Week 7 — Genetic Circuits Part II: Neuromorphic Circuits
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs) What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions? Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal. Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
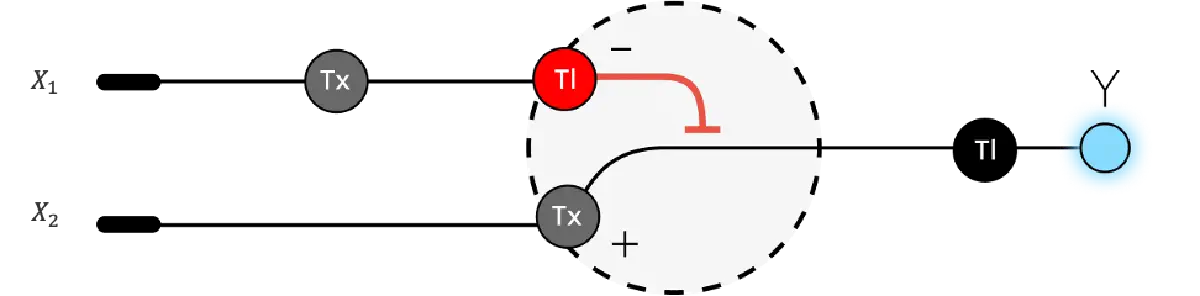
What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
Traditional genetic circuits are built like digital logic gates (AND, OR, NOT). They produce binary outputs—ON or OFF—based on inputs that cross a fixed threshold. Intracellular Artificial Neural Networks (IANNs) take a different approach inspired by biological neurons.
| Feature | Traditional Boolean Genetic Circuits | Intracellular Artificial Neural Networks (IANNs) |
|---|---|---|
| Output type | Binary (ON/OFF) | Continuous (graded, analog) |
| Input integration | Linear; inputs combined via fixed logic (e.g., AND gate) | Weighted summation; each input has a tunable “weight” |
| Nonlinearity | Hard threshold (e.g., repressor titers) | Soft, differentiable activation functions (sigmoidal, similar to neurons) |
| Adaptability | Fixed function; cannot be tuned post-fabrication | Can be trained or tuned by adjusting promoter strengths, ribosome binding sites (RBS), and degradation tags |
| Noise tolerance | Low; small fluctuations can flip the output | High; analog nature averages out molecular noise |
| Function complexity | Complex functions require many parts (multiple gates) | Complex functions can be implemented with fewer parts using weighted summation |
| Biological relevance | Does not mimic natural cellular computation | Mimics how cells naturally integrate multiple signals (e.g., in development, metabolism) |
IANNs allow a cell to make “soft decisions.” For example, instead of a cell producing a drug only when a pathogen is definitively present (Boolean), an IANN could produce the drug in proportion to the severity of infection, conserving resources while still responding effectively.
Describe a useful application for an IANN
Application: Smart Probiotic for Inflammatory Bowel Disease (IBD) Management Goal: Engineer a probiotic bacterium (e.g., E. coli Nissle 1917) that produces an anti-inflammatory drug in proportion to the severity of intestinal inflammation.
Inputs (Biological Signals)
| Input | Molecular Sensor | Biological Meaning |
|---|---|---|
| X₁ | Nitric oxide (NO)-sensitive promoter | NO is produced by immune cells during inflammation; higher concentration = more severe inflammation |
| X₂ | Thiosulfate-sensitive promoter | Thiosulfate is produced by pathogenic bacteria during gut dysbiosis |
| X₃ | pH-sensitive promoter | pH drops during inflammation due to loss of epithelial barrier function |
Processing (Single-Layer Perceptron) Each input is assigned a weight determined by the strength of the promoter and RBS. The weighted sum is computed as:
Z = w₁·[X₁] + w₂·[X₂] + w₃·[X₃]
This weighted sum drives expression of a transcription factor that activates the output gene in a graded, not binary, manner.
Output Behavior
| [X₁] (NO) | [X₂] (Thiosulfate) | [X₃] (pH drop) | Weighted Sum (Z) | Output (Anti-inflammatory drug) |
|---|---|---|---|---|
| Low | Low | Low | < threshold | None |
| Moderate | Low | Low | Moderate | Low dose |
| High | Low | Moderate | High | Medium dose |
| High | High | High | Very high | High dose |
The drug concentration scales with inflammation severity, allowing for adaptive dosing without external monitoring.
