Week 6 HW: Genetic Circuits Part 1
What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Phusion HF PCR Master Mix is a pre-made 2X formulation that contains several key components:
Phusion DNA Polymerase is a high-fidelity, thermostable polymerase fused to a processivity-enhancing domain. It has an error rate roughly 50-fold lower than Taq polymerase, which is critical when you need accurate amplification — as in this mutagenesis lab where only specific, intentional mismatches should be introduced.
dNTPs (deoxyribonucleotide triphosphates — dATP, dTTP, dGTP, dCTP) are the raw building blocks the polymerase uses to synthesize new DNA strands.
MgCl&sub2; provides magnesium ions, which are an essential cofactor for polymerase activity and also influence primer annealing stringency.
HF Buffer maintains optimal pH and salt conditions for the enzyme. The “HF” designation indicates it’s optimized for high-fidelity amplification across a broad range of templates. Some formulations also include detergents or stabilizers that help the enzyme tolerate common inhibitors.
The master mix format is convenient because it reduces pipetting steps and the chance of contamination — you only need to add template, primers, and water.
What are some factors that determine primer annealing temperature during PCR?
The annealing temperature is typically set 2–5°C below the lower melting temperature (Tm) of the two primers in a pair. Several factors determine what that optimal temperature is:
Primer length: Longer primers generally have higher Tm values because more hydrogen bonds stabilize the duplex.
GC content: G-C base pairs form three hydrogen bonds versus two for A-T pairs, so primers with higher GC content (ideally 40–60%) have higher Tm.
Salt/cation concentration: Mg²+ and monovalent cations in the buffer stabilize DNA duplexes; higher concentrations raise the effective Tm.
Mismatches: The color forward primers in this lab contain intentional mismatches at the chromophore region. Mismatches destabilize binding and effectively lower the Tm, which is why the insert fragment PCR uses a lower annealing temperature (53°C) compared to the backbone PCR (57°C).
Primer concentration: Higher primer concentrations shift the equilibrium toward annealing.
Secondary structure in the primer or template: Hairpins or self-dimers compete with proper annealing. The protocol recommends checking for these and keeping Gibbs free energy above −10 kcal/mol.
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Both PCR and restriction enzyme digestion produce linear DNA fragments, but they work through fundamentally different mechanisms and have different strengths.
Protocol Differences
Restriction digestion is simpler: you mix your DNA with the enzyme(s) in the appropriate buffer, incubate (often 37°C for 1 hour), and the enzyme cuts at its specific recognition sequence. PCR requires designing primers, setting up a reaction with polymerase and dNTPs, and running a thermocycling program with denaturation, annealing, and extension steps — a more involved setup that takes about 90 minutes.
Output Differences
Restriction enzymes cut at fixed, naturally occurring (or engineered) recognition sites, giving you no flexibility about exactly where the cut happens unless you’ve previously cloned in a new site. PCR, by contrast, lets you amplify any arbitrary region defined by your primer binding sites, giving you complete control over fragment boundaries. PCR also amplifies — you go from a tiny amount of template to millions of copies — while restriction digestion only cuts what’s already there, so you need more starting material.
Mutagenesis Capability
A key advantage of PCR is that primers can introduce mutations. The color forward primers contain intentional mismatches at the chromophore site, so the amplified product carries the desired mutation. Restriction enzymes can’t introduce new sequence — they only cut existing sequence.
When to Use Each
Restriction digestion is preferable when you have well-placed unique sites in your plasmid, when you want a simple and fast workflow, or when you need to avoid the risk of polymerase errors accumulating over many cycles. It’s the standard approach for traditional cloning into multiple cloning sites.
PCR is preferable when you need to amplify from a small amount of template, define custom fragment boundaries, introduce mutations, or add overhangs for assembly methods like Gibson.
In this lab, PCR is the right choice because you need to both introduce chromophore mutations and add overlapping ends for Gibson assembly — neither of which restriction digestion could accomplish alone.
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
Several verification steps are important:
Overlapping ends: Gibson assembly requires 20–40 bp of complementary sequence between adjoining fragments. You must confirm that your primer design creates these overlaps correctly — the 5′ overhang of each primer should be complementary to the end of the adjacent fragment.
