Week 11 — Bioproduction & Cloud Labs
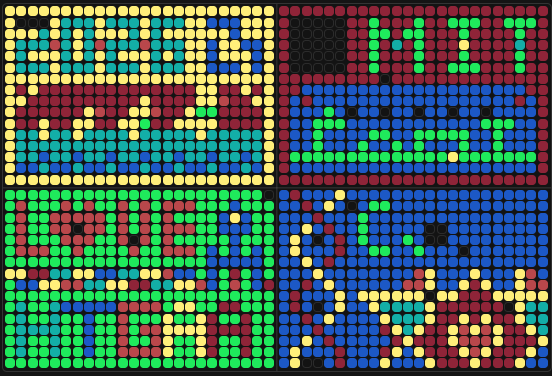
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
Contribute at least one pixel to this global artwork experiment before the editing ends on Sunday 4/19 at 11:59 PM EST.
A personalized URL was sent to the email address associated with your Discourse account, and you can discuss the artwork on the Discourse.
If you did not have a chance to contribute, it’s okay, just make sure you become a TA this fall! 😉
Make a note on your HTGAA webpages including:
what you contributed to the community bioart project (e.g., “I made part of the DNA on the bottom right plate”)
what you liked about the project, and
what about this collaborative art experiment could be made better for next year.
Collective Artwork
1. What I contributed to the community bioart project
Unfortunately, I did not participate in the pixel editing activity. I was unable to contribute because I was in the middle of final exams and other academic commitments during the submission window (April 19 deadline). I did not have time to review the activity or log in to contribute a pixel.
Even though I could not contribute directly, I followed the project outcome and deeply appreciate the effort that went into organizing it.
2. What I liked about the project
What I loved most about this project is how it brought people together from all over the world — different countries, time zones, cultures, and backgrounds — working toward a shared creative and scientific goal.
The project created a real feeling of community. Seeing how everyone coordinated, helped each other, and gave feedback to improve the experiment was inspiring. There are not many activities that manage to unite so many people across such distances in a single collaborative artwork. This was special.
The project also combined synthetic biology, art, automation, and community participation. That mix is exactly what HTGAA is about. Despite the complexity (hundreds of people, cloud lab coordination, DNA templates, fluorescent proteins), everything ran smoothly.
In short, what I liked most was the human connection — the friendship, shared purpose, and the freedom to build something together beyond what any single person could do alone.
3. What about this collaborative art experiment could be made better for next year
First, I want to say that the project is already very well organized. I don’t see major flaws. However, here are a few small suggestions to make it even better:
Extend the editing window or add a second participation round. Many students (like me) have exam periods or overlapping deadlines. A longer window (for example, two weeks instead of one) or a “catch-up weekend” would allow more people to join.
Create a small second phase where latecomers can add a few extra pixels or help analyze the results. This would make the project feel more inclusive without disrupting the main experiment.
Add a live world map showing where each pixel contribution is coming from in real time. Seeing your pixel next to someone’s from the other side of the planet would make the global collaboration more visible and exciting.
Host one optional synchronous “pixel party” (for example, a one-hour Zoom call) where people can contribute together, ask questions, and meet others participating. This would strengthen the sense of community even more.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
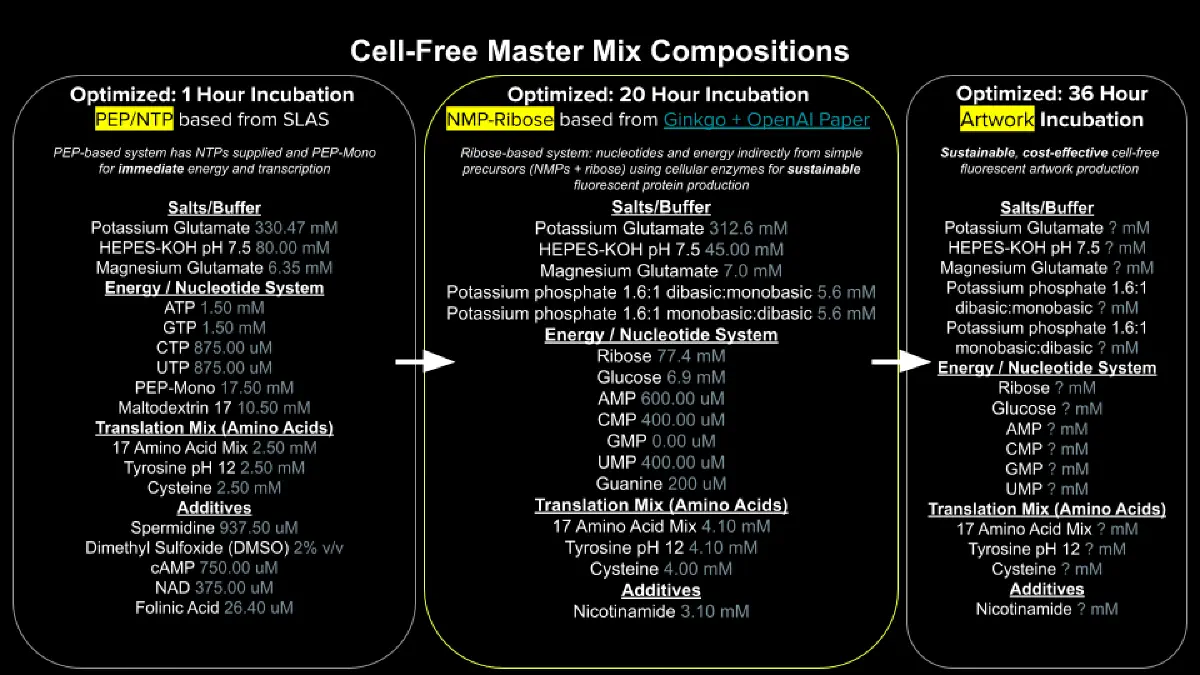
Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
