Week3: Lab Automation
Review this week’s recitation and this week’s lab for details on the Opentrons and programming it.
This week was all about moving from manual pipetting to the world of liquid handling automation. I’ve been diving deep into the Opentrons ecosystem, specifically focusing on how to bridge the gap between writing Python code and seeing the robot actually execute those movements on the deck.
The Technical Workflow The core of my work this week involved getting comfortable with the Opentrons Python API (v2.13). I’ve learned that a solid protocol isn’t just about moving liquid; it’s about defining the environment perfectly. My scripts now follow a strict structure:
Metadata & Requirements: Setting the apiLevel and identifying the project.
Labware Loading: Mapping out the 11-slot deck (e.g., placing the 96-well plate in Slot 1 and the 300µL tip racks in Slot 4).
Instrument Definition: Defining the pipettes on the Left and Right mounts to ensure the robot knows its “limbs.”
The “Opentrons Art” Lab The highlight was the “Opentrons Art” challenge. It sounds fun, but it’s actually a high-stakes lesson in precision and coordinate systems.
Calibration is Everything: I spent a significant amount of time on the Labware Position Check (LPC). Even a 1mm offset in the Z-axis can lead to a crashed tip or an air-gap dispense that ruins the “painting.”
Optimization with Loops: Instead of hard-coding every single movement, I used Python for loops to iterate through the plate’s rows and columns. This makes the code cleaner and allows for more complex patterns with fewer lines of logic.
Fluid Dynamics: I practiced using .top() and .bottom() offsets during the dispense() command to control surface tension and avoid cross-contamination when “painting” with dyes.