Limitations
| Limitation | Explanation |
|---|---|
| Orthogonality | Each sensor must not cross-talk with other cellular processes. Synthetic promoters and engineered transcription factors are needed. |
| Metabolic load | Expressing multiple sensors and a drug biosynthesis pathway can burden the cell, reducing growth and stability. |
| Stochastic noise | Low input levels may produce variable outputs due to gene expression noise. This can be mitigated by using transcriptional amplifiers or negative feedback. |
| Stability in the gut | The probiotic must survive passage through the gastrointestinal tract and maintain the genetic circuit without mutation. |
| Regulatory approval | Genetically engineered probiotics face stringent safety evaluations. |
Draw a diagram for an intracellular multilayer perceptron
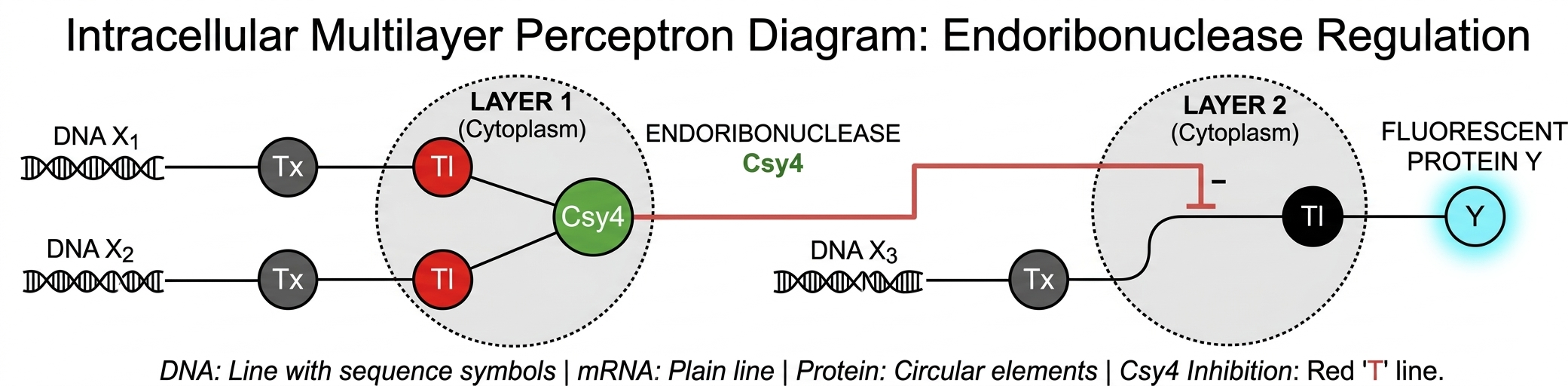

Architectural Breakdown
- Layer 1: Signal Integration and Enzyme Synthesis
The first layer represents the input processing stage. In this biological context:
Genetic Inputs (X1, X2): These are discrete DNA sequences that undergo Transcription (Tx) and Translation (Tl).
Summation/Integration: The system integrates these inputs to produce a specific functional output—the Csy4 endoribonuclease.
Role: In neural network terms, this layer acts as an initial transformation where multiple genetic signals are “compressed” into a single molecular carrier (the enzyme)
- The Inter-layer Link (Molecular Weighting)
The Csy4 produced in Layer 1 does not act as a final output; instead, it functions as a hidden signal. It migrates to the next node where it exerts a regulatory influence. This connection represents the “synapse” between layers, where the presence of the enzyme determines the state of the subsequent node.
- Layer 2: Regulated Output Generation
The second layer governs the final observable phenotype, which is the Fluorescent Protein Y:
Input X3: A separate DNA template is transcribed into mRNA.
Post-transcriptional Regulation: This is the critical “decision” point. The Csy4 from Layer 1 targets a specific recognition site on the mRNA of Layer 2.
Inhibition (The Negative Weight): The red line with a bar (—|) represents an inhibitory operation. The endoribonuclease cleaves the mRNA, effectively preventing its translation (Tl) and silencing the final output Y.
Conclusion
By structuring the circuit this way, we have moved from a simple direct regulation to a cascade-based logic. In this multilayered model, the final output Y is a complex function of the initial inputs $X_1$ and $X_2$, mediated by the “hidden” concentration of Csy4. This mimics the hierarchical depth found in artificial neural networks, allowing for more sophisticated biological computations.