DpnI digestion: After PCR, treating with DpnI destroys the methylated parental template plasmid, ensuring only your newly synthesized, unmethylated PCR products go into the Gibson reaction. Without this step, background colonies from intact template would confound results.
DNA purification: The Zymo Clean & Concentrator step removes primers, dNTPs, polymerase, and buffer salts that could interfere with the Gibson assembly enzymes.
Concentration measurement: Using Nanodrop or Qubit to verify DNA concentration (should be above ~30 ng/μL) ensures you can achieve the proper 2:1 insert-to-vector molar ratio.
Gel electrophoresis: Running a diagnostic gel confirms that your fragments are the expected size. An unexpected band size could indicate mispriming, non-specific amplification, or incorrect primer design.
Sequence verification: Confirming correct orientation (5′→3′) and that overlaps match between fragments prevents assembly failures.
How does the plasmid DNA enter the E. coli cells during transformation?
This lab uses chemical (heat-shock) transformation with chemically competent DH5α cells. These cells have been pre-treated with calcium chloride (CaCl&sub2;), which neutralizes the negative charges on both the cell membrane and the DNA, reducing electrostatic repulsion and allowing DNA to associate with the cell surface.
During the ice incubation (30 minutes), the DNA–cell complexes form at the membrane. The abrupt heat shock at 42°C for 45 seconds creates a thermal imbalance that transiently opens pores in the cell membrane, and the sudden temperature change also creates a concentration gradient that drives DNA into the cell by diffusion. Immediately returning the cells to ice for 5 minutes helps reseal the membrane and stabilize the cells.
After shock, the cells are given SOC medium and incubated at 37°C for 1 hour — this recovery period allows the cells to repair their membranes, begin replicating, and start expressing the antibiotic resistance gene (chloramphenicol resistance in this case) from the plasmid. When plated on selective media containing chloramphenicol, only cells that successfully took up and are expressing the plasmid will survive and form colonies.
Describe another assembly method in detail (such as Golden Gate Assembly).
Golden Gate Assembly is a one-pot, one-step cloning method that uses Type IIS restriction enzymes (most commonly BsaI or BbsI) to create seamless, scarless assemblies of multiple DNA fragments. Unlike conventional restriction enzymes that cut within their recognition site, Type IIS enzymes cut at a defined distance outside their recognition sequence, meaning the recognition site can be positioned so that it is removed from the final product entirely. This allows the designer to specify custom 4-base-pair sticky-end overhangs at each junction, enabling ordered, directional assembly of many fragments simultaneously. The reaction is run as a thermocycling protocol alternating between the restriction enzyme’s optimal temperature (~37°C) and the ligase’s optimal temperature (~16°C), which drives the equilibrium toward the correctly assembled product because correctly ligated junctions no longer contain the enzyme recognition site and cannot be re-cut. Golden Gate can efficiently assemble 10+ fragments in a single reaction, making it particularly powerful for combinatorial library construction or modular cloning systems like MoClo and PhytoBricks. Compared to Gibson assembly — which relies on sequence homology overlaps and works best with 2–6 fragments — Golden Gate offers more precise control over junction sequences and higher efficiency with many fragments, though it requires that the Type IIS recognition site not appear internally in any of your fragments.
