Week 7 HW: Genetic Circuits Part 2
Part 1: Intracellular Artificial Neural Networks (IANNs)
What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
IANNs offer three advantages over Boolean genetic circuits. They operate on graded, continuous intracellular signals rather than discrete ON/OFF states, enabling weighted summation, nonlinear activation, and universal function approximation. Weiss-coauthored neuromorphic circuits demonstrated these capabilities through analog computation, soft majority voting, and ternary switching in living cells. IANNs also permit tunable decision boundaries without topological redesign because effective weights and biases can be adjusted by modifying stoichiometry, promoter strength, or recognition-site placement. The PERSIST endoRNase system illustrates this: the same RNase acts as a repressor or activator depending on 5′-UTR versus 3′-UTR target-site positioning. Finally, multilayer IANNs have greater expressive power per circuit, representing smooth classifiers and nonlinearly separable response surfaces that Boolean truth tables cannot efficiently encode.
Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
An autonomous cell-state classifier for stem-cell differentiation would be a strong use case. The IANN would integrate sensors for an endothelial-intermediate RNA signature (x₁), residual pluripotency (x₂), and off-target lineage markers (x₃), computing a weighted sum z = w₁x₁ − w₂x₂ − w₃x₃ + b passed through a nonlinear output node driving a fluorescent reporter or differentiation factor. Weiss and colleagues used endoRNase-mediated miRNA sensors in a similar fashion to monitor cell-state transitions and guide multistep hiPSC differentiation toward a hematopoietic lineage. Limitations include resource loading in mammalian cells (Weiss’s 2020 work showed competing modules can reduce unregulated gene expression by up to 70%), RNase saturation and cross-cleavage at high enzyme ratios as observed in PERSIST cascades, stochastic weight variation across cells from poly-transfection, and the 650 ng total-DNA constraint imposed by the class protocol, which the supplied two-layer design already saturates.
Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.
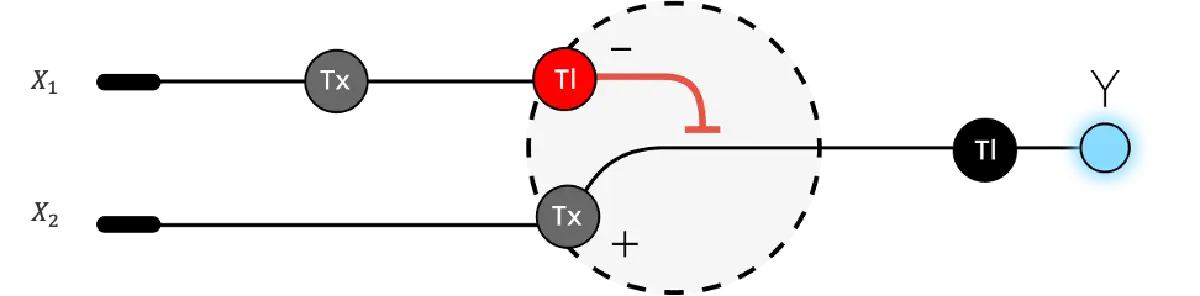

Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
The diagram below shows a two-layer intracellular perceptron built from the supplied parts. In Layer 1, input DNA X1 encodes Csy4, which is transcribed and translated into Csy4 protein that cleaves the Csy4 recognition site on the hidden-layer transcript (Csy4_rec_CasE), repressing it and producing the hidden-node output H = CasE. In Layer 2, CasE protein acts on the CasE recognition site in the output transcript (CasE_rec_mNeonGreen), repressing it to produce the fluorescent output Y = mNeonGreen. Both RNase links are drawn as repression to match the supplied single-layer example. In PERSIST-style designs, the sign of each edge can be inverted by repositioning the recognition site from a 5′-UTR OFF configuration to a 3′-UTR ON configuration. In the provided spreadsheet, this design corresponds to X1 = Csy4 + mKO2, X2 = Csy4_rec_CasE + eBFP2, and Bias = CasE_rec_mNeonGreen, and it consumes the full 650 ng class DNA limit.
Part 2: Fungal Materials
What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Most fungal materials are mycelium-based. Mycelium packaging (used by Dell as a Styrofoam replacement) is made by inoculating agricultural waste with fungal spores and molding it into custom shapes. Mycelium leather from MycoWorks and Bolt Threads (Mylo) uses roughly 70% less water and emits 68% fewer greenhouse gas emissions than cattle leather. Other products include fire-resistant construction and insulation panels with favorable thermal conductivity and sound absorption, compostable foams (Ecovative), and fungal protein food products (Mycorena). The common advantages over traditional counterparts are biodegradability, use of waste feedstocks, and reduced environmental impact. The common disadvantages are limited mechanical performance (mycelium compressive strength around 0.1 to 0.2 MPa versus 17 to 28 MPa for concrete), moisture susceptibility, batch-to-batch variability, scaling difficulties, and in the case of leather substitutes, affordability problems that have forced some manufacturers to shut down.
What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Engineering targets in fungi include modifying cell-wall biosynthetic genes (chitin synthase, alpha-glucan synthase, acetyltransferases) to tune material properties at the genome level, activating silent secondary-metabolite gene clusters through synthetic transcription factors or heterologous expression in hosts like Aspergillus oryzae, producing non-native compounds (cannabinoids, biofuels, therapeutic proteins), and embedding synthetic gene circuits into mycelium to create stimulus-responsive living materials. Fungi have several advantages over bacteria for synthetic biology. As eukaryotes, they perform post-translational modifications (glycosylation, disulfide bonds, proteolytic processing) that are needed for functional human therapeutic proteins. Filamentous fungi secrete 10 to 1,000 times more protein than bacterial hosts. They harbor secondary-metabolite pathways with large intron-containing gene clusters that bacterial systems cannot properly express. Mycelium grows into three-dimensional networks that can be used directly as structural materials, which no bacterial system offers. They also thrive on lignocellulosic waste streams that most bacteria cannot degrade. The tradeoffs are slower growth rates, less well-characterized genetics, and a synthetic biology toolkit that remains less mature than what is available for E. coli, though recent efforts like the Fungal Modular Cloning Toolkit (96 standardized parts for filamentous fungi) are narrowing the gap.
Part 3: First DNA Twist Order
All draft Round 0 constructs have been deposited in Benchling at https://benchling.com/seanmurp/f_/KopGo3fSDI-htgaa_final_project/, organized into sub-folders for Round 0 constructs (by reporter: sfGFP, mCherry, and NanoLuc) and controls.
The library comprises 80 unique 20-nt T7 promoter-spacer variants (+1 to +20) each paired with three reporters (sfGFP, mCherry, NanoLuc), yielding 240 test constructs plus 9 controls (dead-promoter negatives, no-RBS negatives, and synonymous codon-variant sfGFP controls). Spacers are drawn from five design categories: published reference variants (WT T7, T7Max, T7c62, T7#4), systematic ITS mutagenesis at positions +1 to +6, RBS/translation efficiency variants, context-interaction variants designed to produce reporter-dependent expression differences, and random space-filling variants for unbiased landscape coverage.
All constructs are designed as linear DNA fragments. The construct architecture is: 5’-[59 bp buffer]-[T7 consensus promoter]-[20 nt variable spacer]-[reporter CDS]-[T7 terminator]-[59 bp buffer]-3’, with total lengths ranging from ~720 bp (NanoLuc) to ~920 bp (sfGFP). The 59 bp flanking buffers provide protection against residual RecBCD exonuclease activity in the BL21 Star lysate, per Ginkgo’s recommendation of 50–80 bp padding for linear DNA templates in their cell-free system. Constructs will be synthesized as linear gene fragments (e.g., via Twist Bioscience) and used directly as CFPS templates at 15–20 nM without plasmid cloning.