Week 2 HW: DNA read, write and edit
Part 1
Here is the image that I managed to make using electrophoresis gel art:
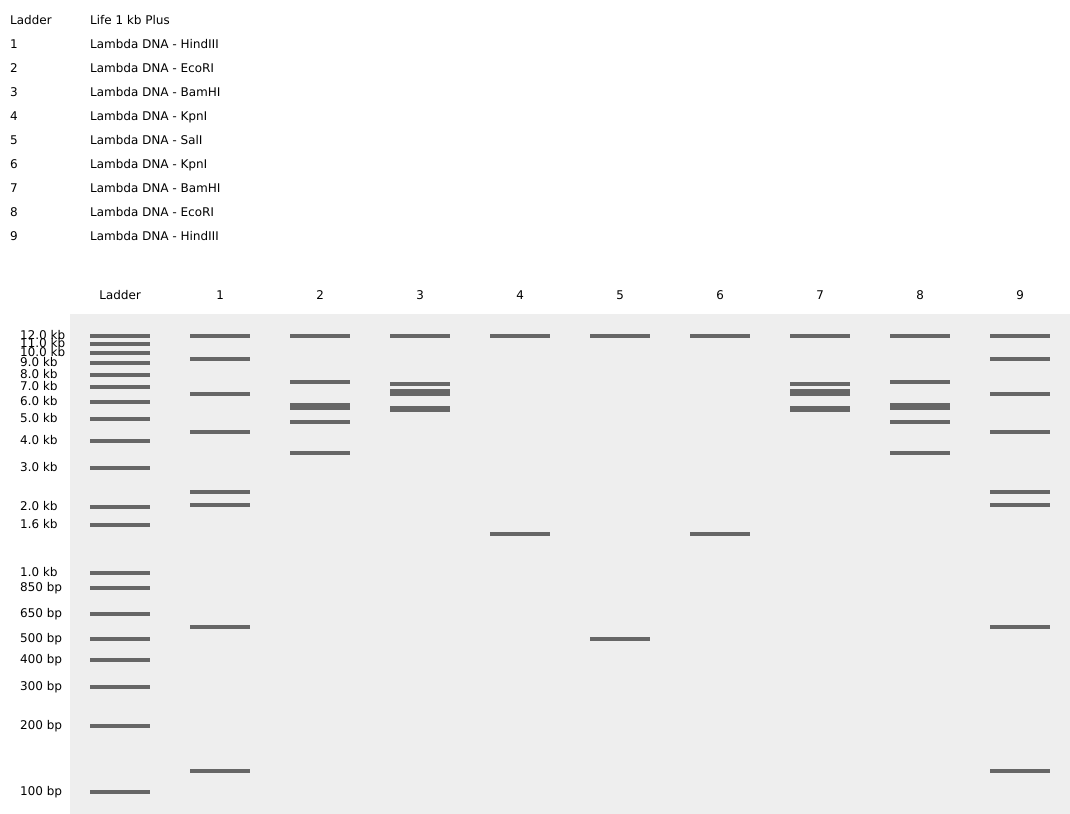


vs


It’s not really the same but if you imagine the emoji having long hair you can sort of see it.
A friend of mine said that it reminds it of Frisks face from undertale and ever since I’ve started seeing it aswell.