E. coli Lysate
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) Salts/Buffer
Potassium Glutamate
HEPES-KOH pH 7.5
Magnesium Glutamate
Potassium phosphate monobasic
Potassium phosphate dibasic
Energy / Nucleotide System
Ribose
Glucose
AMP
CMP
GMP
UMP
Guanine
Translation Mix (Amino Acids)
17 Amino Acid Mix
Tyrosine
Cysteine
Additives
Nicotinamide
Backfill
Nuclease Free Water
Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
Bonus question: How can transcription occur if GMP is not included but Guanine is?
Cell-Free Protein Synthesis — Cell-Free Reagents
1. Role of each component in the cell-free reaction
E. coli Lysate (BL21 (DE3) Star Lysate, includes T7 RNA Polymerase)
The lysate provides all the endogenous machinery needed for transcription and translation, including ribosomes, tRNAs, initiation factors, elongation factors, and termination factors. The BL21 (DE3) strain is engineered to express T7 RNA polymerase, which specifically recognizes the T7 promoter on the DNA template, enabling high-yield transcription of the target gene.
Salts/Buffer (Potassium Glutamate, HEPES-KOH pH 7.5, Magnesium Glutamate, Potassium phosphate monobasic, Potassium phosphate dibasic)
These maintain optimal pH (7.5) and ionic strength for enzymatic activity. Potassium and magnesium are essential cofactors for ribosome function, RNA polymerase activity, and proper protein folding. The phosphate system helps regenerate ATP and maintains energy homeostasis throughout the reaction.
Energy / Nucleotide System (Ribose, Glucose, AMP, CMP, GMP, UMP, Guanine)
This system provides both the energy currency (ATP, GTP, etc.) and the nucleotide building blocks for RNA synthesis. Glucose and ribose are metabolized to generate ATP via glycolysis and the pentose phosphate pathway. AMP, CMP, GMP, UMP are converted to their triphosphate forms (ATP, CTP, GTP, UTP) by endogenous kinases. Guanine serves as a precursor for GTP synthesis through the salvage pathway.
Translation Mix (17 Amino Acid Mix + Tyrosine + Cysteine)
This provides all 20 amino acids required for protein synthesis. The 17-amino acid mix contains the standard set, while tyrosine and cysteine are added separately because they are often limiting or prone to degradation. Without all 20 amino acids, the ribosome would stall during translation.
Additives (Nicotinamide)
Nicotinamide is a NAD+ precursor that helps maintain redox balance and energy metabolism. It also inhibits certain nucleases and proteases that could degrade the DNA template or the synthesized protein, thereby extending the reaction lifetime.
Backfill (Nuclease Free Water)
Nuclease-free water is used to bring the reaction to the final volume without introducing contaminating nucleases (RNases or DNases) that would degrade the DNA template, mRNA, or tRNA, destroying the reaction.
2. Main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix
The PEP-NTP master mix uses pre-charged nucleotide triphosphates (NTPs) and phosphoenolpyruvate (PEP) as a rapid energy source, enabling fast protein synthesis that reaches peak fluorescence within about 1 hour. In contrast, the NMP-Ribose-Glucose master mix uses nucleotide monophosphates (NMPs) plus ribose and glucose, which must be converted to NTPs through endogenous metabolic pathways, resulting in slower energy release that sustains protein synthesis for up to 20 hours. The NMP-based mix is therefore better for long-term experiments, while the PEP-NTP mix is better for quick results.
