Week 2 HW: DNA-Read Write and Edit
Part 1: Benchling & In-silico Gel Art
Import lambda DNA
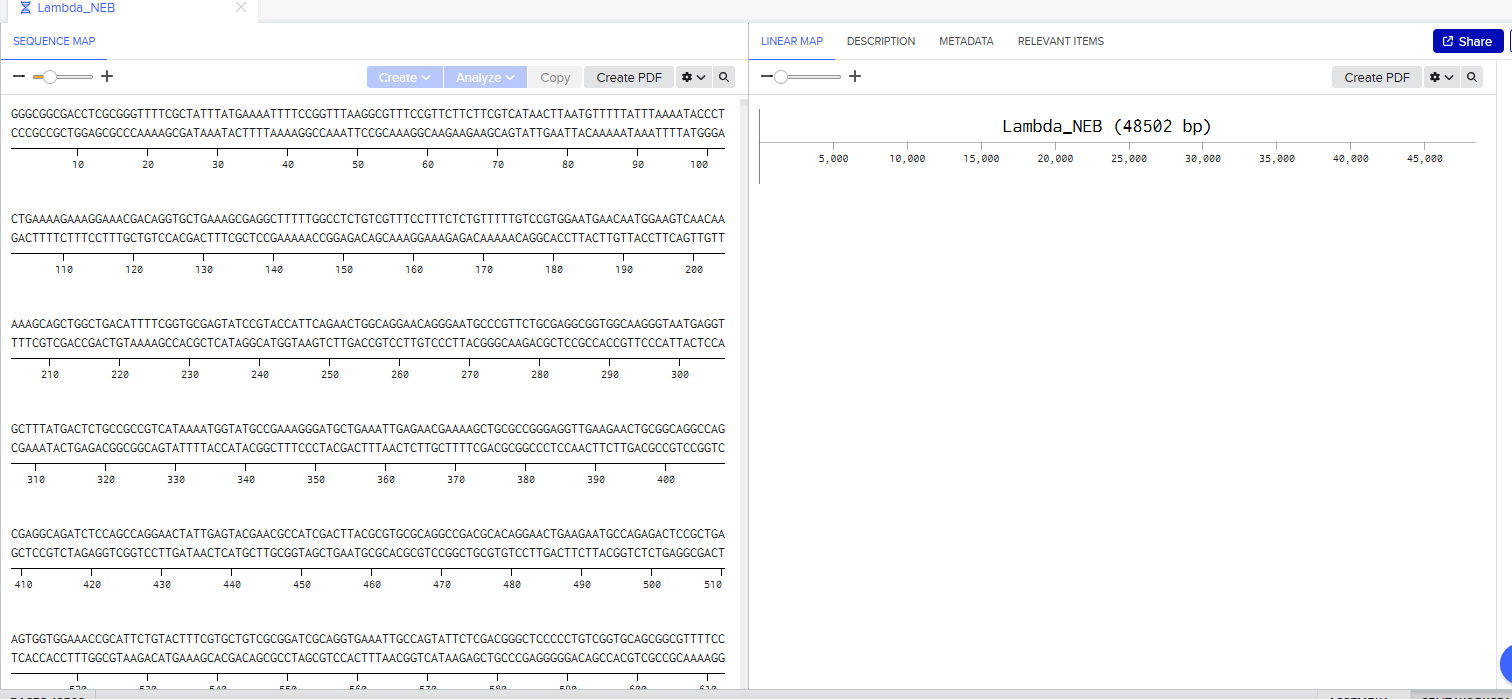

Courtesy:NCBI - O’Leary NA, Cox E, Holmes JB, Anderson WR, Falk R, Hem V, Tsuchiya MTN, Schuler GD, Zhang X, Torcivia J, Ketter A, Breen L, Cothran J, Bajwa H, Tinne J, Meric PA, Hlavina W, Schneider VA. Exploring and retrieving sequence and metadata for species across the tree of life with NCBI Datasets. Sci Data. 2024 Jul 5;11(1):732. doi: 10.1038/s41597-024-03571-y. PMID: 38969627; PMCID: PMC11226681.