Part 3
I chose fibroin, which is one of the two main proteins found in silk. Silk in its raw state consists of two main proteins, sericin and fibroin, with a glue-like layer of sericin coating two singular filaments of fibroin called brins. I used uniprot to find a sequence for fibroin that is produced by the Aliatypus thompsoni, also know as the trapdoor spider: SQ SEQUENCE 193 AA; 18898 MW; 94DFC97AA25F9796 CRC64; ASSASGASSS IGVASSKGVA SSSKTATKAR ISAGSSGSST STKSSSSAST AVPTNLSGSR SHALSSSNSG QDNTVGDDFG LGYISGGILP VNTPALNFPS DLGSLTSGLL SSLDGPVLPS VEYRITSLTS SVLSLLSTSG GAFNYSSFAK NLAILAYQIS VSNPGLSVSQ VVSETLLESV GALIHILVSS QVG
After I reverse translated, the protein looks like this (using https://www.genecorner.ugent.be/rev_trans.html): GCGAGCAGCGCGAGCGGCGCGAGCAGCAGCATTGGCGTGGCGAGCAGCAAAGGCGTGGCG AGCAGCAGCAAAACCGCGACCAAAGCGCGCATTAGCGCGGGCAGCAGCGGCAGCAGCACC AGCACCAAAAGCAGCAGCAGCGCGAGCACCGCGGTGCCGACCAACCTGAGCGGCAGCCGC AGCCATGCGCTGAGCAGCAGCAACAGCGGCCAGGATAACACCGTGGGCGATGATTTTGGC CTGGGCTATATTAGCGGCGGCATTCTGCCGGTGAACACCCCGGCGCTGAACTTTCCGAGC GATCTGGGCAGCCTGACCAGCGGCCTGCTGAGCAGCCTGGATGGCCCGGTGCTGCCGAGC GTGGAATATCGCATTACCAGCCTGACCAGCAGCGTGCTGAGCCTGCTGAGCACCAGCGGC GGCGCGTTTAACTATAGCAGCTTTGCGAAAAACCTGGCGATTCTGGCGTATCAGATTAGC GTGAGCAACCCGGGCCTGAGCGTGAGCCAGGTGGTGAGCGAAACCCTGCTGGAAAGCGTG GGCGCGCTGATTCATATTCTGGTGAGCAGCCAGGTGGGC
Afterwards I had to optimize the codon sequence, which results in this: GCATCTAGCGCGTCTGGTGCGAGCAGCTCTATTGGTGTGGCAAGCTCTAAGGGCGTTGCGTCTAGCAGCAAAACCGCGACCAAAGCGCGTATTAGCGCGGGTAGCAGCGGCAGCTCTACCAGCACCAAGTCTAGCAGCTCTGCAAGCACTGCGGTTCCGACCAACTTATCTGGCTCTCGTAGCCATGCATTAAGCTCTTCTAACAGCGGCCAGGACAACACTGTTGGTGATGATTTTGGCCTGGGCTATATTAGCGGTGGCATCCTGCCAGTTAACACCCCAGCGCTGAATTTTCCAAGCGATTTAGGCTCTTTAACTAGCGGCCTGCTGAGCTCTTTAGATGGCCCAGTGTTACCGTCTGTTGAGTATCGTATCACTTCTTTAACTTCTTCTGTTTTAAGCCTGCTGAGCACTAGCGGCGGTGCGTTTAACTACTCTAGCTTCGCGAAAAACCTGGCAATCCTGGCGTACCAAATCAGCGTTAGCAACCCGGGTCTGAGCGTGTCTCAGGTGGTTTCTGAGACCCTGTTAGAATCTGTTGGCGCACTGATCCATATCCTGGTGAGCAGCCAGGTGGGT
The question still stands… How can I produce this protein using my DNA?
Well obvioulsy I can’t make the sequence straight from my DNA since my body doesn’t produce silk. What I can do instead is use my DNA as a backbone (which I can just extract from my cells) and edit it so it has the fibroin sequence using CRIPS-based editing or PCR-based gene assembly. After I have the right sequence for fibroin production, it will be insedrted into a plasmid and then transported into E-coli cells. Inside a cell, RNA polymerase reads the DNA sequence and it produces mRNA copy of it. Ribosomes translates it into the polypeptide chain which is then folded into it’s desired shape (in our case silk fibers).
Part 4
4.2
This is the given example with the E-coli glowing phosphorescent green under UV light.