3. Bonus question: How can transcription occur if GMP is not included but Guanine is?
Transcription can still occur because guanine is converted to GMP by the enzyme guanine phosphoribosyltransferase (also called hypoxanthine-guanine phosphoribosyltransferase, HPRT), which transfers a phosphoribosyl group from phosphoribosyl pyrophosphate (PRPP) to guanine, producing GMP. Once GMP is formed, cellular kinases (GMP kinase and nucleoside diphosphate kinase) phosphorylate it to GDP and then to GTP, which is the direct substrate for RNA polymerase during transcription. The cell-free lysate contains these endogenous salvage pathway enzymes, so guanine can serve as the starting point for GTP synthesis even when GMP is not directly provided.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
sfGFP
mRFP1
mKO2
mTurquoise2
mScarlet_I
Electra2
The amino acid sequences are shown in the HTGAA Cell-Free Benchling folder.
Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here.
Important In order to be eligible for this, make sure that your final project slide is in the “2026 Committed Listener ONE FINAL PROJECT IDEA” slide deck.
The final phase of this lab will be analyzing the fluorescence data we collect to determine whether we can draw any conclusions about favorable reagent compositions for our fluorescent proteins. This will be due a week after the data is returned (date TBD!). The reaction composition for each well will be as follows:
6 μL of Lysate 10 μL of 2X Optimized Master Mix from above 2 μL of assigned fluorescent protein DNA template 2 μL of your custom reagent supplements Total: 20 μL reaction
Planning the Global Experiment — Cell-Free Master Mix Design
1. Biophysical or functional property of each fluorescent protein that affects expression or readout in cell-free systems
sfGFP (superfolder GFP)
sfGFP has been engineered to fold rapidly and efficiently even when fused to poorly folding partners, which is advantageous in cell-free systems because it minimizes fluorescence loss due to misfolding. However, it still requires molecular oxygen for chromophore maturation, which can be limiting in anaerobic or poorly oxygenated cell-free reactions.
mRFP1 (monomeric red fluorescent protein 1)
mRFP1 matures relatively quickly (around 60 minutes to half-maximal fluorescence) but has a tendency to form dimers at high concentrations, which can affect its solubility and fluorescence readout in crowded cell-free lysates. It is also sensitive to acidic pH, and cell-free reactions can become acidic over time due to metabolic byproducts, potentially reducing its signal.
mKO2 (monomeric Kusabira Orange 2)
mKO2 has an unusually fast maturation rate (approximately 20 minutes to half-maximal fluorescence), making it ideal for short-term cell-free experiments. However, it is moderately sensitive to reducing conditions, and cell-free lysates contain reducing agents like DTT or glutathione that could impair chromophore oxidation if not carefully controlled.
mTurquoise2
mTurquoise2 is a cyan fluorescent protein with high brightness and photostability, but it has a slow maturation time (around 60-90 minutes) and requires complete oxidation of its chromophore to achieve full fluorescence. In cell-free reactions, slow maturation means that fluorescence continues to increase for many hours, which is good for long experiments but problematic for early timepoint measurements.
mScarlet_I
mScarlet_I is one of the brightest monomeric red fluorescent proteins, but it has a relatively long maturation time (approximately 90 minutes to half-maximum) and is prone to aggregation when expressed at high levels. In cell-free systems, protein aggregation can reduce soluble fluorescence and complicate readout, especially at high DNA template concentrations.
Electra2
Electra2 is a recently developed green-yellow fluorescent protein that is highly tolerant of acidic pH and oxidative conditions, which is beneficial for cell-free reactions that may drift in pH over time. However, it has a slower maturation rate compared to sfGFP and requires proper calcium ion balance for optimal chromophore formation, and cell-free lysates may not provide ideal calcium concentrations unless supplemented.
2. Hypothesis for adjusting reagents to improve fluorescence over 36 hours
Protein selected: mTurquoise2
Reagent(s) to adjust: Add a controlled oxygen-releasing system (e.g., glucose oxidase and catalase with slow-release glucose) and increase the concentration of reducing agent scavengers such as oxidized glutathione (GSSG).
Expected effect: mTurquoise2 has a slow maturation time (~60-90 minutes) and requires complete chromophore oxidation for full fluorescence. In a standard cell-free reaction, oxygen is gradually depleted over 36 hours, and reducing conditions can inhibit proper chromophore oxidation. By adding a slow oxygen-release system (glucose oxidase generates hydrogen peroxide, which catalase converts to water and oxygen, slowly replenishing O₂), and by increasing oxidized glutathione to buffer the redox environment, I expect that mTurquoise2 will continue to mature for the full 36-hour incubation rather than plateauing early. This should result in significantly higher maximum fluorescence compared to the unmodified master mix.
