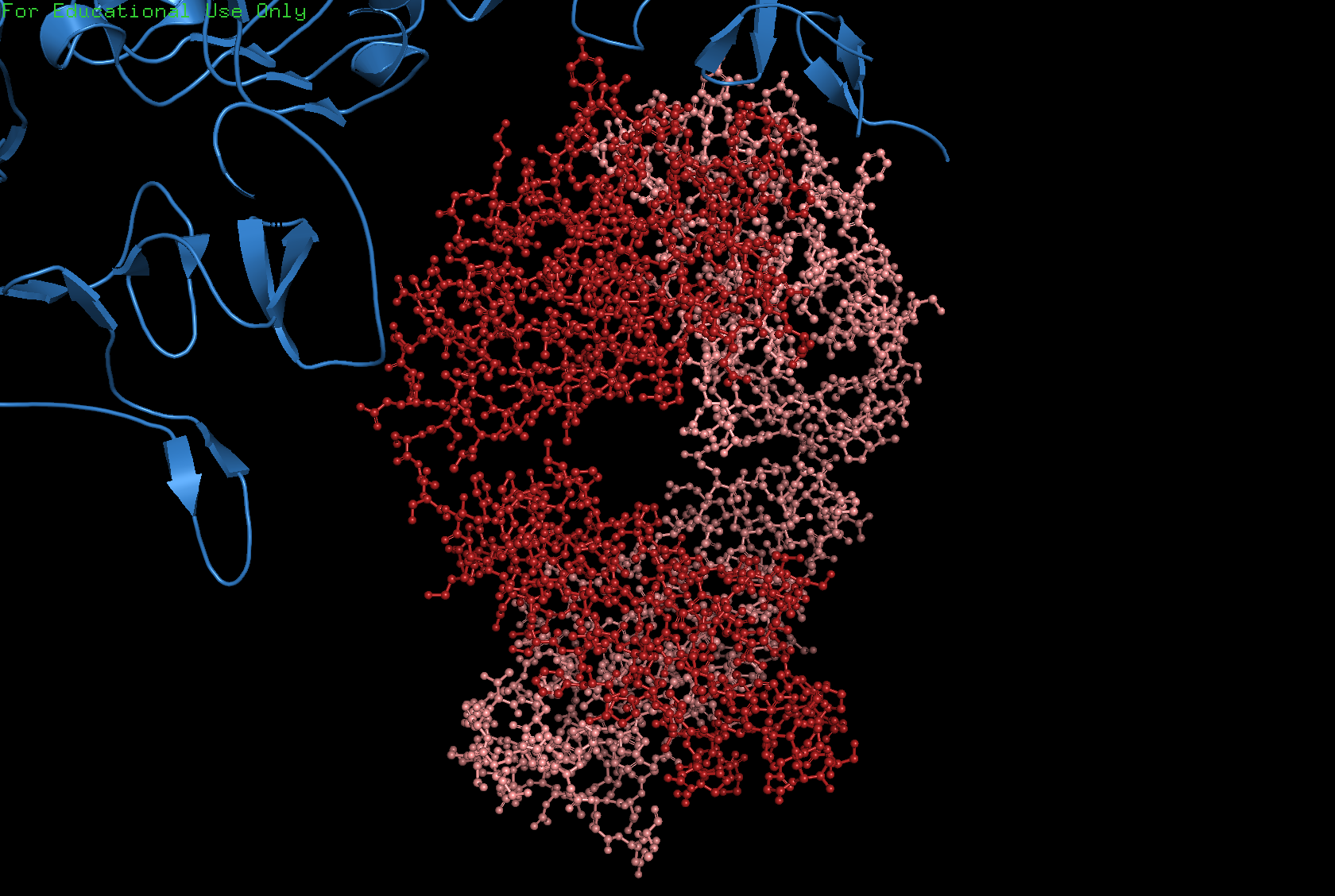

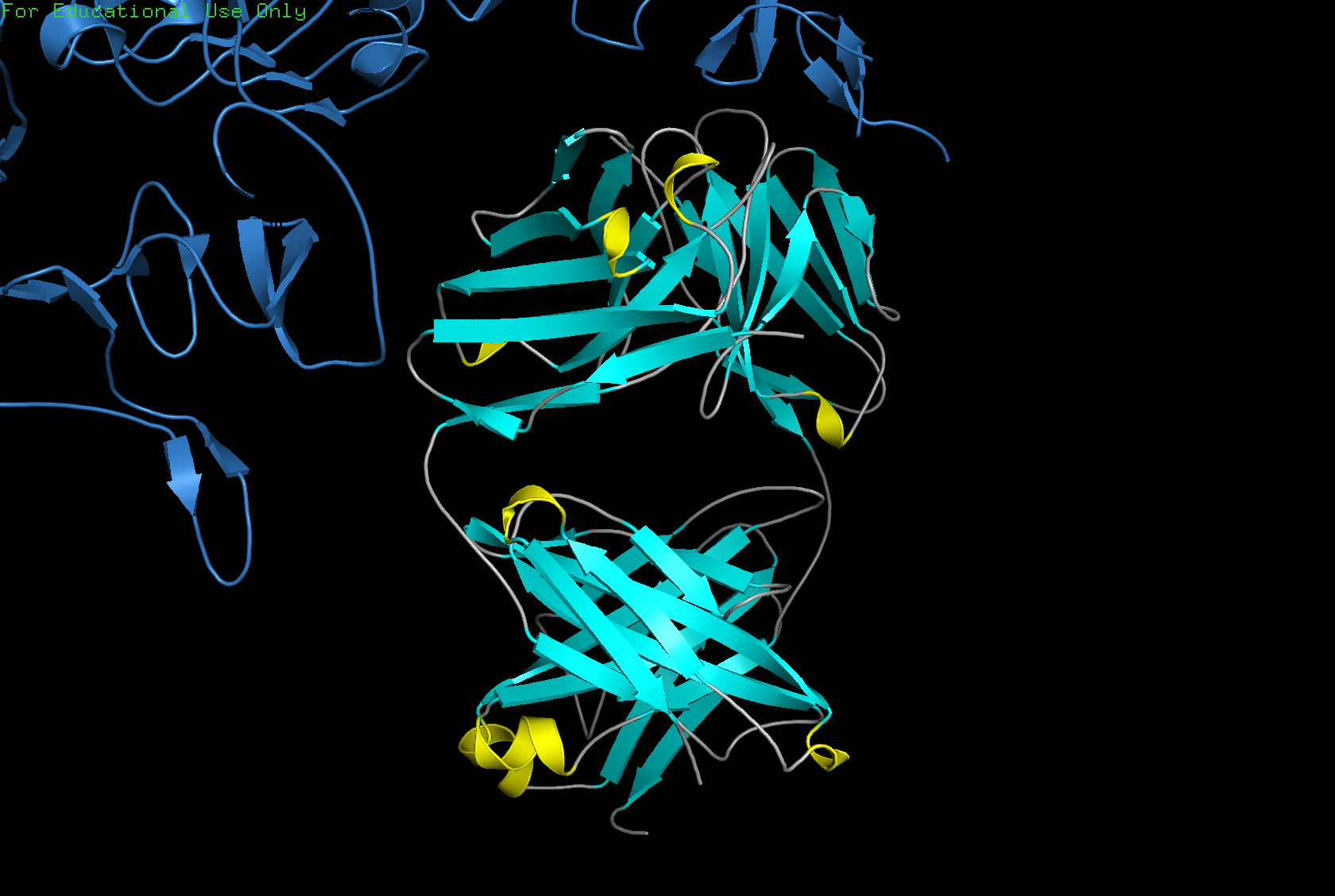
Color the protein by secondary structure. Does it have more helices or sheets?
The structure has more sheets than helices — specifically 215 atoms in sheets vs 30 atoms in helices.

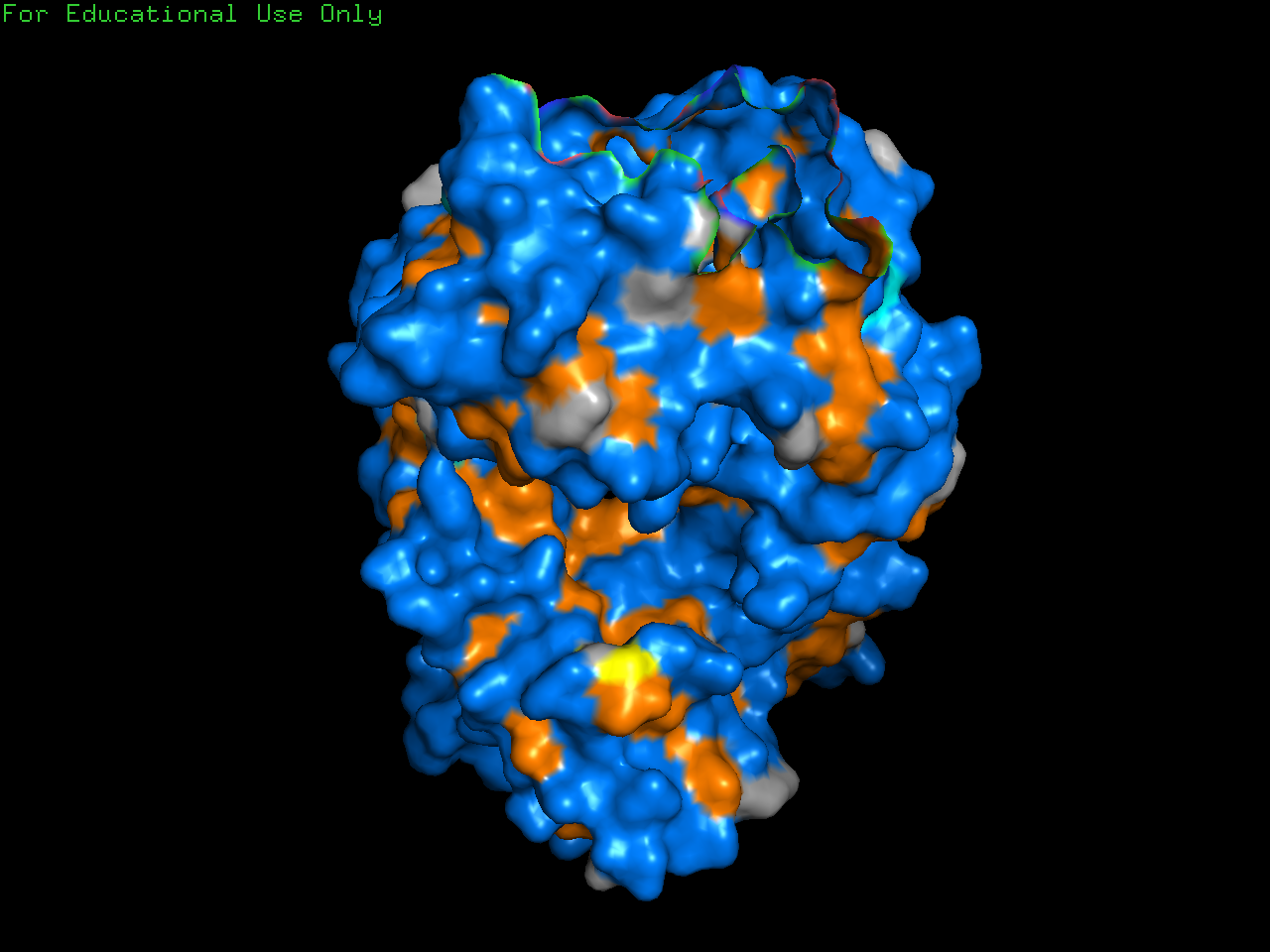
Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues?
Generally, proteins have a hydrophobic core with a hydrophilic surface, and trastuzumab follows this pattern. The immunoglobulin fold is a beta sandwich where:
- Hydrophobic residues (orange) point inward
- Hydrophilic residues (blue) point outward
(This is hard to see in the visualization because the inward and outward surfaces are not so distinct.)
However, the CDR loops — the tips that contact the target HER2 — are mixed: aromatic hydrophobics (Trp, Tyr) provide shape complementarity, while polar and charged residues form hydrogen bonds and salt bridges with the antigen.

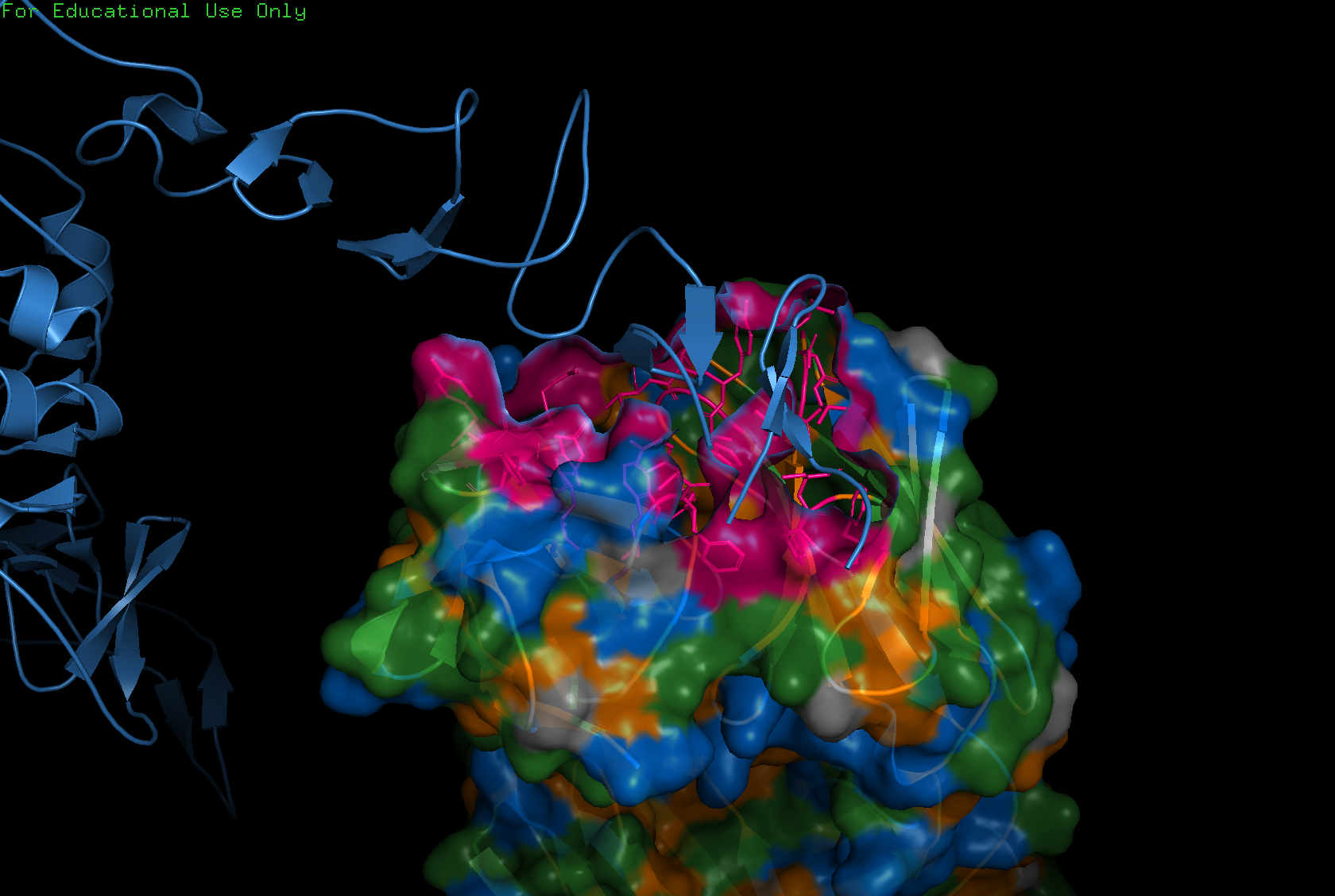
Visualize the surface of the protein. Does it have any “holes” (aka binding pockets)?