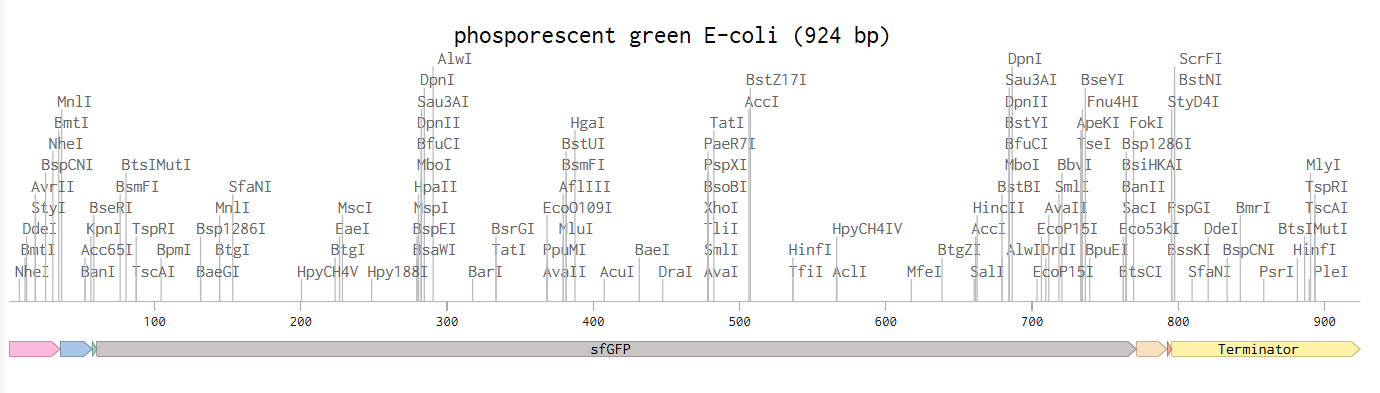

4.3-6
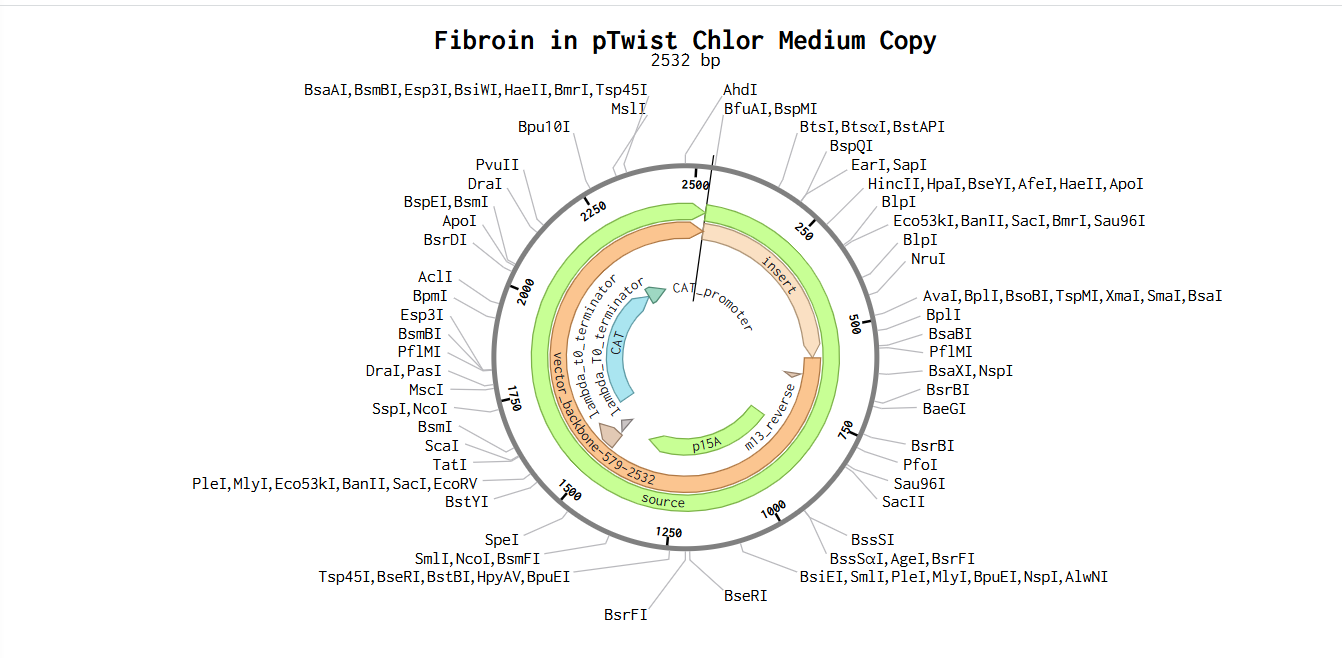

Part 5
5.1 DNA READ
i) Since I’m a computer engineer at base, I would love to be able to make a synthetic DNA data storage. Other than my personal preference for this conjucture of amazing subjects, DNA data storage is the most efficient way of storing information since it doesn’t need any energy after creation and can store insane ammounts of information (~215 petabytes in 1 gram of DNA) for long periods of time in minimal spaces. However sequencing is critical due to the errors that can occur during encoding. Since DNA stores information passively, sequencing is crucial when decoding aswell.
ii) I would most likely need the Third-generation sequencing mostly because the DNA sequences that are supposed to be stored are long. Not only that, but since ONT can read continious sequences of DNA molecules and give me real-time answers clearly makes it the superior option. In addition, it does not require PCR amplification to create a signal, meaning it bypasses the “vicious” cycle of amplification biases that can sometimes lead to errors in GC-rich regions of the genome.