Generate an artistic design using the GUI at opentrons-art.rcdonovan.com.
I want to make hourglass pattern and also ı want to add colors my pattern so ı used sfGFP and mRFP1 bacterias together.
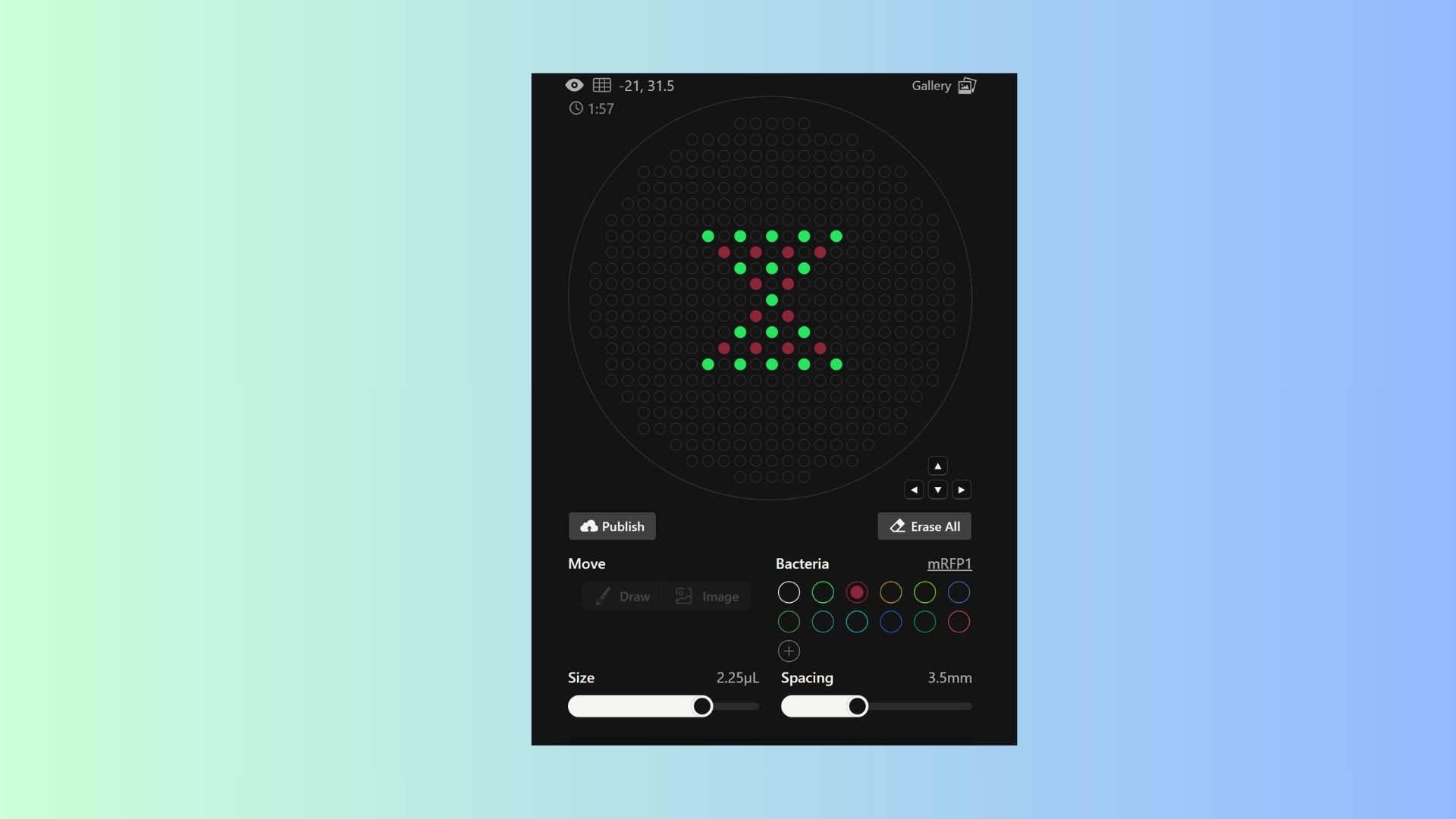

Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.
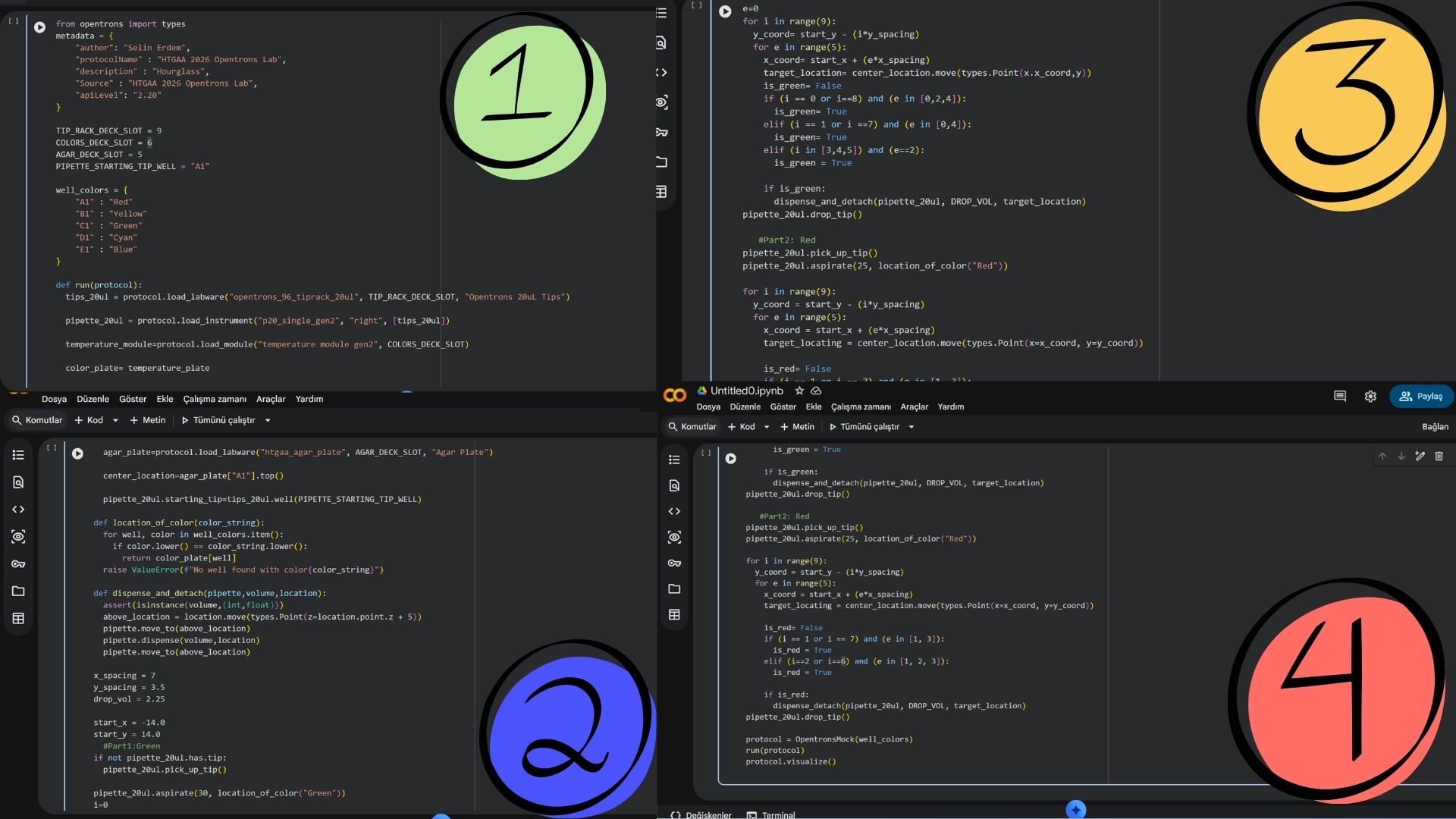

Post-Lab Questions:
1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Paper Title: Opentrons for automated and high-throughput viscometry (Soh et al., 2025)
https://www.sciencedirect.com/org/science/article/pii/S2635098X25000348
We always think that opentrons only transition liquid form but this article that completeley flips the script.Thay achieved that ınstead of just using the robot to prep samples, they actually turned the robot itself into the measuring tool. In bioengineering, measuring how hard to a liquid is because of the its viscosity. You need expensive rheometers, and doing it manually for hundreds of hydrogel or protein samples takes forever.These researchers had a great idea that why not use the physics of the pipette itself? They realized that the speed at which a liquid enters or leaves a pipette tip tells you exactly how viscous it is. They hacked their Opentrons OT-2 by clearing a spot on the deck for a high-precision scale. As the robot dispensed the liquid, they tracked the weight changes every 0.2 seconds. Then, they fed all that timing and weight data into a machine-learning model. So we didn’t have to do anything that much. Basically,the robot wasn’t just “moving” stuff anymore; it was “sensing” it. It could prep a brand-new biomaterial mix and immediately tell you its physical properties without a human ever touching a beaker.Therefore, ıt’s a perfect example of how automation isn’t just about saving time—it’s about doing science in a way that’s literally impossible for a human to do by hand.