How Golden Gate Works (Step by Step)
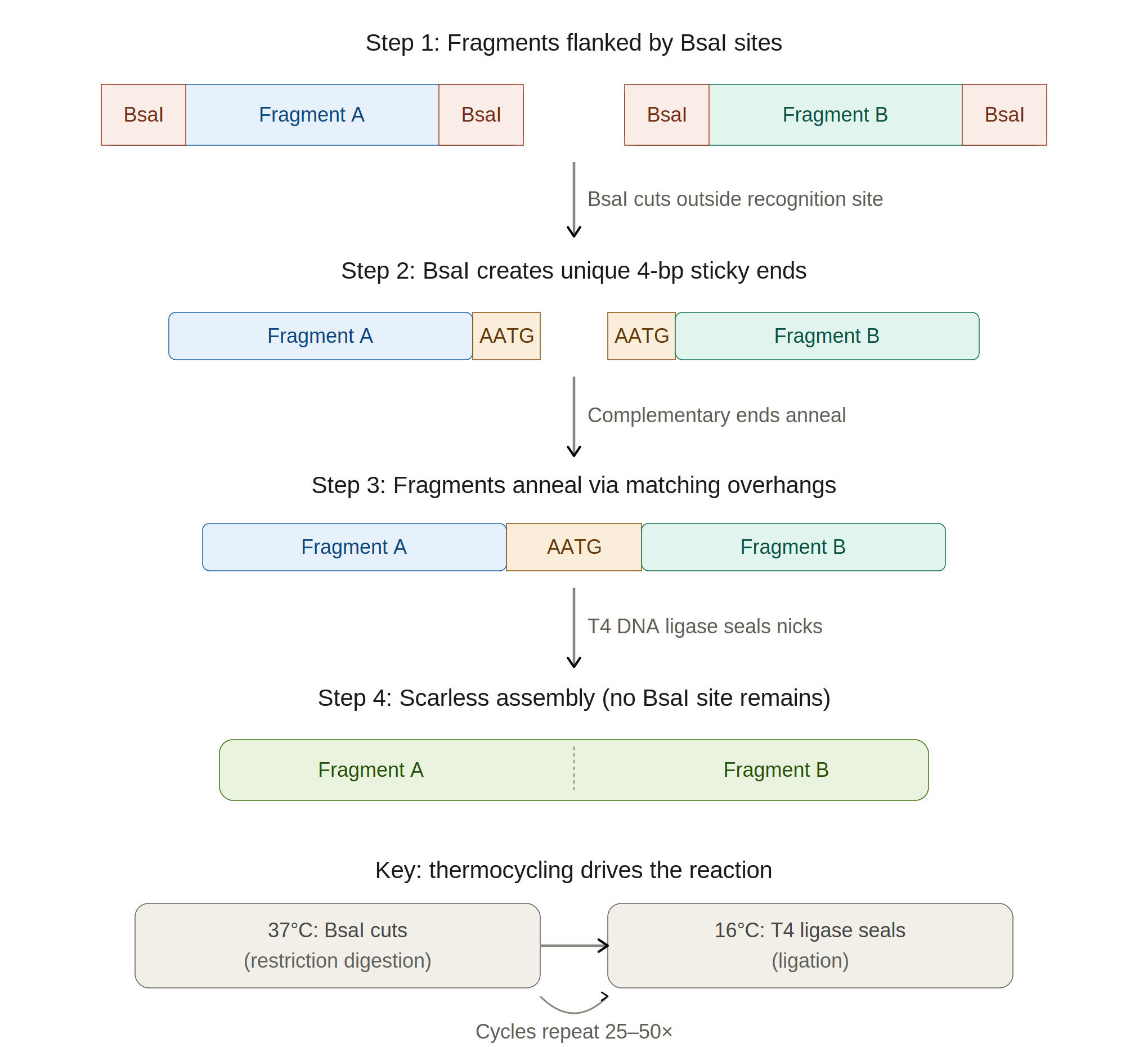

Comparing Golden Gate to Gibson (from this lab)
Gibson uses homologous overlaps (20–40 bp) and an isothermal reaction at 50°C with exonuclease, polymerase, and ligase. Golden Gate uses short 4-bp overhangs generated by restriction enzymes and alternates temperatures. Gibson is simpler for 2–3 fragment assemblies (like this chromophore lab), while Golden Gate excels at assembling many fragments (10+) in a defined order, since each junction can have a unique 4-bp overhang acting as an “address.”