5.2 DNA write
i) In my opinion, it would be a dream to be able to produce a synthetic genetic construct that allows a host organism to efficintly produce spider silk proteins. The production of silk should most likely be regulated so I know exactly the length of each strand, or why not make them arbitrarily. Why you may ask? Silk is one of the strongest biodegradable materials known to man, and it also has a lot of uses in medicine, textile industry and engineering. For some kids out there maybe the “I want to become Spiderman” wish might come true one day.
ii) So what technologies would I need?
If I wanted to synthesize a spider‑silk gene, I’d start by ordering commercially made oligonucleotides and then assemble them using methods like Gibson Assembly or overlap‑extension PCR. These approaches are well‑suited for building long, repetitive genes and let me incorporate codon optimization along with any regulatory elements I need, such as promoters, ribosome‑binding sites, and terminators.
To verify the construct, I’d rely on a combination of Sanger sequencing and long‑read next‑generation sequencing (like PacBio or Oxford Nanopore). Sanger is extremely accurate (around 99.99%) but it’s slow and not ideal for very long or highly repetitive sequences, so I’d use it mainly to confirm individual repeat units. Long‑read NGS, on the other hand, can capture the entire gene in one go and handles repetitive regions well, though each read is a bit less accurate and requires some bioinformatic assembly.
Using both methods together gives a high level of confidence that the full‑length gene was synthesized correctly, striking a good balance between accuracy, speed, and scalability.
5.3 DNA edit
i) When I think about valuable DNA edits, I’m most interested in changes that improve health, resilience, and environmental sustainability, rather than enhancements for their own sake. In humans, this could mean correcting disease-causing mutations, like inherited metabolic disorders or certain forms of blindness, where fixing a single faulty gene could prevent suffering without changing a person’s identity. Another promising area is reducing age-related decline, helping people stay healthier as they get older rather than trying to extend lifespan indefinitely. To me, the worst part about aging isn’t my appearance changing, but me losing my senses (poorer eyesight and hearing), so something of great interest to me would be to manage to slow down something like eyesight loss.
I’m cautious about human enhancement, such as boosting intelligence or physical ability, because of ethical concerns around fairness and consent. But using DNA editing to prevent disease, strengthen ecosystems, and support sustainable agriculture seems like a direction with clear benefits and manageable risks.
ii) What technology or technologies would you use to perform these DNA edits and why?
If the goal is to reduce eyesight degradation, particularly conditions driven by inherited mutations or age‑related cellular decline—the most appropriate technologies would be those that offer high precision and minimal disruption to the genome. In this context, CRISPR‑based tools remain the leading candidates, but not the traditional cut‑and‑repair versions. Instead, I would look toward base editing and prime editing, which are designed to make extremely targeted changes without creating double‑strand breaks. That matters because retinal cells are delicate, slow‑dividing, and difficult to replace, so any editing approach needs to be as gentle and predictable as possible.
Base editors are useful when eyesight degradation stems from a single incorrect DNA letter, because they can convert one nucleotide into another with high specificity. Prime editors go a step further, allowing slightly larger corrections or small insertions and deletions, which could be valuable for conditions where the underlying mutation is more complex. Both technologies aim to correct the genetic cause of degeneration before it leads to irreversible damage.
For age‑related vision loss, where the problem isn’t a single mutation but a gradual decline in cellular function, the focus would shift toward editing pathways that support cellular resilience, mitochondrial health, or protective responses to oxidative stress. Again, precision tools like prime editing would be preferable, because they allow subtle adjustments to gene regulation without fundamentally altering the identity or behavior of retinal cells.
Across all of these possibilities, the priority is safety: technologies that minimize off‑target effects, avoid unnecessary DNA breaks, and allow careful control over the extent of the edit. The retina is one of the most sensitive tissues in the body, so any intervention must be both scientifically justified and ethically grounded