Yes, binding pockets are visible on the surface.

Part C: ML-Based Protein Design Tools
For this exercise, I chose the 6M0J SARS-CoV-2 Spike Receptor Binding Domain.
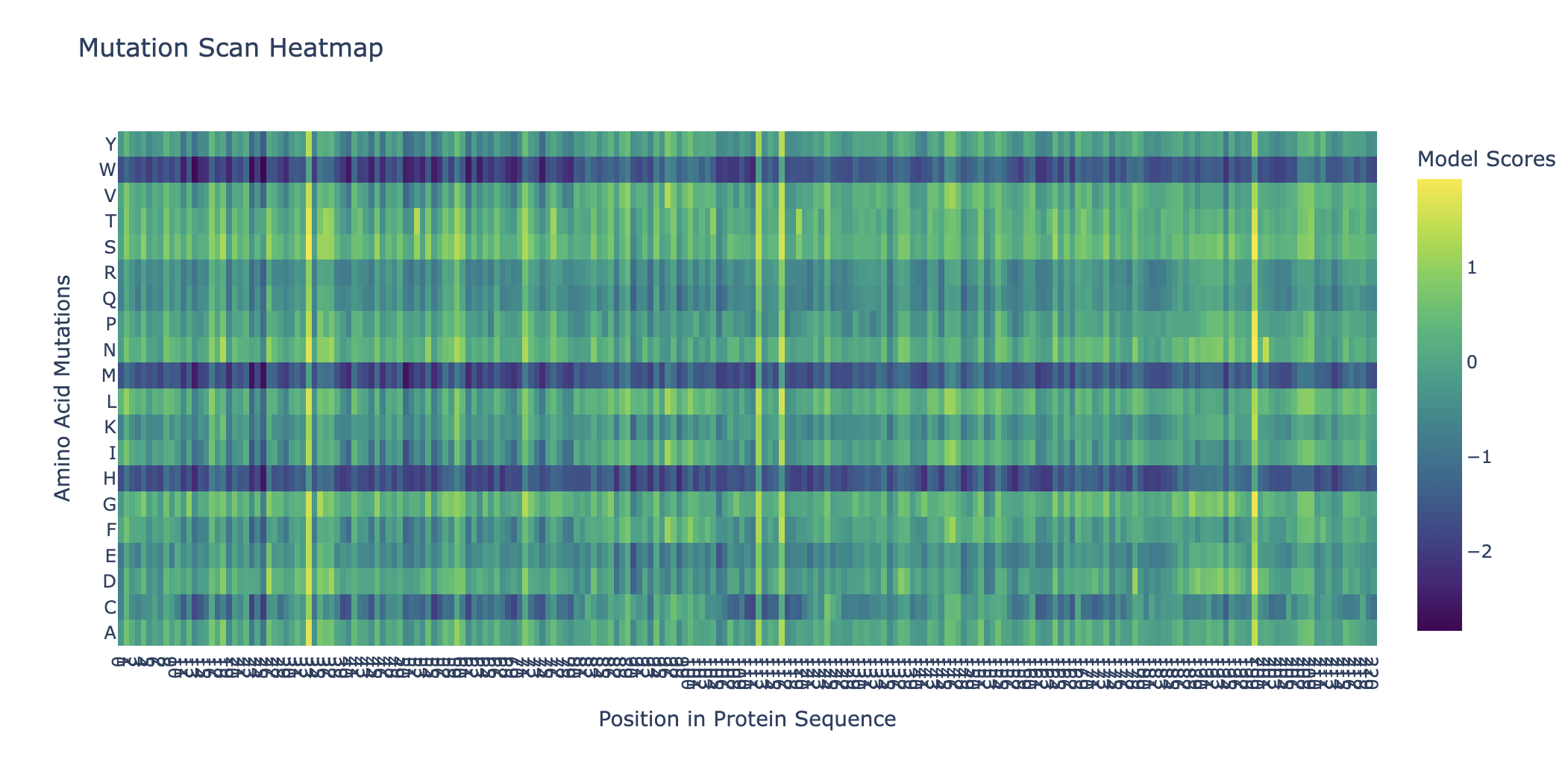
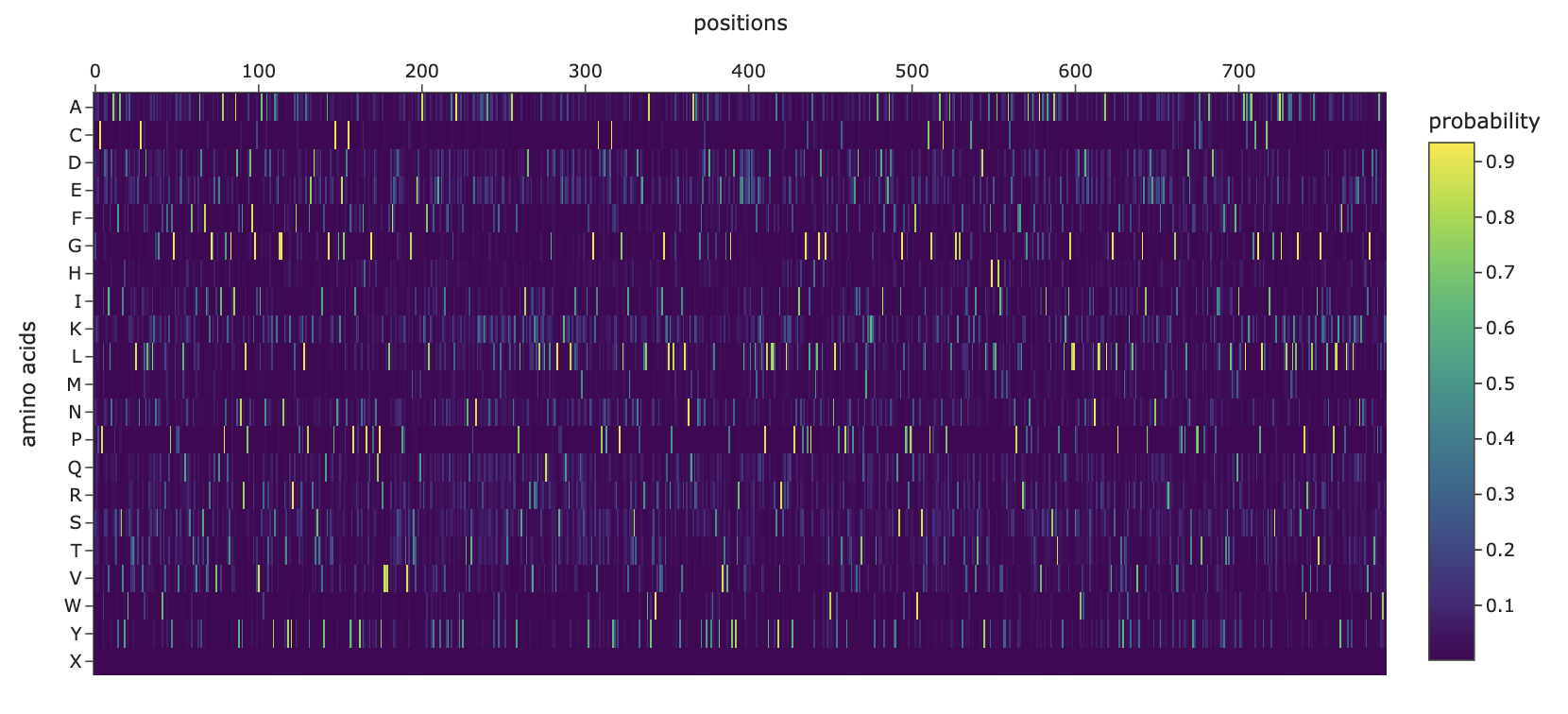
Deep Mutational Scans
Can you explain any particular pattern?
Horizontal patterns (rows): The rows for tryptophan (W), histidine (H), and methionine (M) are consistently darker across nearly all positions. These are large, bulky, or chemically complex amino acids that are difficult to accommodate at arbitrary positions without disrupting the protein’s fold. In contrast, small, simple amino acids like alanine or serine are more easily tolerated as substitutions, which is why their rows appear lighter overall.
Vertical patterns (columns): The most striking pattern is the dark purple vertical stripes at specific positions. These correspond to cysteine residues, which form disulfide bonds that hold the shape together so it can bind the human ACE2 receptor. Because ESM2 learned from millions of protein sequences that these cysteines are almost never substituted in nature, it heavily penalizes any mutation at those positions. The darkest scores appear when cysteine is mutated to something like tryptophan or proline, since these would not only break the disulfide bond but also create additional structural problems.

Latent Space Analysis
Analyze the different formed neighborhoods: do they approximate similar proteins?
Generally, the proteins are clustered tightly. There are some distinct clusters on the edges, which likely share a common evolutionary ancestor.