3. Experimental design summary (for reference when data is returned)
The reaction composition for each well will be:
6 μL of Lysate
10 μL of 2X Optimized Master Mix
2 μL of assigned fluorescent protein DNA template
2 μL of custom reagent supplements (as hypothesized above)
Total reaction volume: 20 μL
Incubation time: 36 hours
Readout: Fluorescence measured at regular intervals (specific wavelengths for each protein)
Next steps: Once I receive my assigned artwork wells with specific fluorescent proteins (by email by April 24), I will define the precise reagent concentrations for my custom supplements. After the data is collected and returned, I will analyze whether my hypothesis was correct and whether the modified master mix improved fluorescence for mTurquoise2 over the 36-hour incubation.
Part D: Build-A-Cloud-Lab | (optional) Bonus Assignment
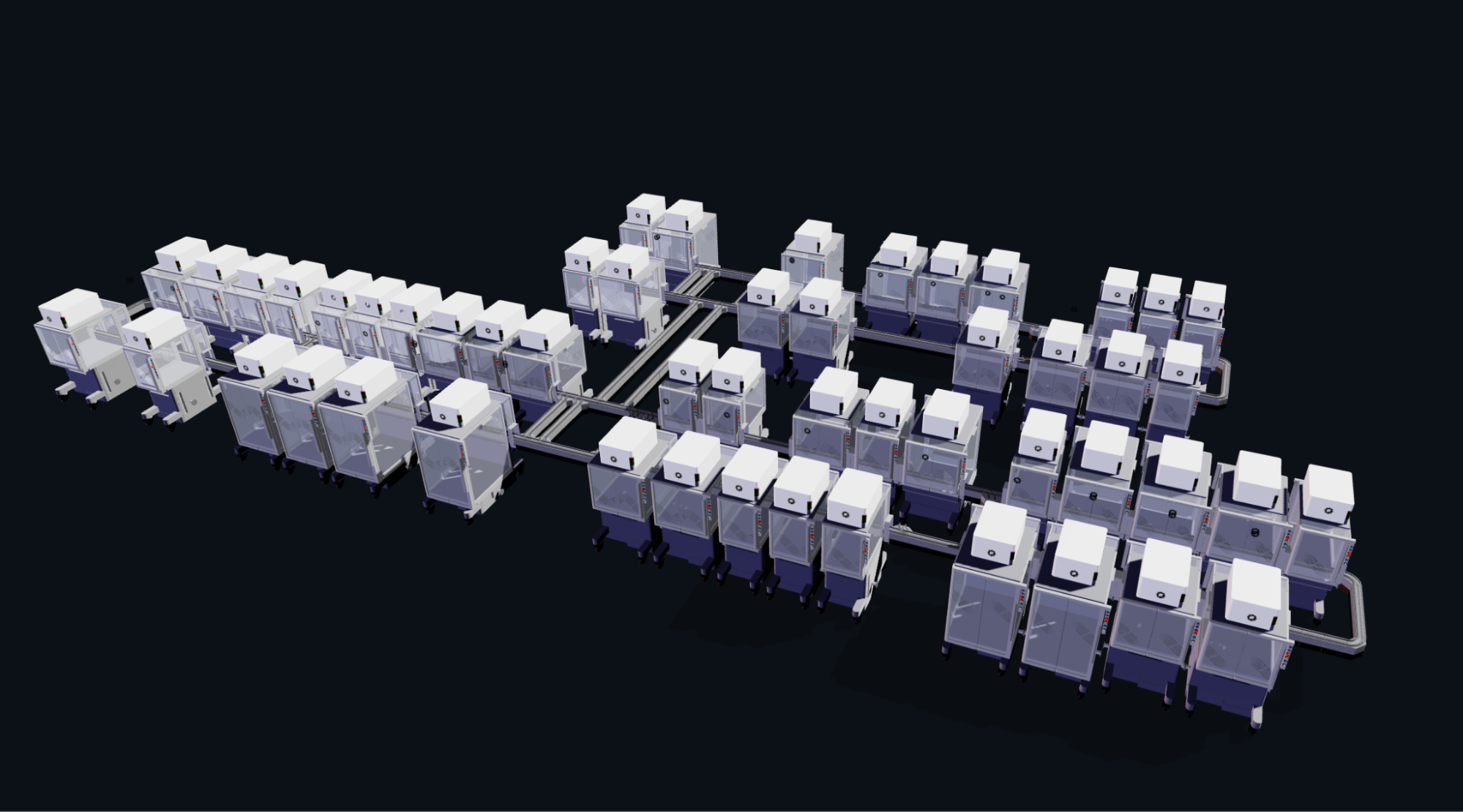

Ginkgo Nebula Cloud Laboratory Rendering, 2025 Ginkgo Nebula Cloud Laboratory Rendering, 2025
Use this simulation tool to create an interesting looking cloud lab out of the Ginkgo Reconfigurable Automation Carts. This is just a minimal implementation so far, but I would love to see some fun designs!