2.Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
For my final project, I’m tackling a pretty complex challenge: developing a synthetic promoter that can sense when a “hallucination state” is occurring in a cell and then automatically suppress specific schizophrenia risk genes. Since I need to test a huge library of different promoter sequences to see which one reacts most precisely, doing this by hand would be a nightmare. This is where the Opentrons comes in.
So what is the plan?I want to use the robot as the “brain” of my high-throughput screening. Instead of manually pipetting 96 different versions of a promoter, I’ll program the Opentrons to handle the Golden Gate Assembly. The robot will mix the DNA parts, run the transformation into cells, and—most importantly—handle the dynamic dosing. I want to simulate “hallucination signals” by adding specific chemical inducers at different concentrations across a plate to see exactly at what threshold my synthetic promoter “turns on” the suppression. Also ı am thinking other tools to using for my project. For example, -Custom Labware: I’m thinking about 3D printing a custom rack to hold my specific microfuge tubes on the Opentrons deck so I don’t waste any of my expensive synthetic DNA.
-Ginkgo Nebula: Since designing synthetic promoters involves a lot of trial and error, I plan to use Ginkgo Nebula to synthesize the most promising DNA sequences I find. I’ll send my digital designs to them, get the DNA back, and then use the Opentrons to run the final verification experiments locally.
By using automation, I’m not just “doing an experiment”—I’m building a system that can find the perfect genetic switch for schizophrenia much faster and more accurately than I ever could manually.
Final Project Ideas