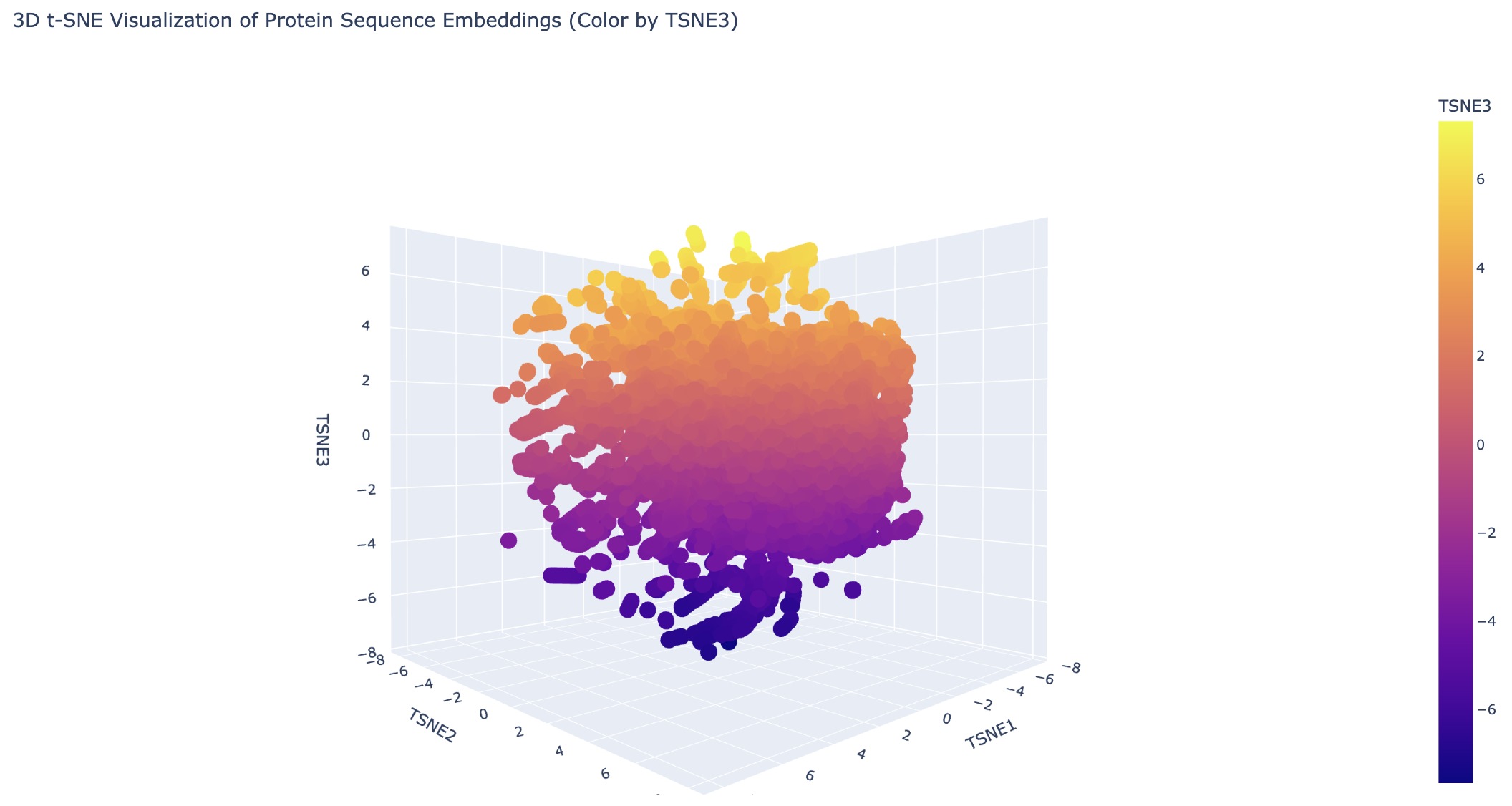

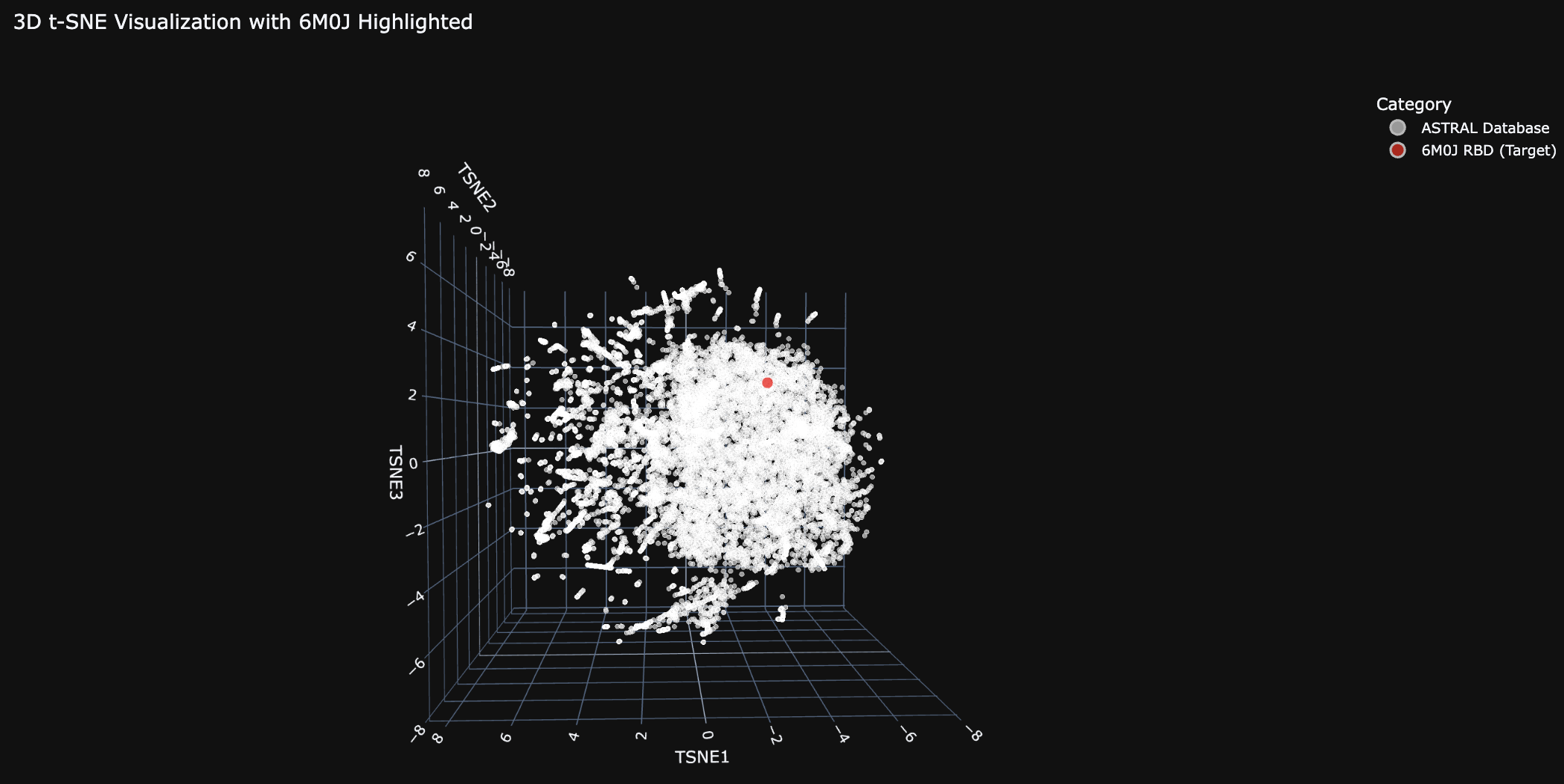
Place your protein in the resulting map and explain its position and similarity to its neighbors.
The 6M0J protein falls within the main cluster.

Folding a Protein
Fold your protein with ESMFold. Do the predicted coordinates match your original structure?
| Metric | Score | Interpretation |
|---|---|---|
| pLDDT (local confidence, 0–100) | 25.516 | Low — local structure unlikely to match the true structure |
| pTM (global fold confidence, 0–1) | 0.129 | Low — global topology prediction unreliable |
This is likely because the 6M0J viral protein is normally attached to a massive Spike protein complex. The SARS-CoV-2 Spike RBD is by itself very unstable.

Try changing the sequence. Is your protein structure resilient to mutations?
The original protein is not very resilient, given its poor pLDDT and pTM scores. However, after refolding with ProteinMPNN, the structure became much more stable:
| Metric | Original | After ProteinMPNN |
|---|---|---|
| pLDDT | 25.516 | 92.095 |
| pTM | 0.129 | 0.881 |
Note: while the structural metrics improved dramatically, this could result in a functionally incorrect protein — stability does not guarantee biological activity.
Inverse Folding
Analyze the predicted sequence probabilities and compare the predicted sequence vs the original one.