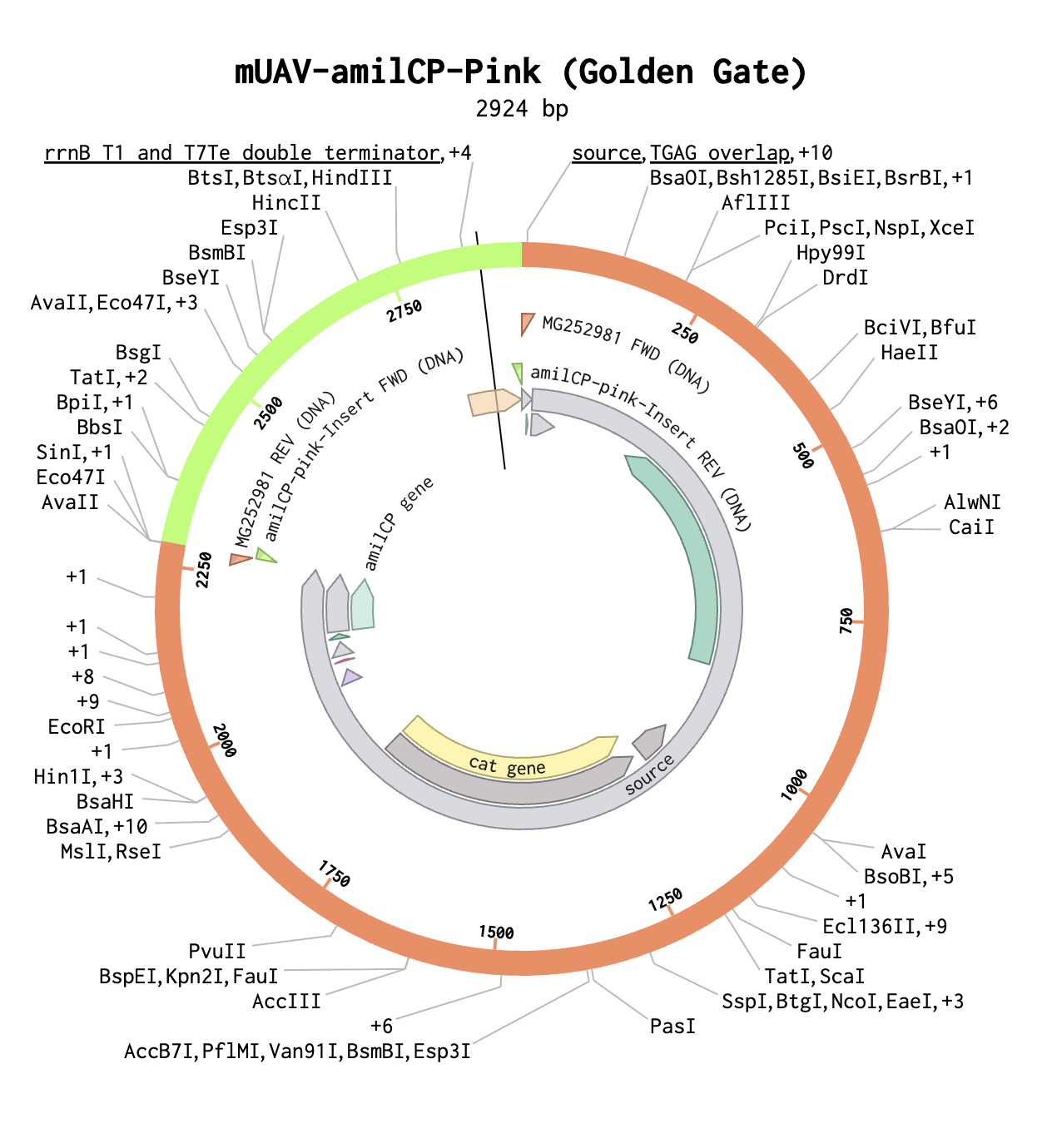
Model this assembly method with Benchling or Asimov Kernel!
This is the completed GGA with pink insert.

Asimov Kernel
Create a Repository and Notebook
Explore the Bacterial Demos Repo
Explore the devices in the Bacterial Demos Repo to understand how the parts work together by running the Simulator on various examples, following the instructions for the simulator found in the “Info” panel (click the “i” icon on the right to open the Info panel).
Recreate the Repressilator
Create a blank Construct and save it to your Repository. Recreate the Repressilator in that empty Construct by using parts from the Characterized Bacterial Parts repository — search the parts using the Search function in the right menu and drag and drop them into the Construct. Confirm it works as expected by running the Simulator (“play” button) and compare your results with the Repressilator Construct found in the Bacterial Demos repository.
Build Three Custom Constructs
Build three of your own Constructs using the parts in the Characterized Bacterial Parts Repo. For each construct, explain how you think it should function, run the simulator, and share the results. If the results don’t match your expectations, speculate on why and see if you can adjust the simulator settings to get the expected outcome.
Construct 1:
Construct 2:
Construct 3: