Week 2 HW: DNA r/w/e
Part 0: Basics of Gel Electrophoresis Attend or watch all lecture and recitation videos. Optionally watch bootcamp.
Part 1: Benchling & In-silico Gel Art See the Gel Art: Restriction Digests and Gel Electrophoresis protocol for details. Overview:
Make a free account at benchling.com Import the Lambda DNA. Simulate Restriction Enzyme Digestion with the following Enzymes: EcoRI HindIII BamHI KpnI EcoRV SacI SalI Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks. You might find Ronan’s website a helpful tool for quickly iterating on designs!
Part 0: Basics of Gel Electrophoresis
Part 0 reviews the fundamental biological principles that support the rest of this project. Understanding how genetic information flows inside cells is essential for designing and interpreting molecular biology experiments.
DNA as the Information Storage Molecule
DNA (deoxyribonucleic acid) is the molecule that stores genetic information in living organisms. It consists of two complementary strands arranged in a double helix. Each strand is made of nucleotides containing four bases: adenine (A), thymine (T), cytosine (C), and guanine (G).
The sequence of these bases encodes instructions for building proteins. DNA is chemically stable, making it ideal for long-term information storage. During experiments such as restriction digests and gel electrophoresis, we manipulate DNA directly to analyze or modify genetic information.
RNA and Transcription
RNA (ribonucleic acid) is a temporary copy of genetic instructions. During transcription, an enzyme called RNA polymerase reads a DNA template strand and synthesizes messenger RNA (mRNA).
RNA differs from DNA in three key ways:
It contains ribose sugar instead of deoxyribose
It uses uracil (U) instead of thymine (T)
It is usually single-stranded
mRNA carries genetic instructions from DNA to ribosomes, where proteins are produced.
Proteins and Translation
Proteins are functional molecules that perform most cellular tasks, including catalysis, structure, and signaling. During translation, ribosomes read mRNA in groups of three nucleotides called codons. Each codon corresponds to a specific amino acid.
A chain of amino acids folds into a three-dimensional structure that determines the protein’s function. In this project, designing DNA sequences ultimately aims to control which proteins are produced.
The Central Dogma of Molecular Biology
The relationship between DNA, RNA, and protein is summarized by the central dogma:
DNA → RNA → Protein
This directional flow explains how genetic information is expressed inside cells. All molecular biology techniques used in this assignment — including cloning, restriction digests, and gene expression — rely on manipulating this pathway.
Restriction Enzymes and DNA Manipulation
Restriction enzymes are proteins that cut DNA at specific sequences. These enzymes allow scientists to divide DNA into predictable fragments. By selecting particular enzymes, researchers can design DNA pieces that generate specific band patterns during gel electrophoresis.
This precise cutting ability is the foundation of genetic engineering and is essential for both analytical and creative gel art design.
Gel Electrophoresis Principles
Gel electrophoresis separates DNA fragments by size. Because DNA carries a negative charge, it migrates toward the positive electrode in an electric field.
Smaller fragments move faster through the agarose gel matrix, while larger fragments move more slowly. This separation produces visible bands that correspond to fragment length.
By comparing observed bands to predicted fragment sizes, researchers can verify DNA structure and confirm successful restriction digests.
Part 1: Benchling & In-silico Gel Art
Part 1 focuses on designing a gel electrophoresis experiment using virtual simulation tools before performing any physical lab work.
The primary goal of this design phase is to create a controlled DNA banding pattern through selective restriction enzyme digestion. Instead of randomly cutting DNA, the experiment is planned so that specific fragment sizes generate a visual composition on an agarose gel.
This approach transforms gel electrophoresis from a purely analytical technique into a hybrid scientific and artistic exercise. At the same time, it reinforces essential molecular biology concepts such as enzyme specificity, fragment prediction, and experimental reproducibility. Benchling’s virtual digest tool is used to simulate how restriction enzymes cut a known DNA substrate. By testing different enzyme combinations digitally, predicted fragment lengths can be analyzed without consuming physical reagents.
After creating a free account on benchling.com and importing the Lambda DNA, restriction enzyme digestion was simulated using the following enzymes:
EcoRI
HindIII
BamHI
KpnI
EcoRV
SacI
SalI
Resulting in:
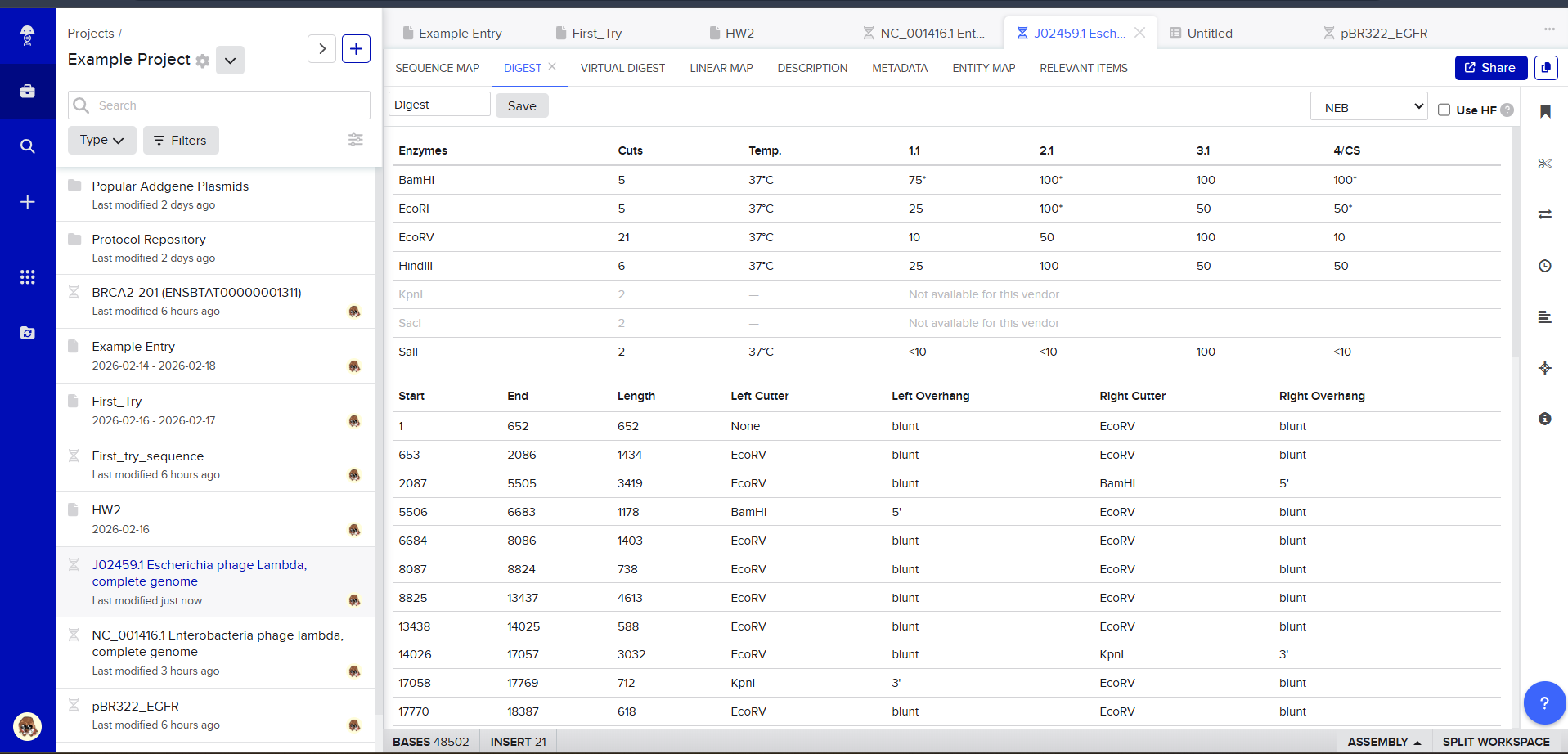

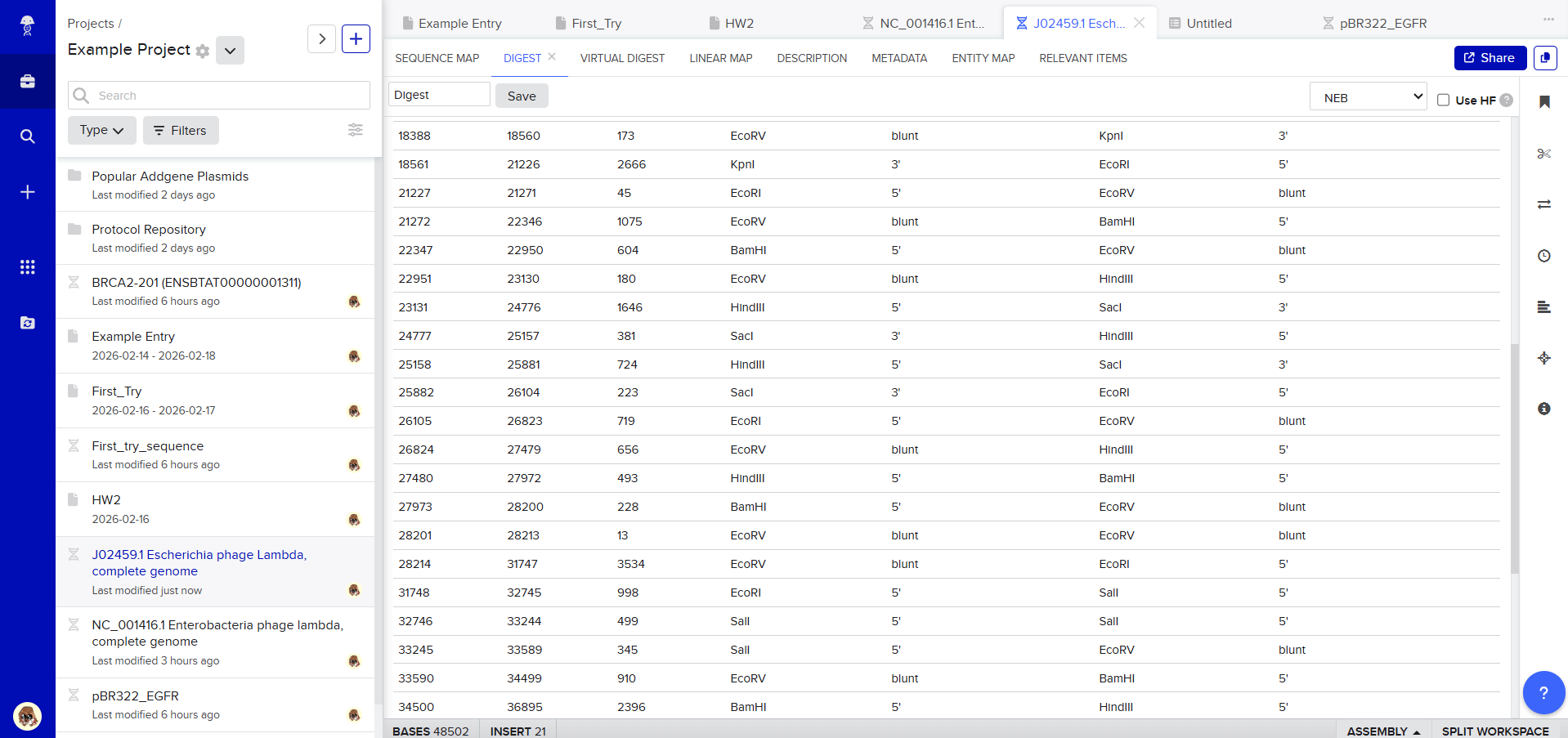

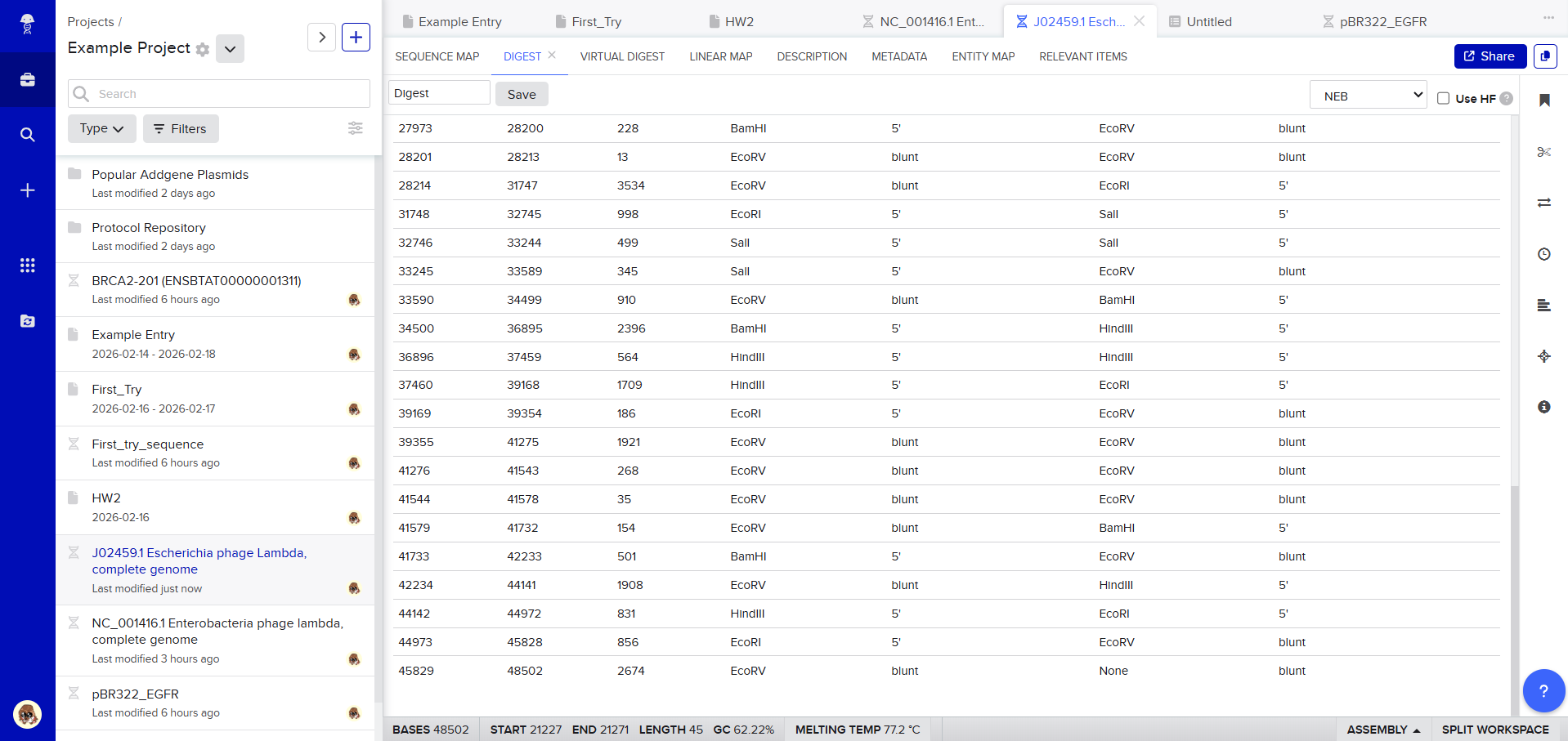

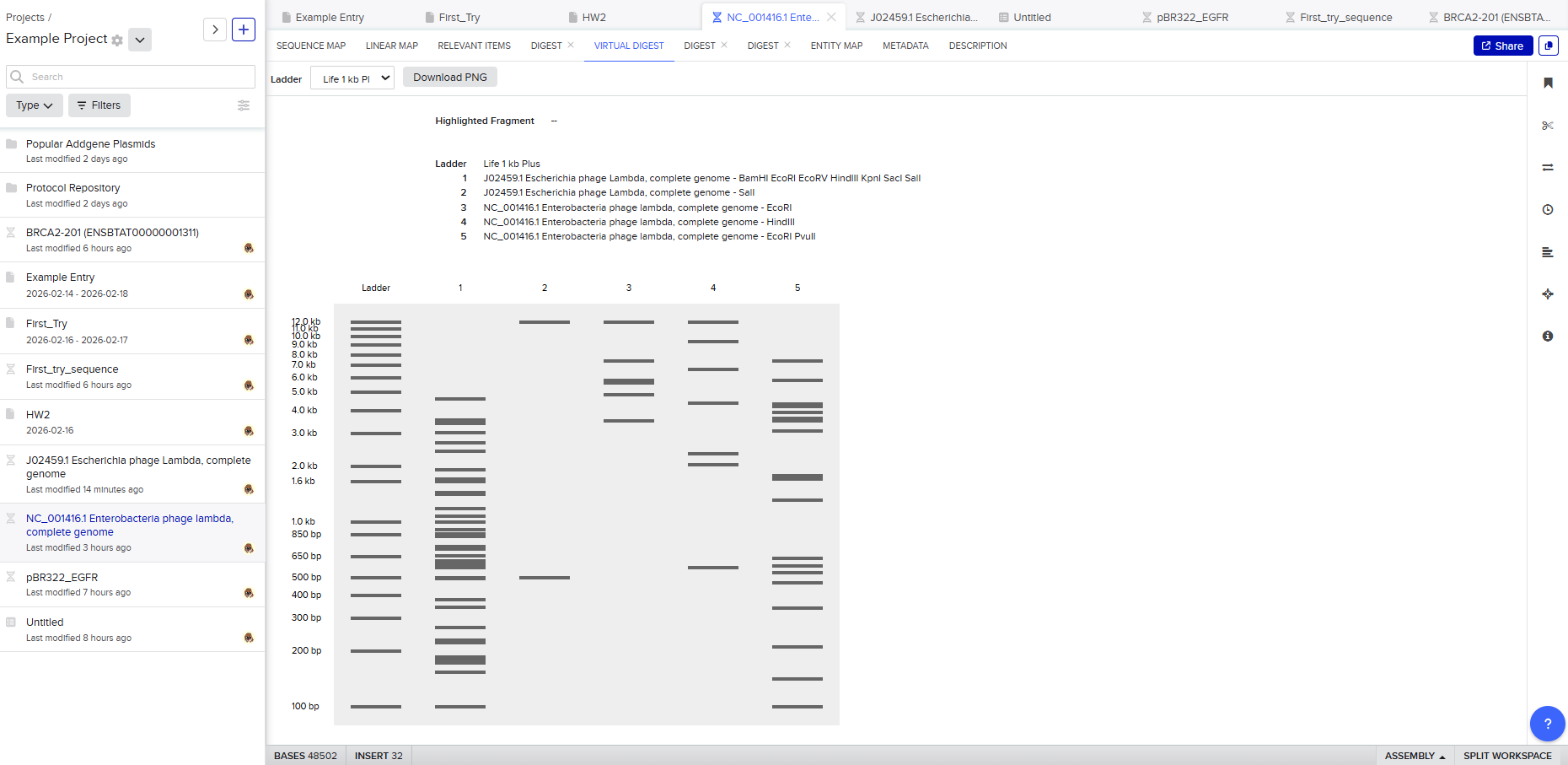
Then, go to the virtual digest tab to see how the digest looks. This visualization uses all the enzymes on the list.
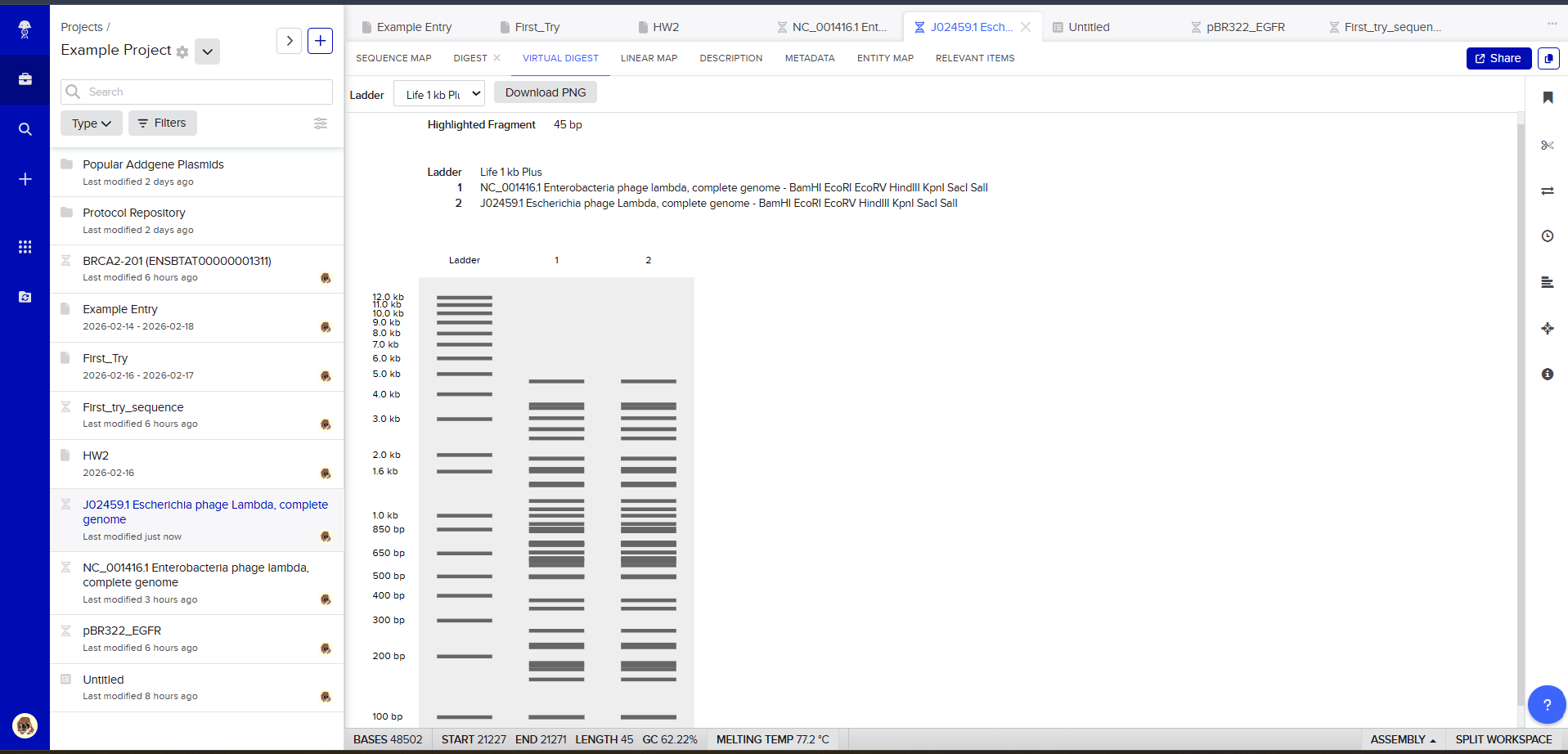

After seeing what could be done with the enzymes, I continued testing more combinations. For faster iteration, I used Ronan’s website to get more images. After several attempts, I ended up with the following iteration:
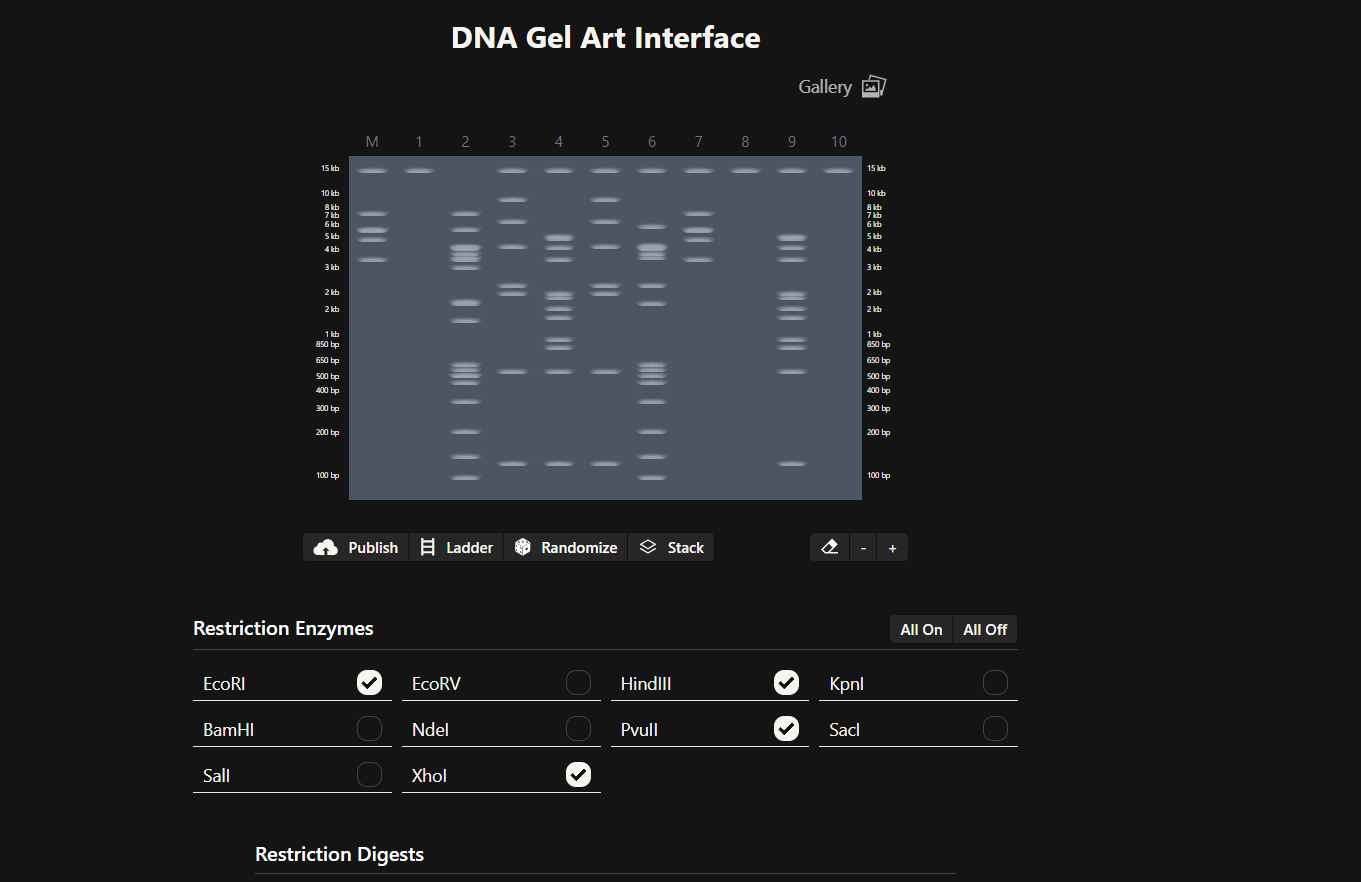

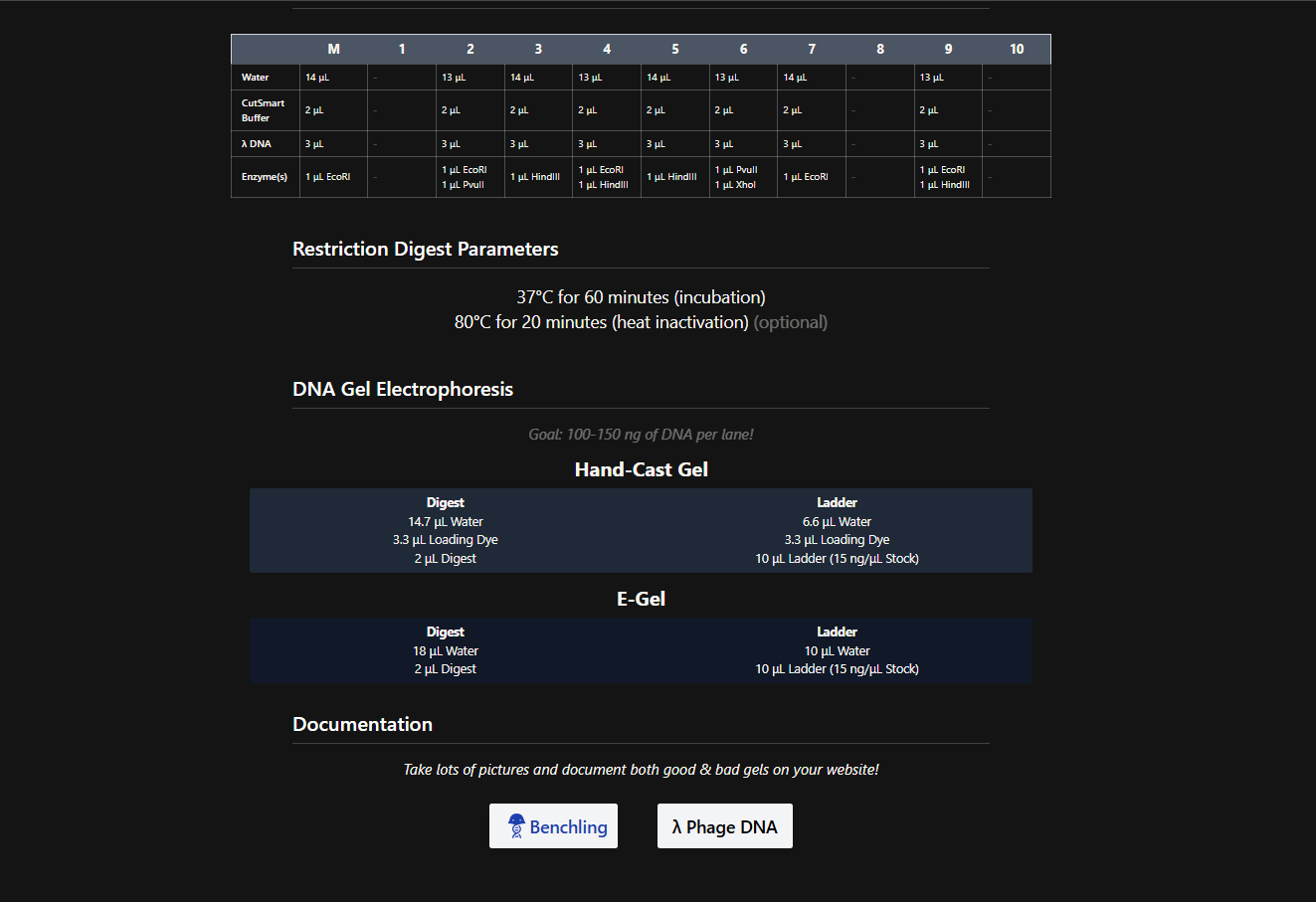

I liked it a lot because when I saw it, I don’t know why, a sculpture of the ancient Incas came to mind at that moment.


I don’t know if you see it too, but here are a few lines to see if it makes it easier to detect.


Anyway, I tried to make that drawing look like Paul Vanouse’s Latent Figure Protocol artwork. But I didn’t know how to do it, so I decided to ask Gemini how I could do it. This is the result:
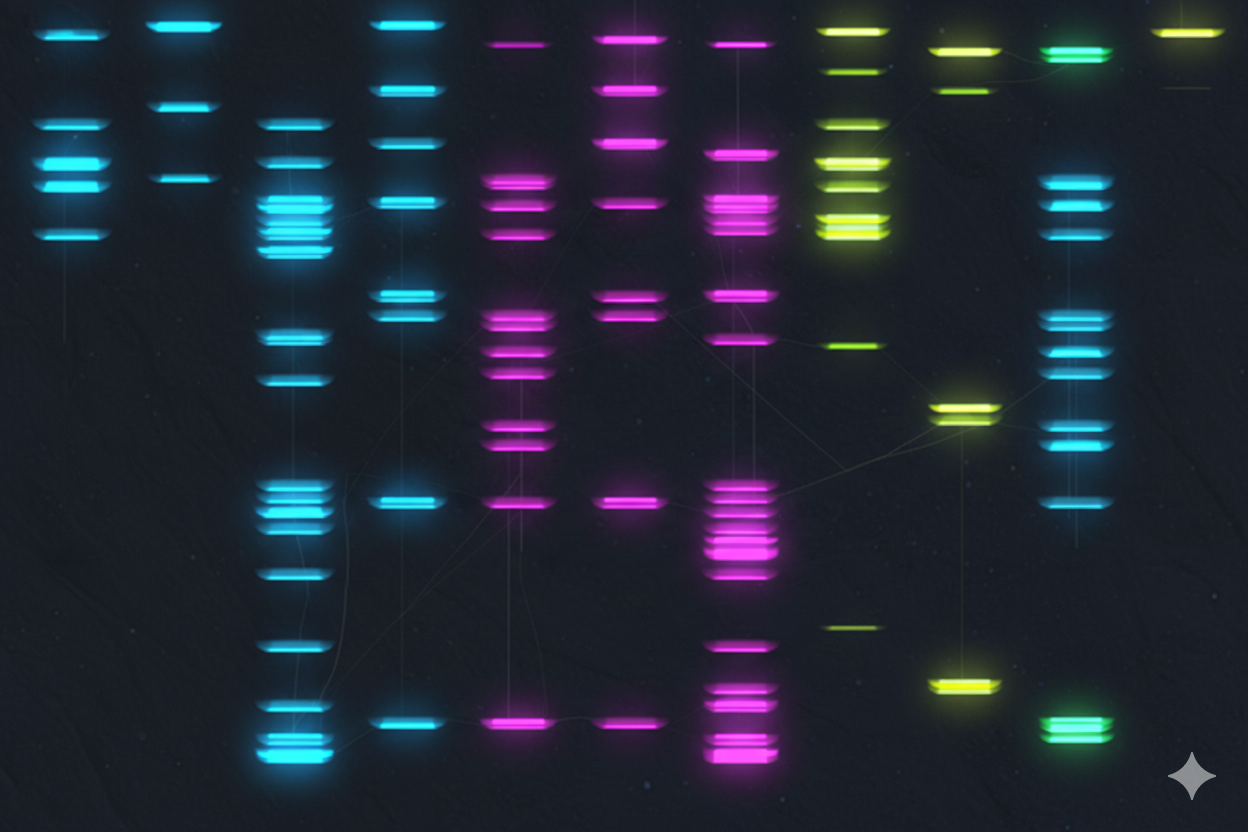


It’s not exactly what I expected; it doesn’t really resemble that style of art, but I ended up liking it.
Then, I tried to replicate it in Benchling using the enzymes the website mentioned. The bad thing is that it didn’t turn out as I expected. I’m still not sure what went wrong, but I didn’t make many attempts to recreate it; I didn’t have much time.

But if you look closely, it could easily resemble a level you’d find while playing Mario Maker. Well, that’s what I can see; I don’t know what you all think.
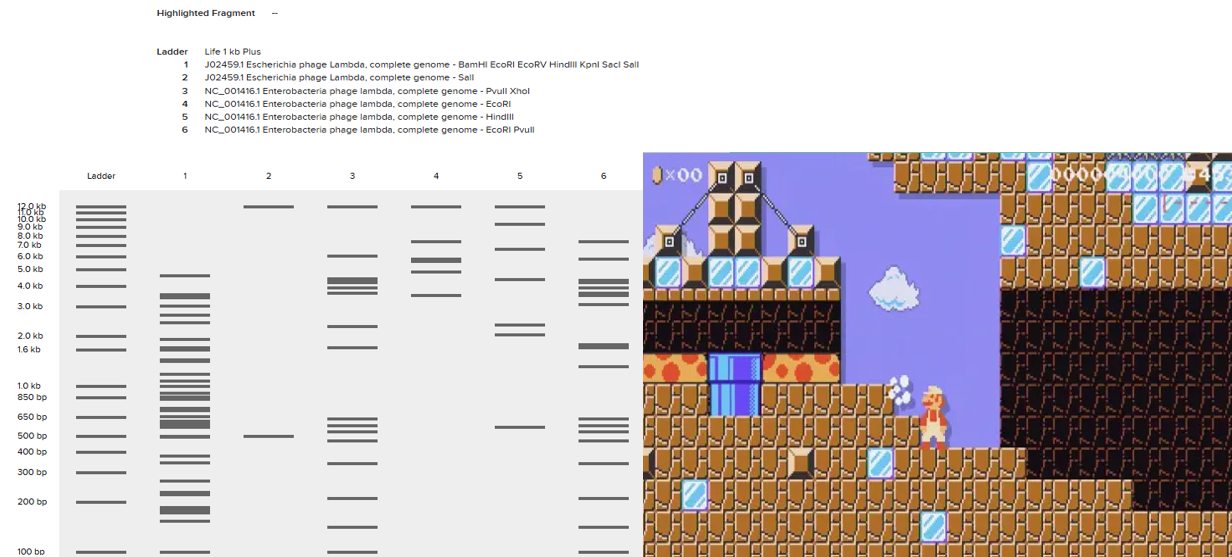

In the end, it’s a tool I need to practice more, but I really liked how it works. But let’s leave opinions aside and move on to the rest of HW2.
Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
Perform the lab experiment you designed in Part 1 and outlined in the Gel Art: Restriction Digests and Gel Electrophoresis protocol.
Part 2: Gel Electrophoresis Experiment (Simulation and Analysis)
There was no lab available at my node this week, so I couldn’t complete this part. Instead, I completed a detailed virtual simulation of the protocol using Benchling and theoretical analysis of the expected outcomes. This allowed me to understand the experimental workflow and interpret how restriction digests generate DNA fragment patterns that can be visualized as gel art.
The experiment would begin with designing a restriction digest of Lambda DNA using selected high-fidelity restriction enzymes. By importing the Lambda DNA sequence into Benchling and running virtual digests, I tested different enzyme combinations to predict fragment sizes and design a gel pattern inspired by gel art. This simulation demonstrated how enzyme selection directly influences the final banding pattern.
If performed in a physical laboratory, the next step would involve preparing a 1% agarose gel in TAE buffer and staining it with a fluorescent dye. The digested DNA samples would be mixed with loading dye and pipetted into the gel wells. When an electric field is applied, negatively charged DNA fragments migrate toward the positive electrode. Smaller fragments move faster through the agarose matrix, resulting in size-based separation.
After electrophoresis, the gel would be imaged using a blue light transilluminator. The resulting band pattern would be compared with the virtual digest predictions. Agreement between expected and observed fragment sizes would confirm successful restriction digestion and validate the DNA design used to create the gel art.
Although I did not physically run the gel, performing the simulation reinforced key molecular biology concepts, including restriction enzyme specificity, fragment size prediction, and electrophoretic separation. This exercise highlights how computational tools can effectively model laboratory experiments and support experimental planning in situations where physical lab access is limited.
Part 3: DNA Design Challenge 3.1. Choose your protein.
In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of >the tools described in recitation (NCBI, UniProt, google), obtain the protein sequence for the protein you chose.
[Example from our group homework, you may notice the particular format — The example below came from UniProt]
sp|P03609|LYS_BPMS2 Lysis protein OS=Escherichia phage MS2 OX=12022 PE=2 SV=1 METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLL EAVIRTVTTLQQLLT
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The Central Dogma discussed in class and recitation describes the process in which DNA sequence becomes transcribed and translated into protein. The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from. Using one of the tools discussed in class, NCBI or online tools (google “reverse translation tools”), determine the nucleotide sequence that corresponds to the protein sequence you chose above.
[Example: Get to the original sequence of phage MS2 L-protein from its genome phage MS2 genome - Nucleotide - NCBI]
Lysis protein DNA sequence atggaaacccgattccctcagcaatcgcagcaaactccggcatctactaatagacgccggccattcaaacatgaggattacccatgtcgaagacaacaaagaagttcaactctttatgtattgatcttcctcgcgatctttctctcgaaatttacca>atcaattgcttctgtcgctactggaagcggtgatccgcacagtgacgactttacagcaattgcttacttaa
3.3. Codon optimization.
Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
[Example from Codon Optimization Tool | Twist Bioscience while avoiding Type IIs enzyme recognition sites BsaI, BsmBI, and BbsI]
Lysis protein DNA sequence with Codon-Optimization ATGGAAACCCGCTTTCCGCAGCAGAGCCAGCAGACCCCGGCGAGCACCAACCGCCGCCGCCCGTTCAAACATGAAGATTATCCGTGCCGTCGTCAGCAGCGCAGCAGCACCCTGTATGTGCTGATTTTTCTGGCGATTTTTCTGAGCAAATTCACCAACCAGCTGCTGCTGAGCCTGCTGGAAGCGGTGATTCGCACAGTGACGACCCTGCAGCAGCTGCTGACCTAA
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
3.5. [Optional] How does it work in nature/biological systems?
Describe how a single gene codes for multiple proteins at the transcriptional level. Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!! See example below. [Example shows the biomolecular flow in central dogma from DNA to RNA to Protein] Special note that all “T” were transcribed into “U” and that the 3-nt codon represents 1-AA.
3.1 Choose your protein
For this assignment, I selected the Damage Suppressor protein (Dsup) from tardigrades. Dsup is a remarkable protein that has been shown to protect cellular DNA from radiation and oxidative stress. Tardigrades are microscopic extremophiles capable of surviving severe environmental conditions, including intense radiation, dehydration, extreme temperatures, and even the vacuum of space. Their resilience has attracted significant interest in bioengineering and astrobiology.
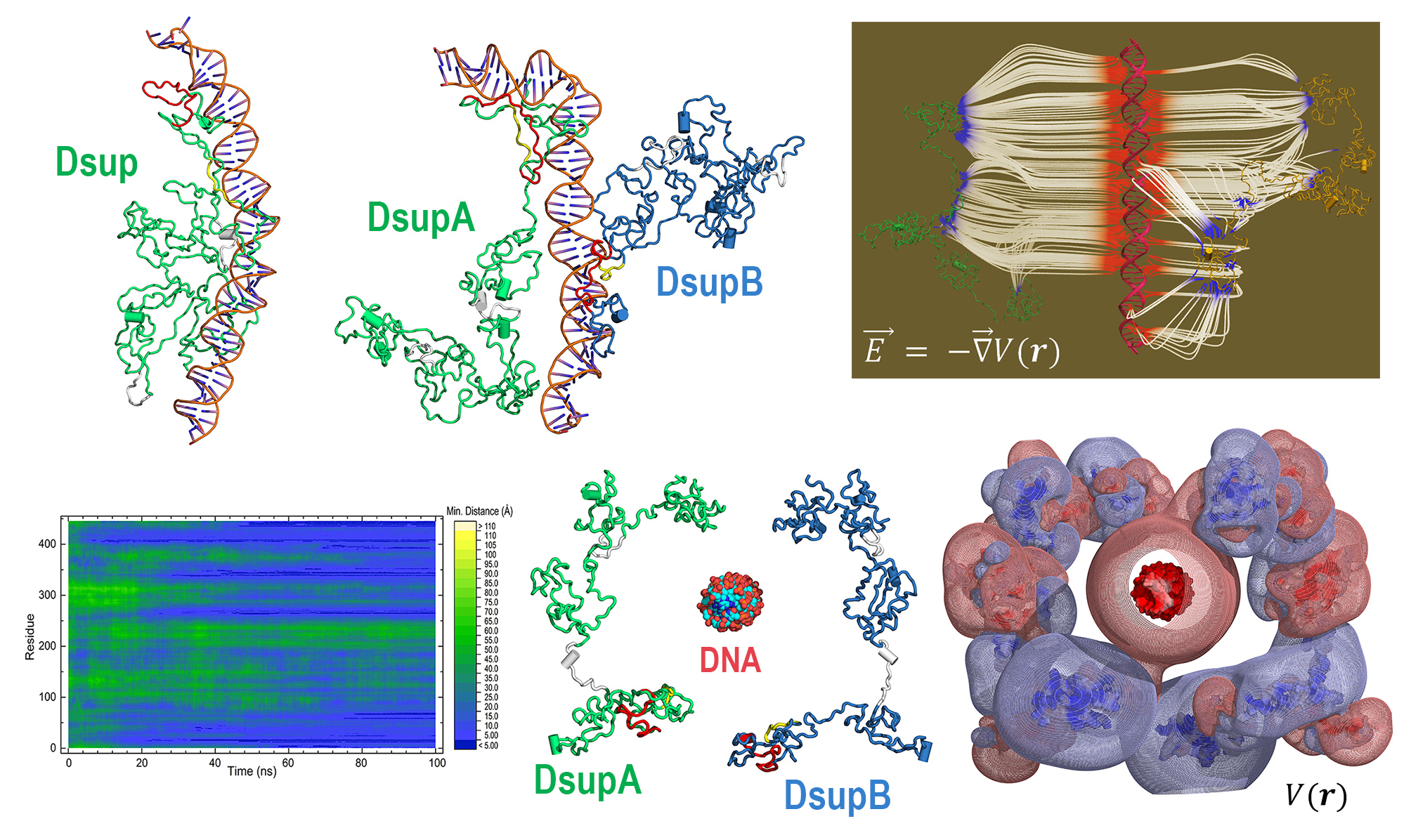

I chose Dsup because it represents a compelling intersection between fundamental biology and applied biotechnology. Its protective properties suggest potential applications in radiation protection for human cells, improvement of stress resistance in engineered microorganisms, and future space exploration where biological systems are exposed to harsh environments. Studying and expressing this protein could contribute to the development of more robust biological systems.
Using the UniProt protein database, I obtained the amino acid sequence of the Dsup protein. UniProt provides curated protein information, including functional annotations and sequence data. The protein sequence used for this project is shown below in FASTA format.
Protein sequence (excerpt): I downloaded the sequence to import it into Benchling and be able to view it better.
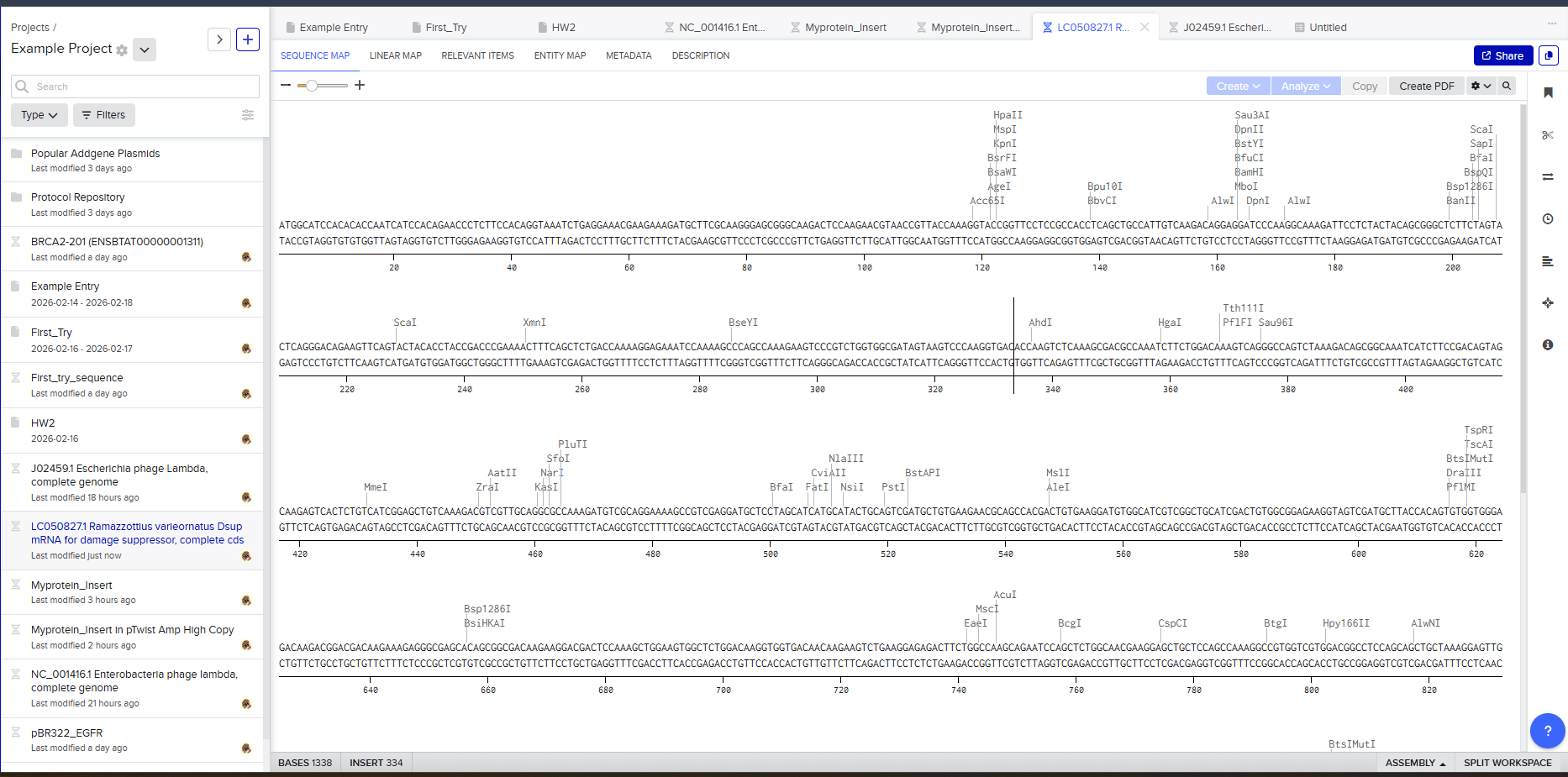

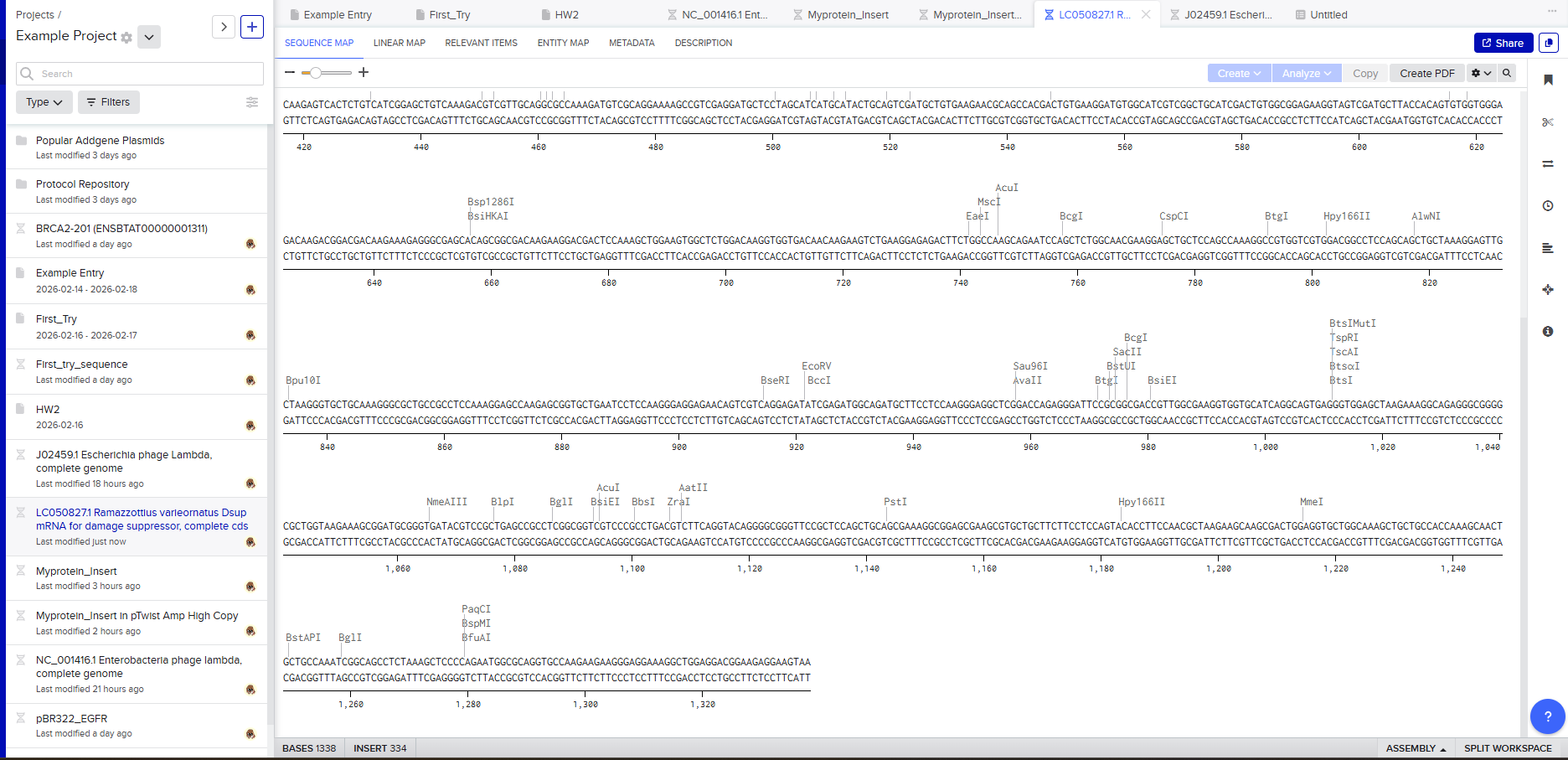

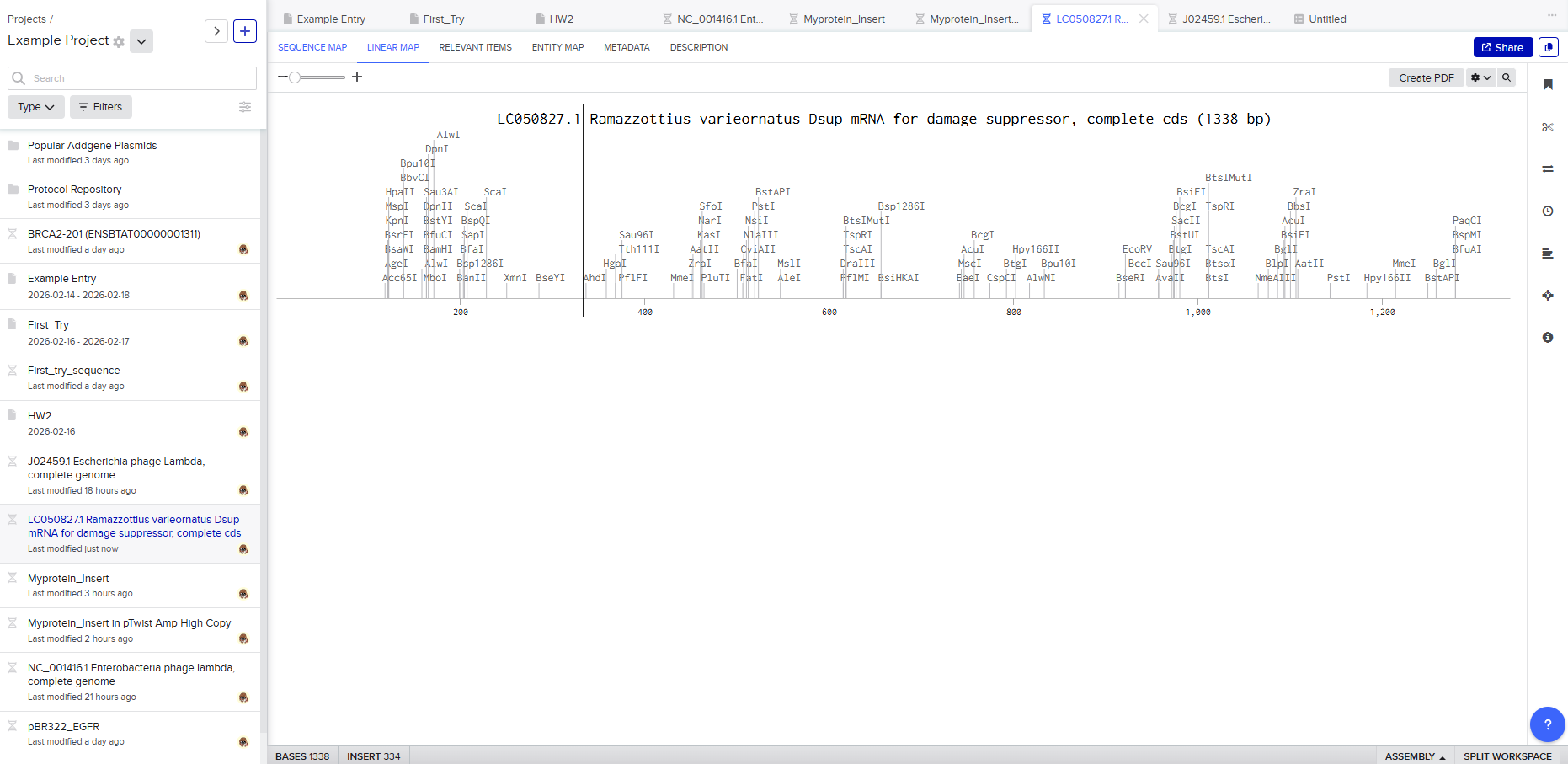

Anyway, if you want to download the complete sequence, you can find it at NIH.

3.2 Reverse translation (protein to DNA)
To express this protein in a laboratory system, the amino acid sequence must be converted into a DNA sequence. Using reverse translation tools based on the genetic code, I generated a nucleotide sequence corresponding to the Dsup protein.
Reverse translation assigns a codon to each amino acid. Because the genetic code is degenerate, meaning that most amino acids are encoded by multiple codons, there are many possible DNA sequences that can produce the same protein. The reverse-translated sequence represents one valid encoding of the protein.
Reverse-translated DNA sequence (excerpt): For this process I used Reverse Translate, in case you want to try it yourself:


This sequence serves as an initial template that can be further optimized for expression in a specific host organism. The result is much longer; you can verify this for yourself (I didn’t know how to put the entire sequence here 😅).
3.3 Codon optimization
Although many DNA sequences can encode the same protein, not all sequences are expressed equally well in every organism. Different species show preferences for certain codons, a phenomenon known as codon bias. If a gene uses rare codons for the host organism, translation can become inefficient, reducing protein yield.
I optimized the Dsup DNA sequence for expression in Escherichia coli, a widely used host in biotechnology. E. coli is preferred because it grows rapidly, is cost-effective, and has a well-characterized genetic system. Codon optimization improves translation efficiency by matching the codon usage to the host’s tRNA abundance.
This optimization enhances protein production by improving ribosome speed and accuracy, increasing mRNA stability, and reducing the likelihood of translation stalling. The resulting sequence is designed to maximize reliable expression in E. coli.

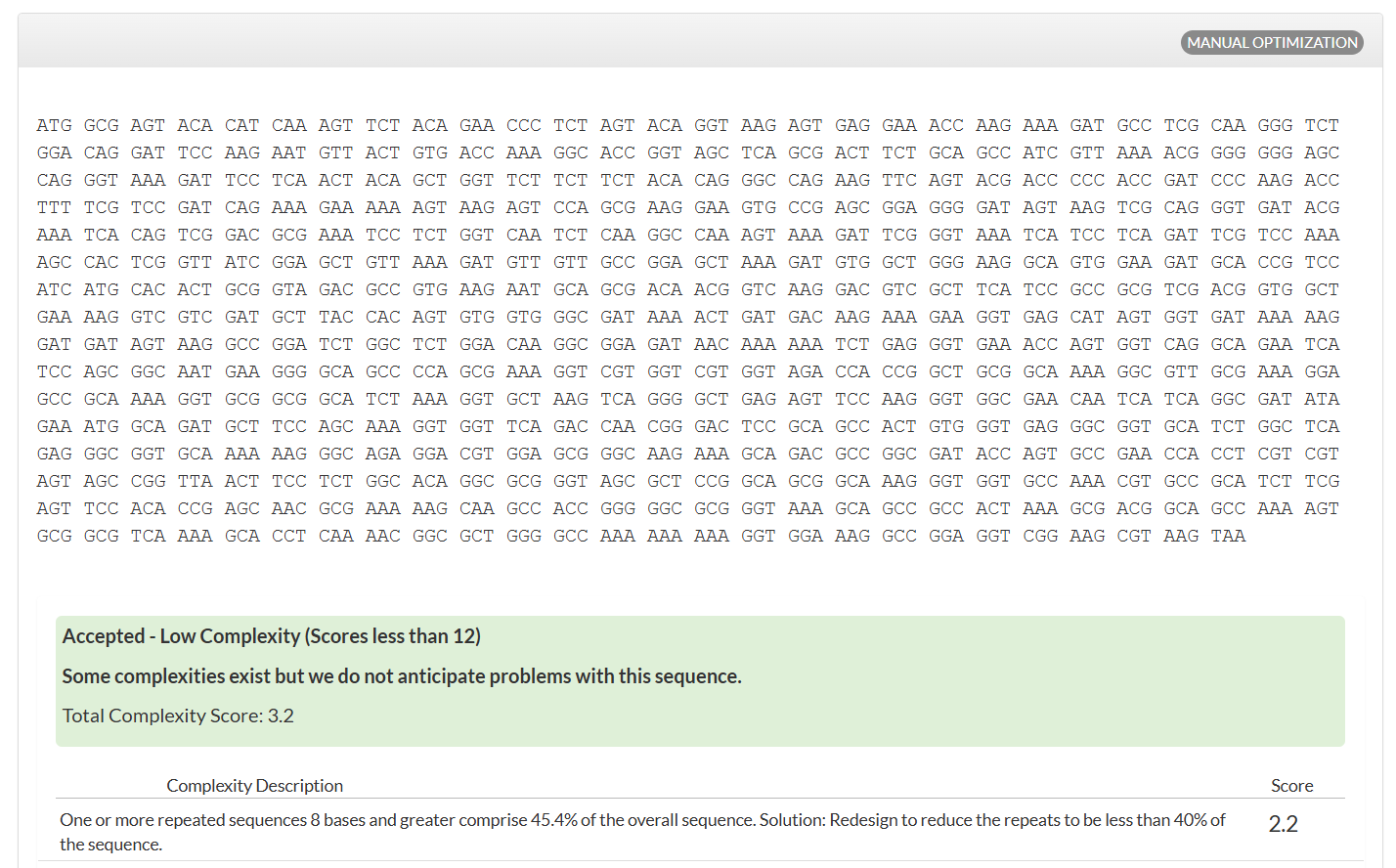

Analyzing optimization in E. coli sparked my curiosity, and I wanted to test how this would work in humans.

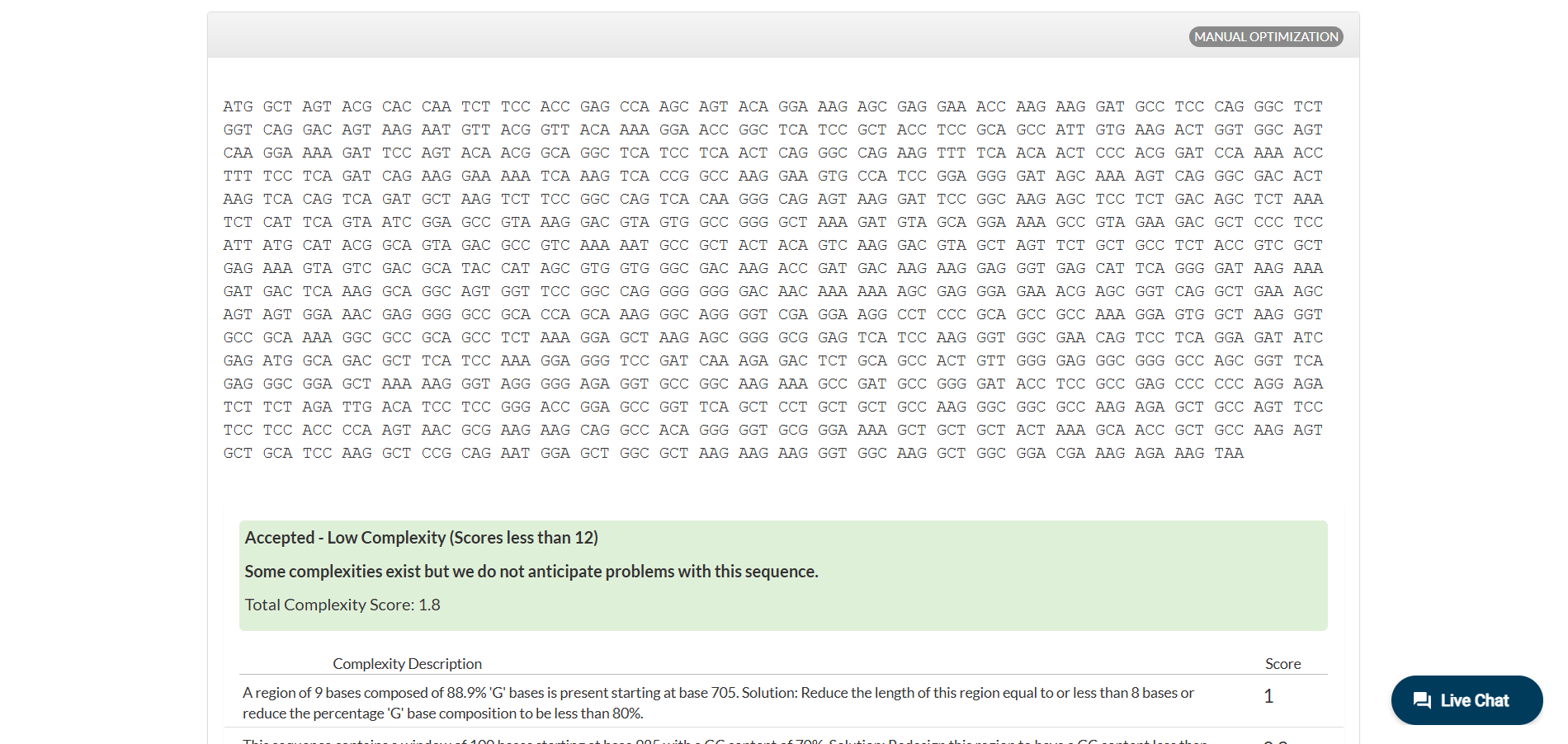

Anyway, there are more cases to analyze different optimizations, which you can see for yourself in IDT (the tool I used for this part).
3.4 You have a sequence. Now what?
Once the codon-optimized DNA sequence is obtained, it can be used to produce the Dsup protein through standard molecular biology techniques.
In a cell-dependent expression system, the DNA is inserted into a plasmid vector and introduced into bacterial cells through transformation. Inside the cell, RNA polymerase transcribes the DNA into messenger RNA. Ribosomes then translate the mRNA into a polypeptide chain, which folds into the functional Dsup protein. This method is commonly used for large-scale protein production in research and industry.
Alternatively, the DNA can be used in a cell-free expression system. These systems contain purified transcription and translation machinery extracted from cells. By adding the DNA template directly to this mixture, proteins can be synthesized rapidly without living cells. Cell-free systems are especially useful for rapid prototyping and synthetic biology applications.
Both approaches follow the central dogma of molecular biology, in which genetic information flows from DNA to RNA and finally to protein.
3.5 Optional: How it works in biological systems
3.5 Optional: How it works in biological systems
In natural biological systems, a single gene can give rise to multiple protein products through several regulatory mechanisms. These include alternative transcription start sites, RNA processing events such as alternative splicing, and post-translational modifications that alter protein function.
A simple example of the central dogma can be illustrated by aligning a short DNA sequence with its RNA transcript and resulting protein.
A short fragment of the Dsup gene illustrates the central dogma of molecular biology. The DNA sequence:
ATG GCA TCC ACA CAC CAA TCA TCC ACA GAA CCC TCT
is transcribed into RNA by replacing thymine with uracil:
AUG GCA UCC ACA CAC CAA UCA UCC ACA GAA CCC UCU
During translation, each codon corresponds to one amino acid, producing the protein fragment:
Met–Ala–Ser–Thr–His–Gln–Ser–Ser–Thr–Glu–Pro–Ser.
Each group of three nucleotides, called a codon, specifies one amino acid. During transcription, thymine is replaced by uracil in RNA. During translation, ribosomes read these codons to assemble the corresponding amino acid sequence, demonstrating how genetic information is converted into functional proteins.
Part 4: Prepare a Twist DNA Synthesis Order This is a practice exercise, not necessarily your real Twist order!
4.1. Create a Twist account and a Benchling account 4.2. Build Your DNA Insert Sequence
For example, let’s make a sequence that will make E. coli glow fluorescent green under UV light by constitutively (always) expressing sfGFP (a green fluorescent protein):
In Benchling, select New DNA/RNA sequence
Give your insert sequence a name and select DNA with a Linear topology (this is a linear sequence that will be inserted into a circular backbone vector of our choosing).
Go through each piece of the given DNA sequences highlighted below (Promoter, RBS, Start Codon, Coding Sequence, His Tag, Stop Codon, Terminator) and paste the sequences into the Benchling file one after the other (replacing the coding sequence with your codon optimized DNA sequence of interest!). Each time you add a new piece of the sequence, make sure to annotate by right clicking over the sequence and creating an annotation that describes what each piece (e.g., Promoter, RBS, etc.) is (see image below).
Promoter (e.g. BBa_J23106): TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGC RBS (e.g. BBa_B0034 with spacers for optimal expression): CATTAAAGAGGAGAAAGGTACC Start Codon: ATG Coding Sequence (your codon optimized DNA for a protein of interest, sfGFP for example): AGCAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTTGTTGAATTAGATGGTGATGTTAATGGGCACAAATTTTCTGTCCGTGGAGAGGGTGAAGGTGATGCTACAAACGGAAAACTCACCCTTAAATTTATTTGCACTACTGGAAAACTACCTGTTCCGTGGCCAACACTTGTCACTACTCTGACCTATGGTGTTCAATGCTTTTCCCGTTATCCGGATCACATGAAACGGCATGACTTTTTCAAGAGTGCCATGCCCGAAGGTTATGTACAGGAACGCACTATATCTTTCAAAGATGACGGGACCTACAAGACGCGTGCTGAAGTCAAGTTTGAAGGTGATACCCTTGTTAATCGTATCGAGTTAAAGGGTATTGATTTTAAAGAAGATGGAAACATTCTTGGACACAAACTCGAGTACAACTTTAACTCACACAATGTATACATCACGGCAGACAAACAAAAGAATGGAATCAAAGCTAACTTCAAAATTCGCCACAACGTTGAAGATGGTTCCGTTCAACTAGCAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCCTGTCCTTTTACCAGACAACCATTACCTGTCGACACAATCTGTCCTTTCGAAAGATCCCAACGAAAAGCGTGACCACATGGTCCTTCTTGAGTTTGTAACTGCTGCTGGGATTACACATGGCATGGATGAGCTCTACAAA 7x His Tag (Let’s add a 7×His tag at the C-terminus of the protein to enable protein purification from E. coli): CATCACCATCACCATCATCAC Stop Codon: TAA Terminator (e.g. BBa_B0015): CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA Once you’ve completed this, click on Linear Map to preview the entire sequence. If you intend to have a TA review a sequence in the future, this is a good way to verify that all sections are annotated!
This is not required for this exercise, but to share your design with others, please ensure that link sharing is turned on!(Optional) Share your final sequence link with a TA for review!
This insert sequence you built is commonly referred to as an expression cassette in molecular biology (a sequence you can drop into any vector and it’ll perform its function). Go ahead and download the FASTA file for the sequence you made.
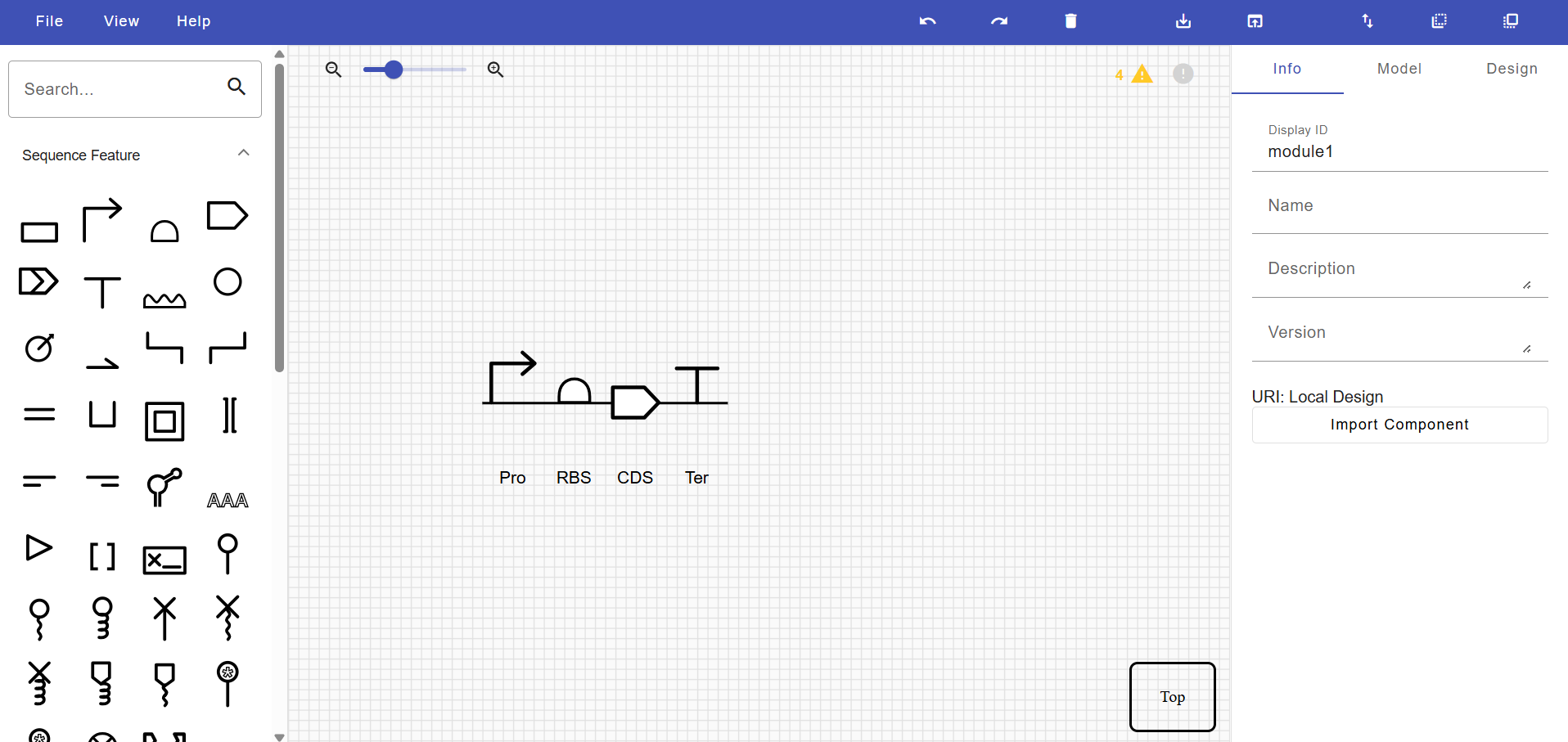
It’s helpful to visualize DNA designs using SBOL Canvas (Synthetic Biology Open Language) to convey your designs. Here’s an example of what you just annotated in Benchling:
4.3. On Twist, Select The “Genes” Option
4.4. Select “Clonal Genes” option
For this demonstration, we’ll choose Clonal Genes. You’ll select clonal genes or gene fragments depending on your final project.
Historically, HTGAA projects using clonal genes (circular DNA) have reached experimental results 1-2 weeks quicker because they can be transformed directly into E. coli without additional assembly.
Gene fragments (linear DNA) offer greater design flexibility but typically require an assembly or cloning step prior to transformation. An advantage is If designed with the appropriate exonuclease protection, gene fragments can be used directly in cell-free expression.
4.5. Import your sequence
You just took an amino acid sequence of interest and converted it into DNA, codon optimized it, and built an expression cassette around it! Choose the Nucleotide Sequence option and Upload Sequence File to upload your FASTA file.
4.6. Choose Your Vector
Since we’re ordering a clonal gene, you will need to refer to Twist’s Vector Catalog to choose your circular backbone. You can think of this as taking your linear expression cassette for your protein of interest, and completing the rest of the circle!
The backbone confers many special properties like antibiotic resistance, an origin of replication, and more. Discuss with your node to decide on appropriate antibiotic options. At MIT/Harvard, you can use Ampicillin, Chloramphenicol, or Kanamycin resistance.
Twist vectors do not contain restriction sites near the insert fragment, so make sure to flank your design with cut sites if you are intending to extract this DNA insert fragment later.
For this demonstration, choose a Twist cloning vectors like pTwist Amp High Copy.
Click into your sequence and select download construct (GenBank) to get the full plasmid sequence:
Go back to your Benchling account. Inside of a folder, click the import DNA/RNA sequence button and upload the GenBank file you just downloaded.
This is the plasmid you just built with your expression cassette included. Congratulations on building your first plasmid!
Part 4: Preparing a Twist DNA Synthesis Order
This exercise simulates the workflow used in modern synthetic biology to design and order custom DNA. Although this is a practice exercise, it mirrors the real process researchers use to synthesize genes for experimental work.
4.1 Creating Twist and Benchling Accounts
The first step is creating accounts on Twist Bioscience and Benchling. These platforms serve complementary roles in DNA engineering.
Benchling functions as a digital molecular biology workspace where DNA sequences can be designed, edited, and annotated. It allows researchers to simulate genetic constructs before ordering them.
Twist Bioscience is a commercial DNA synthesis provider. Once a sequence is finalized in Benchling, it can be uploaded to Twist for physical synthesis.
Creating these accounts establishes the digital pipeline from design to manufacturing.

I hope you like it :)
4.2 Building the DNA Insert Sequence
The goal of this section is to construct an expression cassette — a functional DNA unit that produces a protein inside a host organism.
In Benchling, a new linear DNA sequence is created. The topology is set to linear because this insert will later be placed inside a circular plasmid vector.
The sequence is built from modular components:
Promoter: initiates transcription. It controls how strongly the gene is expressed.
Ribosome Binding Site (RBS): ensures efficient translation by recruiting ribosomes.
Start Codon (ATG): signals the beginning of protein synthesis.
Coding Sequence: contains the codon-optimized gene of Dsup, in my case.
7× His Tag: adds histidine residues to allow protein purification.
Stop Codon: terminates translation.
Terminator: stops transcription and stabilizes mRNA.
Each component is pasted sequentially and annotated in Benchling. Annotation is critical because it documents the function of each region and makes the design interpretable to collaborators.
The final annotated construct represents a complete gene expression system. Viewing the Linear Map confirms the structural organization and ensures no sections are missing.
Exporting the sequence as a FASTA file prepares it for DNA synthesis.
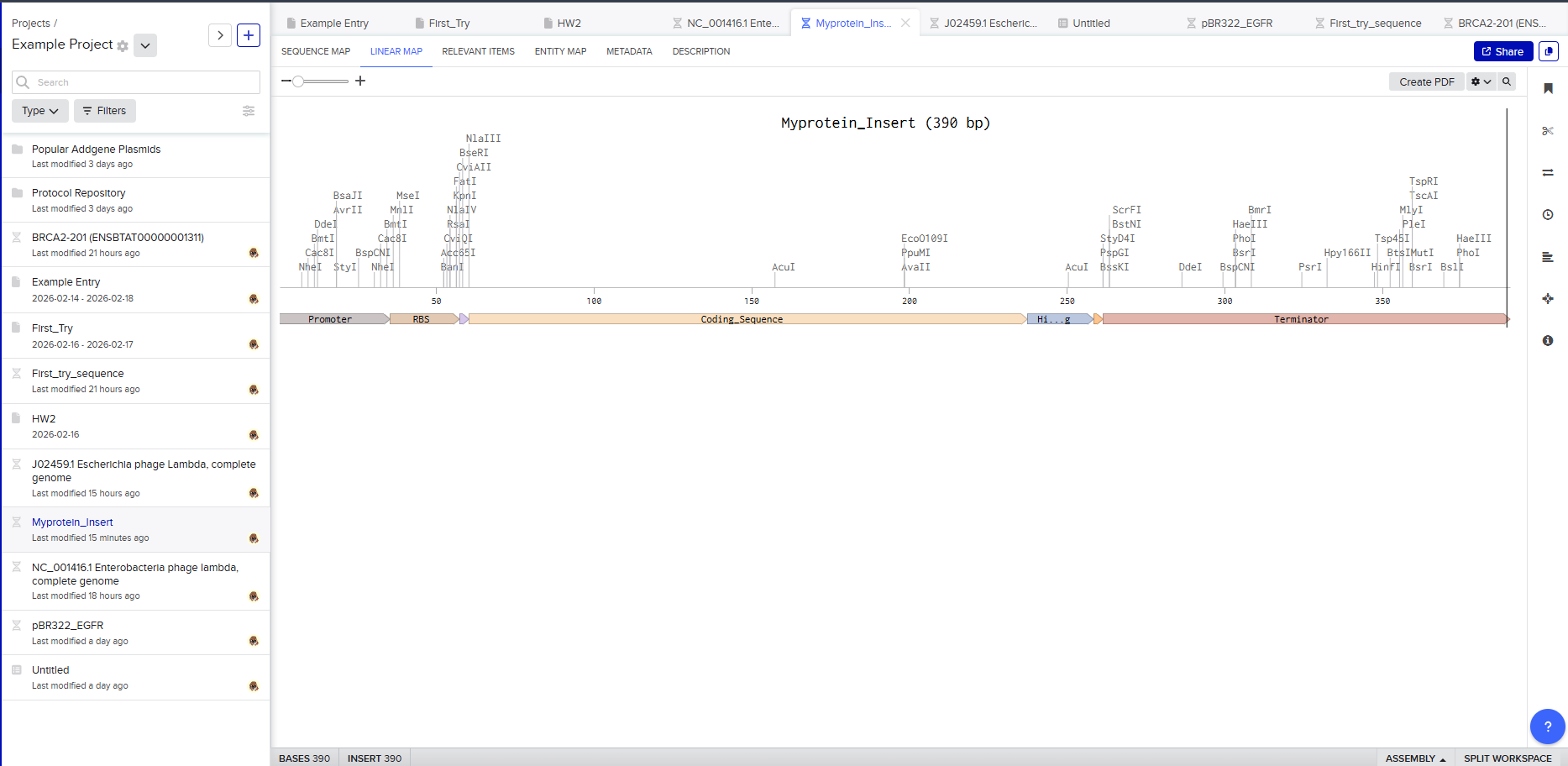

Expression Cassette Concept
The constructed insert is called an expression cassette because it can function independently once inserted into a plasmid. This modular design allows the same cassette to be reused in different vectors or host organisms.
Visualization with SBOL Canvas helps communicate the design using standardized synthetic biology symbols. I don’t know why, but I like this part. I love the SBOL Canvas interface; I think it’s simply because it’s simple. I would like to use more of this interface.

4.3 Selecting the “Genes” Option in Twist
Inside Twist’s ordering interface, selecting the Genes category specifies that a full gene construct is being synthesized rather than short oligonucleotides.
4.4 Choosing Clonal Genes
Clonal genes are circular plasmids delivered ready for transformation into bacteria. This option accelerates experimentation because no additional cloning is required.
In contrast, gene fragments are linear DNA pieces that require assembly before use. While more flexible, they add extra laboratory steps.
Choosing clonal genes prioritizes speed and simplicity.
4.5 Importing the Sequence
The FASTA file exported from Benchling is uploaded to Twist. This step transfers the digitally designed expression cassette into the manufacturing platform.
At this stage, Twist verifies the sequence for synthesis compatibility.
4.6 Choosing a Vector
A vector is a circular DNA backbone that carries the insert into host cells. It contains essential features such as:
an origin of replication (for plasmid copying),
antibiotic resistance markers (for selection),
cloning regions.
Selecting a vector like pTwist Amp High Copy determines how the plasmid behaves inside E. coli.
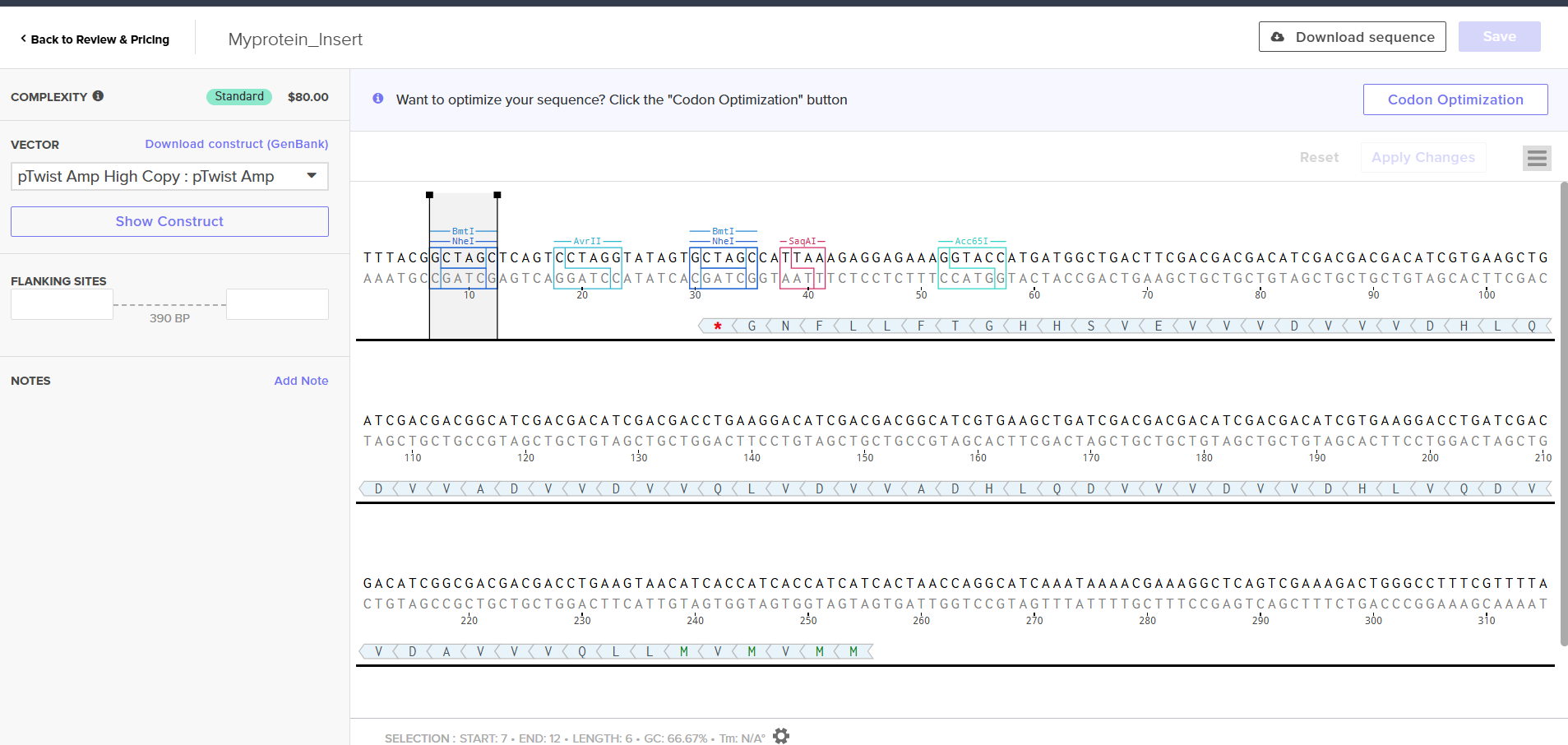

Downloading the full plasmid sequence and re-importing it into Benchling allows visualization of the final construct: the insert integrated into the backbone.
This confirms successful plasmid design.
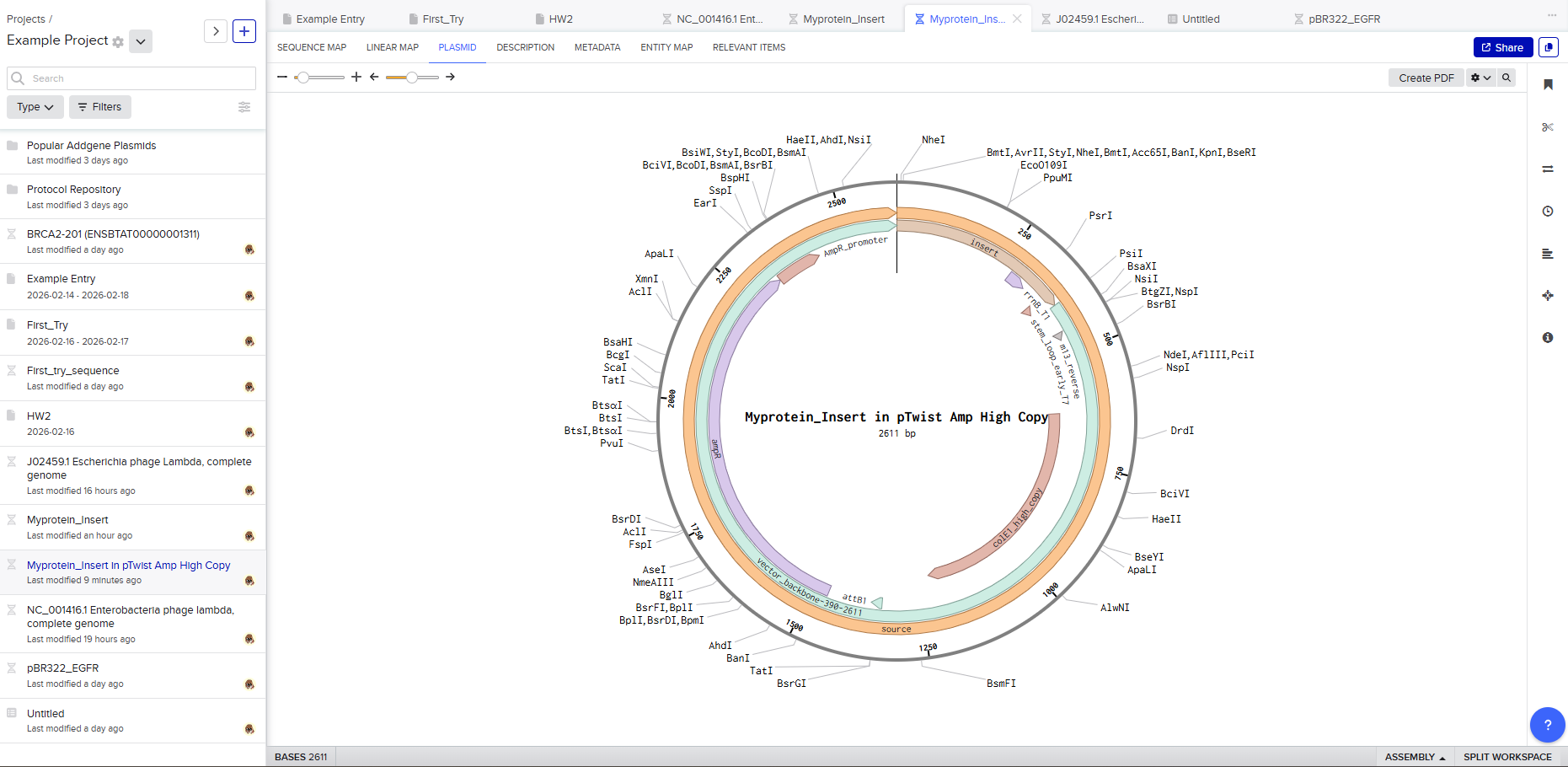

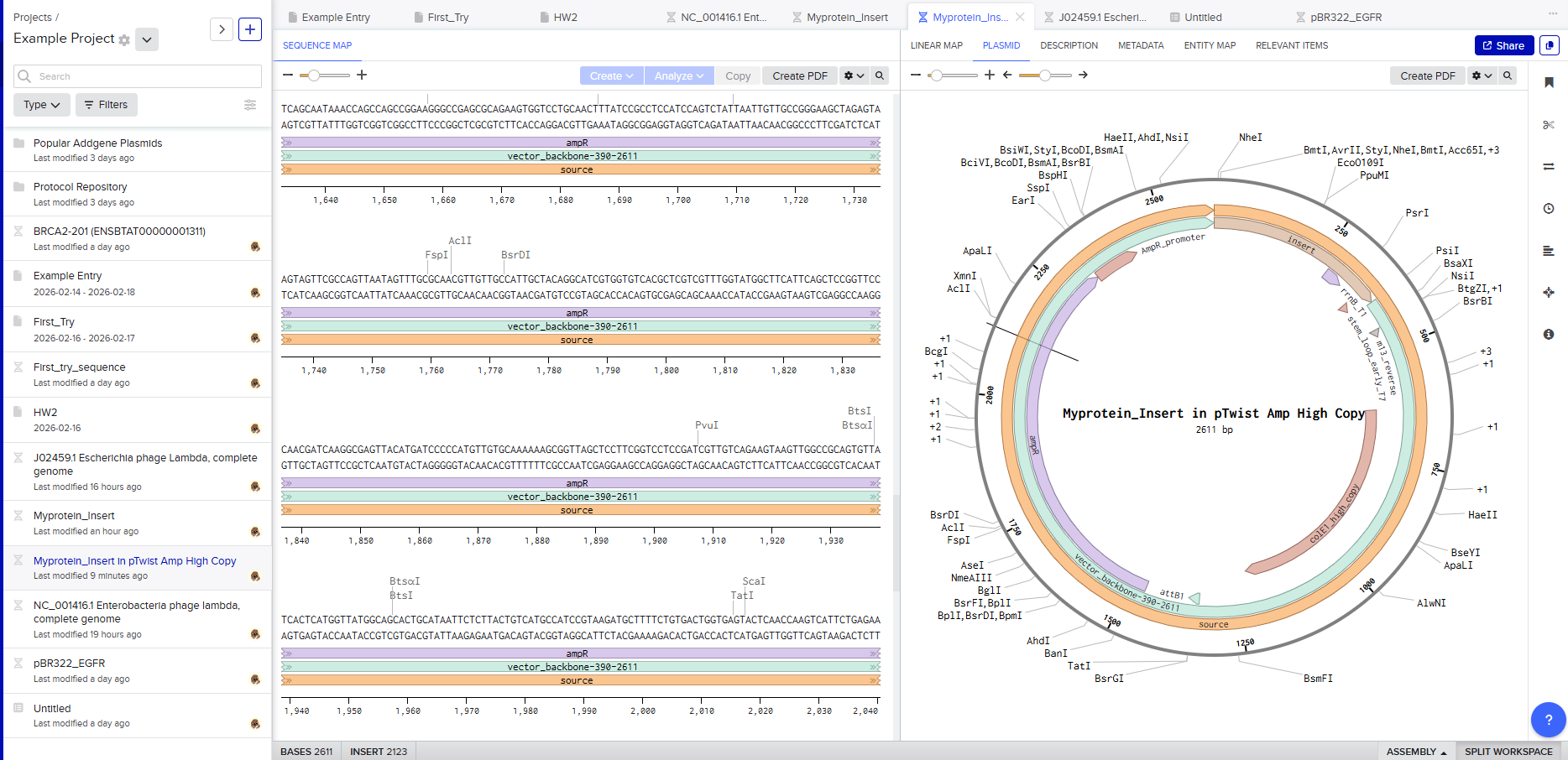

Final Outcome
By the end of this exercise, a fully annotated plasmid construct has been digitally assembled. This workflow demonstrates the complete pipeline of modern DNA engineering:
design → annotation → synthesis preparation → plasmid assembly.
For final projects, both the annotated insert and chosen vector must be clearly documented to ensure reproducibility and successful DNA synthesis.
Part 5: DNA Read/Write/Edit 5.1 DNA Read (i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? Also answer the following questions:
Is your method first-, second- or third-generation or other? How so? What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps. What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)? What is the output of your chosen sequencing technology? 5.2 DNA Write (i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
See some famous examples of DNA design
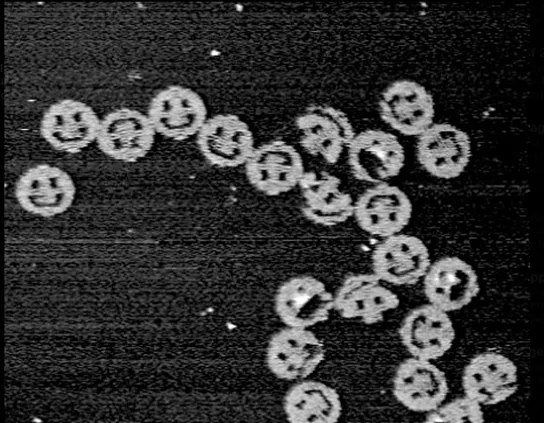
(ii) What technology or technologies would you use to perform this DNA synthesis and why? Also answer the following questions:
What are the essential steps of your chosen sequencing methods? What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability? 5.3 DNA Edit (i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?

(ii) What technology or technologies would you use to perform these DNA edits and why? Also answer the following questions:
How does your technology of choice edit DNA? What are the essential steps? What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing? What are the limitations of your editing methods (if any) in terms of efficiency or precision?
5.1 DNA Read
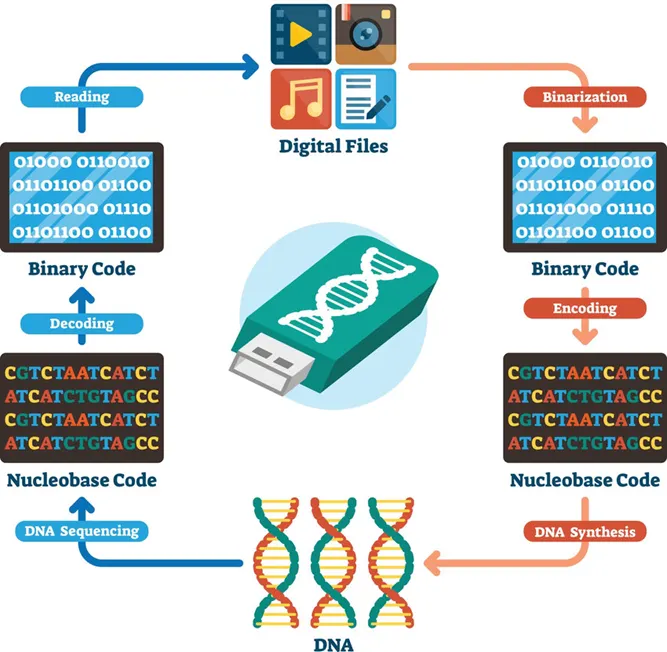
For DNA sequencing, I would choose to read DNA used in DNA-based digital data storage. This technology encodes digital information such as images, text, or scientific data into synthetic DNA molecules. I am interested in sequencing this type of DNA because it represents a bridge between biology and computer science, with the potential to create extremely dense, long-term archival storage systems. DNA is far more stable than conventional storage media and could preserve information for thousands of years. Sequencing stored DNA is essential to verify that the encoded information has not degraded and can be accurately retrieved.
To sequence this DNA, I would use next-generation sequencing (NGS), specifically sequencing-by-synthesis technology developed by Illumina. This method is considered a second-generation sequencing technology because it enables massively parallel sequencing of millions of DNA fragments simultaneously, unlike first-generation Sanger sequencing which reads one fragment at a time.
The input for this method is purified DNA containing encoded data. The preparation steps include fragmenting the DNA into short pieces, attaching adapter sequences to both ends, amplifying the fragments using PCR, and immobilizing them on a flow cell. During sequencing, fluorescently labeled nucleotides are added one base at a time. A camera records the fluorescence emitted as each base is incorporated. Specialized software converts these signals into a nucleotide sequence through a process called base calling.
The output of this technology is a large dataset of short DNA reads in digital format. These reads are assembled computationally to reconstruct the original encoded information. This approach provides high accuracy and scalability, which are critical for reliable data retrieval in DNA storage systems.
5.2 DNA Write
For DNA synthesis, I would design a genetic circuit encoding a radiation-protective protein system, inspired by extremophile organisms. Specifically, I would synthesize a codon-optimized gene encoding the Dsup protein along with regulatory elements that allow controlled expression in bacteria. This DNA could be used to study how protective proteins improve cellular resistance to radiation, which has applications in medicine and space exploration.
An example short segment of the synthesized DNA sequence could look like this:
ATGTCCGACCAGTCCCAGAAGCAGGAGAAGCTGAAGGAGGAGCTGAAGGCCAAGAAG
To synthesize this DNA, I would use commercial gene synthesis technology from Twist Bioscience. This technology relies on high-throughput chemical DNA synthesis using phosphoramidite chemistry and microarray-based oligonucleotide assembly.
The essential steps include chemical synthesis of short oligonucleotides, enzymatic assembly into longer fragments, error correction, and cloning into plasmid vectors. These fragments are then amplified and sequence-verified.
The main limitations of this synthesis method include potential synthesis errors in long sequences, cost for very large constructs, and technical limits on maximum fragment length. However, it offers excellent scalability and precision for gene-level synthesis.
5.3 DNA Edit
For DNA editing, I would focus on modifying genes that improve cellular resistance to radiation damage, similar to research being explored by companies such as Colossal Biosciences in the context of advanced genetic engineering. Editing such genes could have applications in protecting human cells during radiation therapy or long-duration space missions.
To perform these edits, I would use CRISPR-Cas9 genome editing technology. CRISPR works by using a guide RNA to direct the Cas9 enzyme to a specific DNA sequence. Cas9 creates a targeted double-strand break, and the cell’s repair machinery introduces modifications during the repair process.
The essential preparation steps include designing a guide RNA that matches the target sequence, constructing a plasmid or delivery system carrying Cas9 and the guide RNA, and introducing these components into cells. The inputs include the DNA template, Cas9 enzyme, guide RNA, and host cells.
The main limitations of CRISPR editing include off-target effects, incomplete editing efficiency, and challenges in delivering the editing machinery into certain cell types. Despite these limitations, CRISPR remains one of the most powerful and precise genome editing tools available.
 GammaShroom
GammaShroom
I hope you haven’t forgotten about my project proposed in HW1. If you don’t know what I’m talking about, take a look at HW1; it’s above WEEk2. Anyway, I mention this because I’d like to talk about how HW2 could help you better understand how to implement what we saw in HW1. HW2 extends the conceptual ideas introduced in the “gammashroom” proposal from HW1 by translating them into the theoretical and computational foundations of modern genetic engineering, even in the absence of a physical laboratory. While the node did not perform wet-lab experiments, the simulation and design components of HW2 still develop the core competencies required to engineer biological systems like “gammashroom”. By studying how restriction enzymes selectively modify DNA and how virtual gel electrophoresis predicts fragment patterns, we learn how engineered genetic constructs can be analyzed and validated in silico before any real-world implementation. This type of predictive modeling is a critical first step in synthetic biology, where careful planning and verification reduce experimental uncertainty.
More importantly, the DNA read/write/edit framework explored in HW2 directly supports the long-term development of engineered organisms capable of radiation resistance and environmental adaptation. Designing codon-optimized genes, selecting expression systems, and understanding how DNA can be precisely modified provide the technical roadmap for implementing protective genetic features similar to those envisioned in the gammashroom system. Even without executing the laboratory protocol, engaging with these workflows conceptually builds an understanding of how engineered DNA moves from digital design to functional biological systems. In this way, HW2 bridges the gap between speculative bioengineering concepts and the structured methodology required to realize them, reinforcing how computational design and molecular planning underpin any future experimental work.
Prompt used for the task
If you saw my HW1, you’ll have noticed that I also included some of the prompts I used to complete the task. I do this to show that AI is a very useful tool for supporting projects, and it’s something that personally helps me a lot to organize myself much better.
For the homework: “Please organize and synthesize the following information from my assignment (Part 3: DNA Design Challenge and Part 5: DNA Read/Write/Edit) into a clear, structured academic format.
Your goals are:
Group related concepts into logical sections and subsections
Remove redundancy while preserving all important scientific details
Use clear headings and transitions between ideas
Maintain scientific accuracy and an academic tone
Add short explanations that connect concepts when needed
Highlight key technical terms (DNA sequencing, codon optimization, gene expression, etc.)”
“Please rewrite the following scientific text to improve clarity, flow, and academic quality.
Your goals are:
Use more precise scientific vocabulary and appropriate synonyms
Improve sentence structure and transitions
Maintain the original meaning and technical accuracy
Avoid unnecessary repetition
Use a formal academic tone suitable for a university assignment
Keep explanations clear and accessible
Expand brief sections slightly if needed to improve coherence
Do not add new scientific claims — only refine and strengthen the writing.”
For the picture (Gemini):
“Hello, please examine the image I provided. It represents a DNA sequence modified by restriction enzyme digestion, producing a distinct band pattern. Could you generate an artistic image inspired by the visual structure and composition of this pattern?”