Roughly half of the original amino acids were preserved. This is a typical result for ProteinMPNN, as it seeks to optimize the sequence for the given backbone rather than simply mimicking the native sequence.
| Metric | Original | ProteinMPNN |
|---|---|---|
| Energy score | 1.3747 | 0.8107 |
In the context of ProteinMPNN, a lower score suggests that the new sequence is potentially more stable or fits the target backbone more optimally. This matches the pLDDT and pTM improvements noted above.

Input this sequence into ESMFold and compare the predicted structure to your original.
As noted above, the predicted structure after ProteinMPNN had higher pLDDT and pTM compared to the original.
Bacteriophage Engineering
For this exercise, I worked with Alayah Hines and Terry Luo.
Computational Engineering of the MS2 Lysis Protein (L)
The MS2 L protein is a 75-amino-acid polypeptide that lyses E. coli by an incompletely understood mechanism. Its C-terminal transmembrane (TM) domain inserts into the cytoplasmic membrane and oligomerizes, causing depolarization that triggers host autolytic enzymes to degrade the murein layer. Recessive, conservative missense mutations clustered around a conserved LS dipeptide strongly imply L engages an unidentified host protein target rather than simply disrupting the bilayer. The dispensable N-terminal domain binds chaperone DnaJ (with solved PDB structures), modulating lysis timing — its removal causes lysis ~20 min earlier. No experimental structure of L exists.
Goals:
- Stabilize L for more robust membrane accumulation
- Accelerate lysis by bypassing DnaJ-dependent regulatory timing and improving delivery of functional L to the membrane
Because the downstream lytic target is unknown, we do not attempt to enhance per-molecule toxicity at the point of target engagement; we focus on removing regulatory brakes and increasing the supply of functional protein.
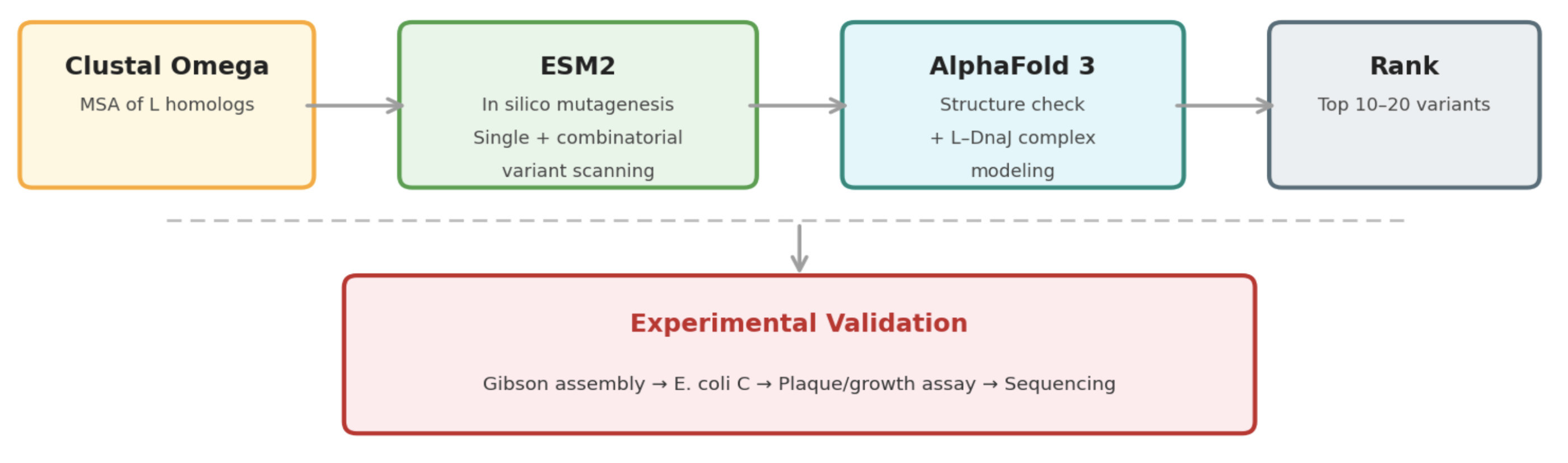
Pipeline: Three Tools, Each Non-Redundant
Clustal Omega (Conservation Map). Align L homologs across Leviviridae (MS2, f2, R17, GA, PP7, AP205, PRR1, M12, KU1, JP34). Conserved C-terminal residues — especially the LS motif — are presumed to mediate the unknown heterotypic interaction and are excluded from mutation. This map constrains all downstream design.
ESM2 + Deep Combinatorial Scanning (Fitness Oracle). Score every single-point mutation by log-likelihood change: increases at mutable positions indicate stabilizing substitutions (Goal 1). N-terminal scanning identifies mutations that disrupt DnaJ binding (Goal 2). A strict preservation rule applies near the LS motif: mutations are evaluated for maintenance of wild-type fitness, not improvement. The genetics show even conservative changes there cause recessive loss of function. Pairwise combinatorial scanning (~2M pairs) captures epistatic synergies at mutable positions.
AlphaFold 3 (Structural Filter + Complex Model). Predicts variant structures as a sanity check (does the TM helix survive?) and models the L–DnaJ complex to verify that N-terminal truncations/mutations disrupt the regulatory interface. Used as a filter, not a design engine. PAE matrix identifies confident interface contacts.
Ranking
Composite score: ESM2 log-likelihood gain (stability) + conservation preservation (all essential residues intact) + AF3-predicted DnaJ-binding disruption (for timing bypass). Top 10–20 variants advance to experimental validation.
Why Not More Tools?
ProteinMPNN is excluded because it is trained on crystallized globular PDB proteins, not predicted structures of disordered membrane peptides. The compute is instead invested in combinatorial ESM2 depth.
Pitfalls
- No experimental structure: All structural reasoning rests on AF3 predictions for a challenging target. Mitigated by treating AF3 as a filter and cross-referencing against the conservation map.
- Unknown lytic target: The central limitation. We cannot optimize target-binding affinity for an unidentified partner; engineering is restricted to upstream properties (stability, membrane delivery, DnaJ bypass).
- Autolysin bottleneck: If lysis rate is limited by host autolytic enzyme activity rather than L accumulation, stabilization gains may show diminishing returns; the plaque assay will reveal this.
Pipeline Schematic