Assignment Part 2: Fungal Materials What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts? What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Examples of existing fungal materials, their uses, advantages, and disadvantages
| Aspect | Fungi (Yeast & Filamentous) | Bacteria (e.g., E. coli, Bacillus) |
|---|---|---|
| Protein secretion | Excellent; naturally secrete high titers of enzymes; eukaryotic secretion pathway handles complex proteins | Limited; often require cell lysis or periplasmic expression for recovery |
| Post-translational modifications | Perform glycosylation, disulfide bond formation, proteolytic processing (essential for many eukaryotic proteins) | No glycosylation; disulfide bonds only in periplasm |
| Growth substrate | Grow on simple, inexpensive carbon sources (lignocellulose, agricultural waste) | Require refined carbon sources (glucose, glycerol) for optimal growth |
| Morphology | Filamentous fungi form pellets or mycelial mats; easy to separate from liquid cultures; can colonize solid substrates | Suspension growth; require centrifugation or filtration for recovery |
| Safety status | Many species (S. cerevisiae, K. phaffii, A. oryzae) have GRAS (Generally Recognized as Safe) status for food and pharmaceutical production | Some species are pathogens or opportunistic pathogens; endotoxin concerns for therapeutic applications |
| Genetic tools | Mature tools exist (CRISPR-Cas9, homologous recombination), but fewer standardized parts than bacteria | Extremely mature synthetic biology toolbox; thousands of standardized parts (BioBricks, MoClo) |
| Scalability | Well-established industrial fermentation (citric acid, antibiotics, enzymes) at 100,000+ L scale | Also scalable, but often require stricter sterility and oxygen transfer management |
| Generation time | Slower (90 min to several hours for yeast; days for filamentous fungi) | Fast (20–40 minutes) |
| Genome complexity | Larger genomes, more challenging to engineer multiple simultaneous modifications | Smaller genomes, easier to stack multiple edits |
What might you want to genetically engineer fungi to do and why? Advantages of synthetic biology in fungi vs. bacteria.
Engineered Application: Fungi for Bioremediation of Heavy Metals and PFAS
What to engineer:
| Genetic Modification | Purpose |
|---|---|
| Overexpress metallothioneins | Bind and sequester heavy metals (cadmium, lead, mercury) |
| Express bacterial PFAS-degrading enzymes (e.g., dehalogenases, peroxidases) | Break down per- and polyfluoroalkyl substances (forever chemicals) |
| Inducible promoter system (e.g., copper-inducible) | Activate remediation genes only in contaminated environments |
| Surface display of metal-binding peptides | Increase metal adsorption efficiency |
Why fungi for this application:
Mycelial networks can colonize large soil volumes and penetrate contaminated groundwater zones.
Fungi secrete powerful oxidative enzymes (laccases, peroxidases) naturally suited for breaking down recalcitrant pollutants.
Many fungi grow on cheap substrates (wood chips, agricultural waste), making deployment cost-effective.
Fungi form symbiotic relationships with plant roots (mycorrhizae), allowing phytoremediation enhancement.
Advantages of synthetic biology in fungi compared to bacteria
| Aspect | Fungi (Yeast & Filamentous) | Bacteria (e.g., E. coli, Bacillus) |
|---|---|---|
| Protein secretion | Excellent; naturally secrete high titers of enzymes; eukaryotic secretion pathway handles complex proteins | Limited; often require cell lysis or periplasmic expression for recovery |
| Post-translational modifications | Perform glycosylation, disulfide bond formation, proteolytic processing (essential for many eukaryotic proteins) | No glycosylation; disulfide bonds only in periplasm |
| Growth substrate | Grow on simple, inexpensive carbon sources (lignocellulose, agricultural waste) | Require refined carbon sources (glucose, glycerol) for optimal growth |
| Morphology | Filamentous fungi form pellets or mycelial mats; easy to separate from liquid cultures; can colonize solid substrates | Suspension growth; require centrifugation or filtration for recovery |
| Safety status | Many species (S. cerevisiae, K. phaffii, A. oryzae) have GRAS (Generally Recognized as Safe) status for food and pharmaceutical production | Some species are pathogens or opportunistic pathogens; endotoxin concerns for therapeutic applications |
| Genetic tools | Mature tools exist (CRISPR-Cas9, homologous recombination), but fewer standardized parts than bacteria | Extremely mature synthetic biology toolbox; thousands of standardized parts (BioBricks, MoClo) |
| Scalability | Well-established industrial fermentation (citric acid, antibiotics, enzymes) at 100,000+ L scale | Also scalable, but often require stricter sterility and oxygen transfer management |
| Generation time | Slower (90 min to several hours for yeast; days for filamentous fungi) | Fast (20–40 minutes) |
| Genome complexity | Larger genomes, more challenging to engineer multiple simultaneous modifications | Smaller genomes, easier to stack multiple edits |
Fungi are superior for applications requiring secretion of complex eukaryotic proteins, growth on low-cost feedstocks, and colonization of solid substrates. Bacteria remain better for rapid prototyping, simple protein expression, and applications requiring very fast growth.
Assignment Part 3: First DNA Twist Order Review the Individual Final Project documentation guidelines. Submit this Google Form with your draft Aim 1, final project summary, HTGAA industry council selections, and shared folder for DNA designs. DUE MARCH 20 FOR MIT/HARVARD/WELLESLEY STUDENTS Review Part 3: DNA Design Challenge of the week 2 homework. Design at least 1 insert sequence and place it into the Benchling/Kernel/Other folder you shared in the Google Form above. Document the backbone vector it will be synthesized in on your website.
The form is not yet complete; final project approval is still pending. Once my project is approved, there will be an update to the task. :)