Simulation by diffrenet restriction enzymes.
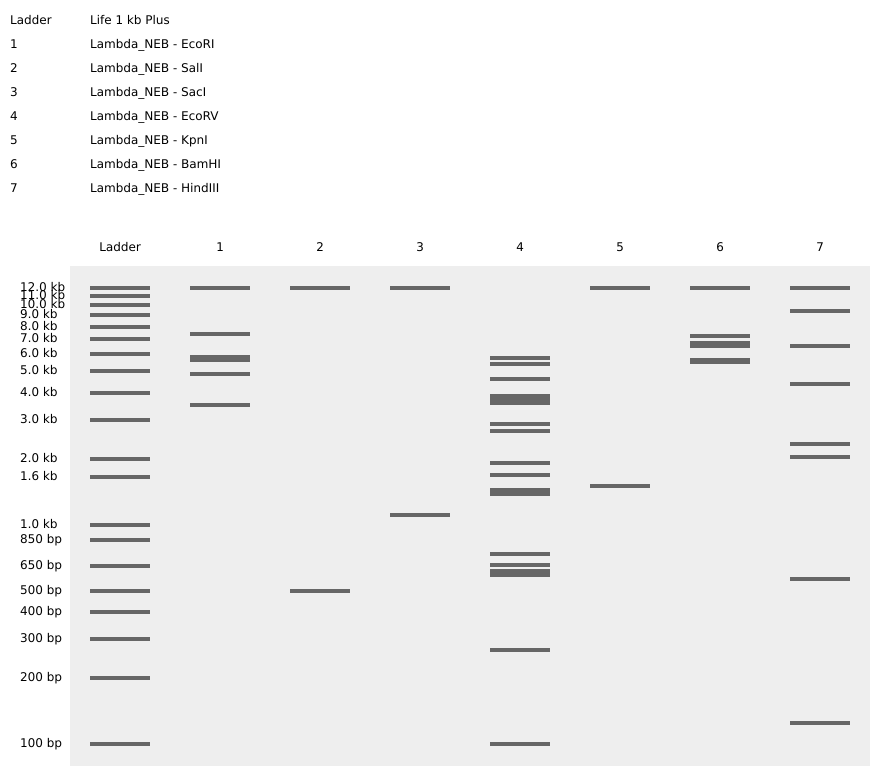


Coutesy:Benchling [Biology Software]. (2026). Retrieved from https://benchling.com.
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
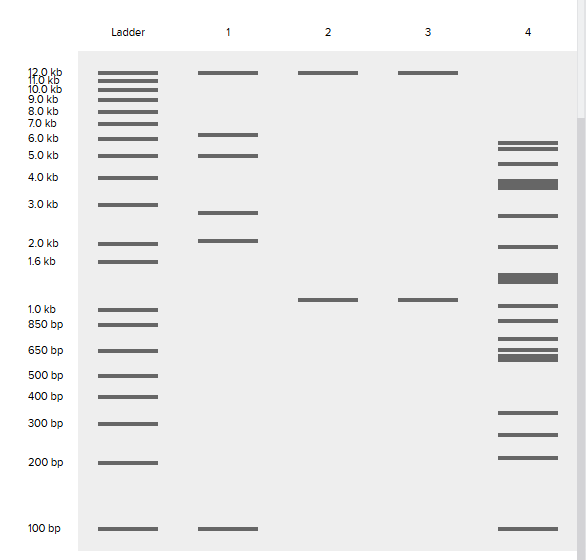




Coutesy:Benchling [Biology Software]. (2026). Retrieved from https://benchling.com.
Part 3: DNA Design Challenge
3.1 Choose your protein.
I have chosen wild type green fluorescent protein. This protein is used as a reporter gene in plasmids to study expression of genes as well as in biosensing, protein localization and also in live cell imaging.
I would like to research on Gfp variants and understand for better use in biosensor field.
Sequence Green fluorescent protein Gene GFP Status UniProtKB reviewed (Swiss-Prot) Organism Aequorea victoria (Water jellyfish) (Mesonema victoria)
FASTA sequence
AAA58246.1 green-fluorescent protein [Aequorea victoria] MSKGEELFTGVVPILVELDGDVNGQKFSVSGEGEGDATYGKLTLKFICTTGKLPVPWPTLVTTFSYGVQC FSRYPDHMKQHDFFKSAMPEGYVQERTIFYKDDGNYKTRAEVKFEGDTLVNRIELKGIDFKEDGNILGHK MEYNYNSHNVYIMADKPKNGIKVNFKIRHNIKDGSVQLADHYQQNTPIGDGPVLLPDNHYLSTQSALSKD PNEKRDHMILLEFVTAAGITHGMDELYK
Courtesy: The UniProt Consortium, “UniProt: the Universal Protein Knowledgebase in 2025,” Nucleic Acids Research, 2025.
3.2 Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
L29345.1 Aequorea victoria green-fluorescent protein (GFP) mRNA, complete cds TACACACGAATAAAAGATAACAAAGATGAGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTT GTTGAATTAGATGGCGATGTTAATGGGCAAAAATTCTCTGTCAGTGGAGAGGGTGAAGGTGATGCAACAT ACGGAAAACTTACCCTTAAATTTATTTGCACTACTGGGAAGCTACCTGTTCCATGGCCAACACTTGTCAC TACTTTCTCTTATGGTGTTCAATGCTTTTCAAGATACCCAGATCATATGAAACAGCATGACTTTTTCAAG AGTGCCATGCCCGAAGGTTATGTACAGGAAAGAACTATATTTTACAAAGATGACGGGAACTACAAGACAC GTGCTGAAGTCAAGTTTGAAGGTGATACCCTTGTTAATAGAATCGAGTTAAAAGGTATTGATTTTAAAGA AGATGGAAACATTCTTGGACACAAAATGGAATACAACTATAACTCACATAATGTATACATCATGGCAGAC AAACCAAAGAATGGAATCAAAGTTAACTTCAAAATTAGACACAACATTAAAGATGGAAGCGTTCAATTAG CAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCCTGTCCTTTTACCAGACAACCATTACCTGTC CACACAATCTGCCCTTTCCAAAGATCCCAACGAAAAGAGAGATCACATGATCCTTCTTGAGTTTGTAACA GCTGCTGGGATTACACATGGCATGGATGAACTATACAAATAAATGTCCAGACTTCCAATTGACACTAAAG TGTCCGAACAATTACTAAATTCTCAGGGTTCCTGGTTAAATTCAGGCTGAGACTTTATTTATATATTTAT AGATTCATTAAAATTTTATGAATAATTTATTGATGTTATTAATAGGGGCTATTTTCTTATTAAATAGGCT ACTGGAGTGTAT
Courtesy:NCBI - O’Leary NA, Cox E, Holmes JB, Anderson WR, Falk R, Hem V, Tsuchiya MTN, Schuler GD, Zhang X, Torcivia J, Ketter A, Breen L, Cothran J, Bajwa H, Tinne J, Meric PA, Hlavina W, Schneider VA. Exploring and retrieving sequence and metadata for species across the tree of life with NCBI Datasets. Sci Data. 2024 Jul 5;11(1):732. doi: 10.1038/s41597-024-03571-y. PMID: 38969627; PMCID: PMC11226681.
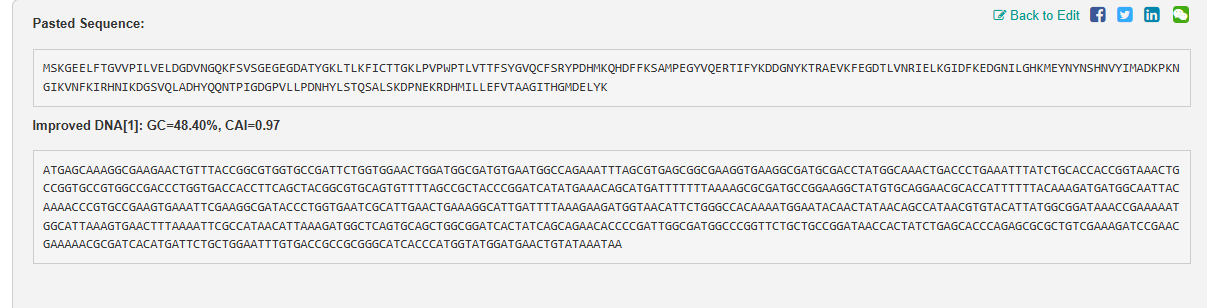
3.3 Codon optimization
Codon optimization is done as each organism has a set of codon preferences for the same amino acids. Due to codon redundancy the same amino acid acn be coded for by multiple sets of codon. This method is used to maximise protein expression based on tailoring the DNA sequence based on tRNa abundance in the host organism. It increases translation efficiency, improves protein yield, and eliminates negative regulatory elements (repressors), which is crucial for producing recombinant proteins, vaccines, and gene therapies.
This was done using Vector Builder {Link: https://www.vectorbuilder.com https://en.vectorbuilder.com/resources/cite.html}.

3.3You have a sequenc e! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
There are many technologies that can be used to produce protein.
1.Recombinant DNA technology
The GFP gene is inserted into plasmid that can replicate inside host cell. They have strong promoters to ensure high levels GFP expression.Based on the expression of the selective marker plasmids are selected and transfected into host cells.
- Heterologous Expression Systems (Host Cells) Bacterial protein expression, primarily using E. coli, is a fast, cost-effective, and scalable method for producing recombinant proteins. It involves :
Cloning: Inserting the GFP gene into an expression vector.(E.coli plasmid)
Transformation: Introducing the plasmid into competent bacterial cells.
Expression: Culturing the cells and inducing GFP production.
Harvesting & Purification: Lysis of cells and purification of the GFP.
- Cell-Dependent Method (In Vivo)
This occurs within the host cell
Transcription (DNA to mRNA): Initiation: The enzyme RNA polymerase binds to a GFP DNA sequence called the promoter, signaling the start of the gene. Elongation: RNA polymerase unwinds the DNA helix and reads one strand (the template strand) in the 3′ to 5′ direction, synthesizing a complementary RNA molecule in the 5′ to 3′ direction. Termination: Upon reaching a “terminator” sequence, the RNA polymerase releases the newly formed pre-mRNA (GFP) strand.
Translation (mRNA to Protein): The mature GFP mRNA moves to the cytoplasm, where it binds to a ribosome. The ribosome reads the codons. Transfer RNA (tRNA) molecules, carrying specific amino acids, match their anticodons to the mRNA codons. The ribosome catalyzes a peptide bond between amino acids, building a polypeptide chain until a stop codon is reached.
Highly regulated, capable of complex post-translational modifications (folding, glycosylation) in eukaryotes, but slow and limited by cell viability.
- Cell-Free Method (In Vitro)
Cell-free protein synthesis (CFPS) harnesses the machinery (ribosomes, tRNAs, enzymes) extracted from cells to produce proteins in a test tube, allowing direct control over the environment.
Preparation: Cells (e.g., E. coli,) are grown, lysed, and centrifuged to remove DNA, cell walls, and debris, leaving only the translational machinery.
Method: The extracted, active machinery is mixed with the GFP DNA template (plasmid or PCR-amplified), amino acids, energy sources (like ATP/GTP), and cofactors.
Process: The system can be coupled (transcription and translation occur together) or uncoupled (using mRNA directly). It bypasses the need for cell viability making it ideal for toxic, membrane, or complex proteins.
Fast (hours instead of days), open system allowing direct manipulation, and capable of producing toxic or high-yield proteins, but can be expensive for large-scale production.
3.5[Optional] How does it work in nature/biological systems?
The protein works as a fluorescence producing mechanism in the jelly fish producing green light.
1. Describe how a single gene codes for multiple proteins at the transcriptional level.
A single gene codes for multiple proteins primarily through alternative splicing of pre-mRNA, where different combinations of exons are joined together after introns are removed.
Pre-mRNA is spliced in multiple ways to include or exclude specific exons. This produces different mRNA transcripts, which are then translated into different protein isoforms.
Alternative Promoters: A gene may have multiple promoters, allowing transcription to start at different points, resulting in mRNA molecules with different 5’ends.
Alternative Polyadenylation: This process alters the end of the mRNA, which can affect mRNA stability and localization, leading to different protein products.
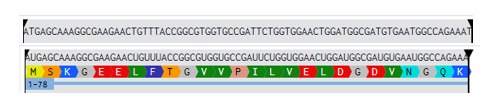
2.Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!!

Part 4: Prepare a Twist DNA Synthesis Order
4.2 Build Your DNA Insert Sequence
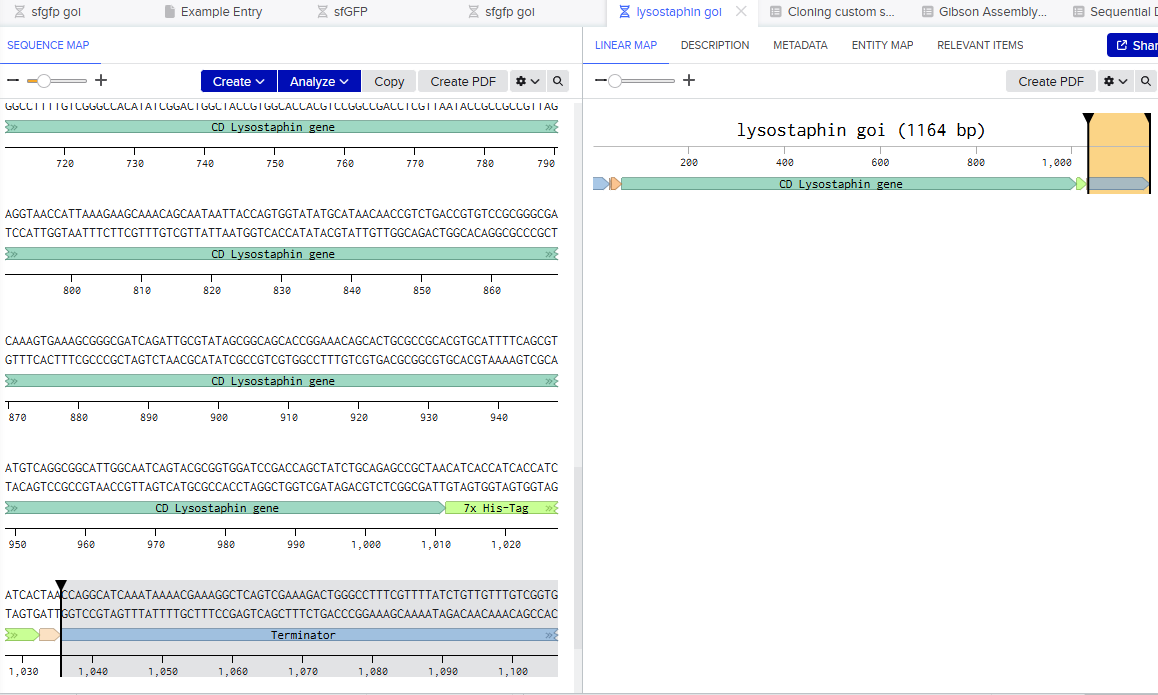

Courtesy:Benchling [Biology Software]. (2026). Retrieved from https://benchling.com.
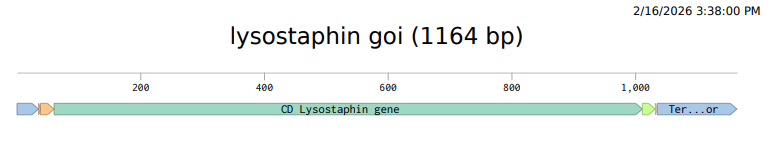


Coutesy:Benchling [Biology Software]. (2026). Retrieved from https://benchling.com.
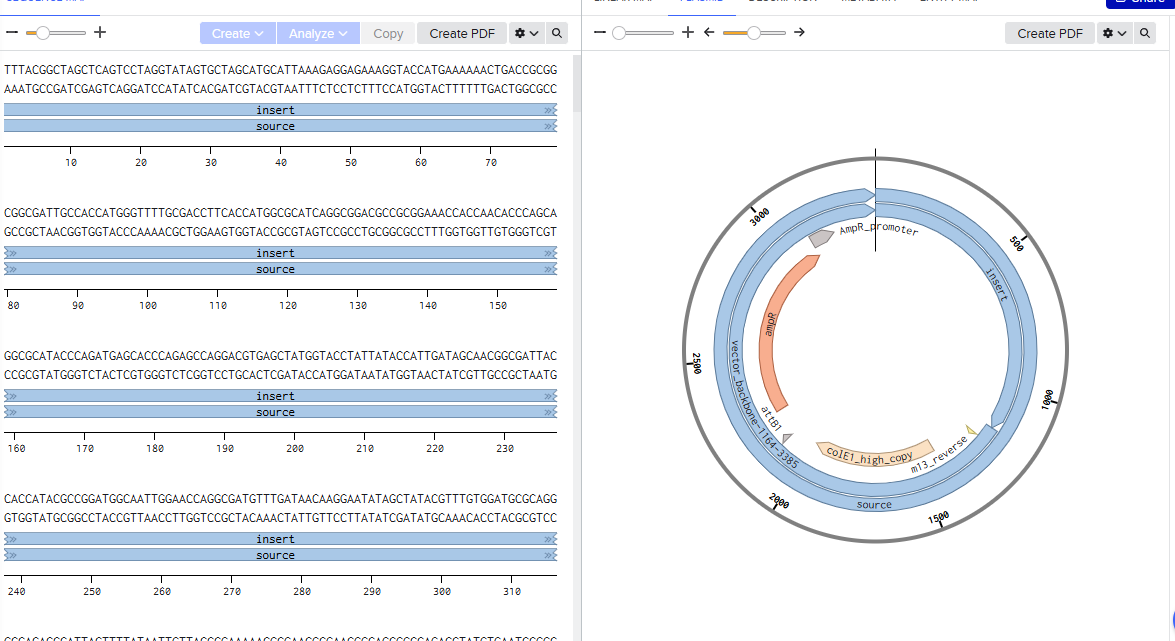

Coutesy: Twist Bioscience
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
I would like to explore DNA of Antibiotic resistant bacteria against penicillin. I want to study their sequence and understand how they become resistant to the antibiotics, study their interaction with antibioticsand compare their genes with their susceptible counterparts to understand which sequence produces which type of resistance to the organism towards antibiotics.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Long-Read Sequencing (Oxford Nanopore Technologies - ONT): Usage: Emerging, real-time sequencing (e.g., MinION) for rapid diagnostics.
Why: Produces very long reads (10 kb to >1 Mb) that can bridge repetitive regions, allowing for the easy reconstruction of plasmids and the identification of the genetic context of resistance genes (e.g., whether they are on a plasmid or chromosome).
1. Is your method first-, second- or third-generation or other? How so?
Oxford Nanopore Technologies (ONT) is considered a third-generation (or sometimes referred to as long-read) sequencing technology. The first-generation (Sanger) or second-generation (Illumina/NGS) methods rely on DNA synthesis and detection of light signals, while ONT measures changes in electrical current as single molecules of DNA/RNA pass through a protein nanopore.
It is categorized as third-generation sequencing due to its ability to sequence long, single molecules of nucleic acids in real-time. It generates very long to ultra-long reads, superior assembly of complex genomes and structural variant detection.
2. What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
The input for the ONT method would be the genomic DNA (gDNA) or plasmid DNA, extracted from pure bacterial cultures or environmental samples.
Steps for Sequencing AMR Genes
Sample Preparation and DNA Extraction:
AMR Bacteria are cultured in LB broth and collected. DNA is extracted using kits for high molecular weight DNA using Qiagen or MagAttract kits. It is quantified and purified using Qubit and Nanodrop respectively.
Library Preparation: Rapid Kits (e.g., Rapid Barcoding Kit SQK-RBK114.24): Transposases fragment and tag DNA with adapters in one step, ideal for quick turnaround. Ligation Kits (e.g., Ligation Sequencing Kit SQK-LSK114): Provides higher output and longer reads for more comprehensive genome coverage.
3.What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
The prepared library is loaded onto an ONT flow cell (e.g., R10.4 or R9.4.1) on a device like the MinION or GridION. As DNA passes through the nanopore, ionic current changes are measured and recorded.
1. Raw Signal Capturing and Preprocessing Signal Acquisition: As DNA/RNA strands pass through a nanopore, they disrupt an electrical current. MinKNOW (ONT’s controlling software) records these changes. File Format Conversion: Raw signals are traditionally stored in .fast5 files but are increasingly saved in the more efficient POD5 file format, which is designed for faster data handling and processing. Data Preparation: Before basecalling, the raw signals are often organized and prepared, potentially involving filtering to remove uninformative data.
2. The Basecalling Process (Neural Network Decoding) Deep Learning Models: Modern ONT basecallers (like Dorado, Guppy, or Bonito) use neural networks (such as Recurrent Neural Networks - RNNs, or Transformer models) to analyze the raw signal data. “Squiggle” to Base Translation: The neural network maps the electrical signal changes to the corresponding nucleotide sequences, usually in real time while the sequencing is still running. Move Tables: The process identifies when a new base enters the pore, producing a “move table” that indicates which part of the signal corresponds to which base. Quality Scoring: Alongside the base sequence, the basecaller assigns a probability score to each base, often represented as a Phred score to indicate confidence in the call.
4.What is the output of your chosen sequencing technology?
Post-Processing and Output FASTQ File Generation: The primary output is a FASTQ file, containing the sequences and their associated quality scores.
BAM/CRAM Output: Alternatively, basecallers can output files in SAM, BAM, or CRAM formats, which can include both the sequence data and the signal-level information.
Demultiplexing (Optional): If multiple samples were mixed in a single run (barcoding), the software identifies which reads belong to which barcode and separates them into individual files.
Polishing (Optional): Additional steps like Medaka or Nanopolish may be used to refine the sequence data further, especially for improving consensus accuracy.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I want to synthesize the Staphylococcus staphylolyticus lysostaphin gene
1 gaaaattcca aaaaaaaacc tactttctta atattgattc atattatttt aacacaatca
61 gttagaattt caaaaatctt aaagtcaatt tttgagtgtg tttgtatatt tcatcaaagc
121 caatcaatat tattttactt tcttcatcgt taaaaaatgt aatatttata aaaatatgct
181 attctcataa atgtaataat aaattaggag gtattaaggt tgaagaaaac aaaaaacaat
241 tattatacga cacctttagc tattggactg agtacatttg ccttagcatc tattgtttat
301 ggagggattc aaaatgaaac acatgcttct gaaaaaagta atatggatgt ttcaaaaaaa
361 gtagctgaag tagagacttc aaaaccccca gtagaaaata cagctgaagt agagacttca
421 aaagctccag tagaaaatac agctgaagta gagacttcaa aagctccagt agaaaataca
481 gctgaagtag agacttcaaa agctccagta gaaaatacag ctgaagtaga gacttcaaaa
541 gctccggtag aaaatacagc tgaagtagag acttcaaaag ctccggtaga aaatacagct
601 gaagtagaga cttcaaaagc cccagtagaa aatacagctg aagtagagac ttcaaaagct
661 ccagtagaaa atacagctga agtagagact tcaaaagctc cggtagaaaa tacagctgaa
721 gtagagactt caaaagcccc agtagaaaat acagctgaag tagagacttc aaaagctcca
781 gtagaaaata cagctgaagt agagacttca aaagctccgg tagaaaatac agctgaagta
841 gagacttcaa aagccccagt agaaaataca gctgaagtag agacttcaaa agccctggtt
901 caaaatagaa cagctttaag agctgcaaca catgaacatt cagcacaatg gttgaataat
961 tacaaaaaag gatatggtta cggtccttat ccattaggta taaatggcgg tatccactac
1021 ggagttgatt tttttatgaa tattggaaca ccagtaaaag ctatttcaag cggaaaaata
1081 gttgaagctg gttggagtaa ttacggagga ggtaatcaaa taggtcttat tgaaaatgat
1141 ggagtgcata gacaatggta tatgcatcta agtaaatata atgttaaagt aggagattat
1201 gtcaaagctg gtcaaataat cggttggtct ggaagcactg gttattctac agcaccacat
1261 ttacacttcc aaagaatggt taattcattt tcaaattcaa ctgcccaaga tccaatgcct
1321 ttcttaaaga gcgcaggata tggaaaagca ggtggtacag taactccaac gcccaataca
1381 ggttggaaaa caaacaaata tggcacacta tataaatcag agtcagctag cttcacacct
1441 aatacagata taataacaag aacgactggt ccatttagaa gcatgccgca gtcaggagtc
1501 ttaaaagcag gtcaaacaat tcattatgat gaagtgatga aacaagacgg tcatgtttgg
1561 gtaggttata caggtaacag tggccaacgt atttacttgc ctgtaagaac atggaataaa
1621 tctactaata ctttaggtgt tctttgggga actataaagt gagcgcgctt tttataaact
1681 tatatgataa ttagagcaaa taaaaatttt ttctcattcc taaagttgaa gcttttcgta
1741 atcatgtcat agcgtttcct gtgtgaaatt gcttagcctc acaattccac acaacatacg
1801 agccggaaca taaagtgcta agcct
Courtesy:NCBI - O’Leary NA, Cox E, Holmes JB, Anderson WR, Falk R, Hem V, Tsuchiya MTN, Schuler GD, Zhang X, Torcivia J, Ketter A, Breen L, Cothran J, Bajwa H, Tinne J, Meric PA, Hlavina W, Schneider VA. Exploring and retrieving sequence and metadata for species across the tree of life with NCBI Datasets. Sci Data. 2024 Jul 5;11(1):732. doi: 10.1038/s41597-024-03571-y. PMID: 38969627; PMCID: PMC11226681.
Lysostaphin is highly active against antibiotic-resistant S. aureus (MRSA) and intermediately vancomycin-susceptible S. aureus (VISA), as its mechanism of action differs from traditional antibiotic resistance mechanisms. Lysostaphin is a potent zinc-dependent metalloendopeptidase (specifically a glycylglycine endopeptidase) produced by Staphylococcus simulans. It acts as an antibacterial agent (bacteriocin) with high efficiency against Staphylococcus aureus (including MRSA).
ii) What technology or technologies would you use to perform this DNA synthesis and why?
1.What are the essential steps of your chosen sequencing methods?
Codon Optimization: To increase expression efficiency, the native lss gene is often codon-optimized to match the preferences of the host microorganism, such as E. coli.
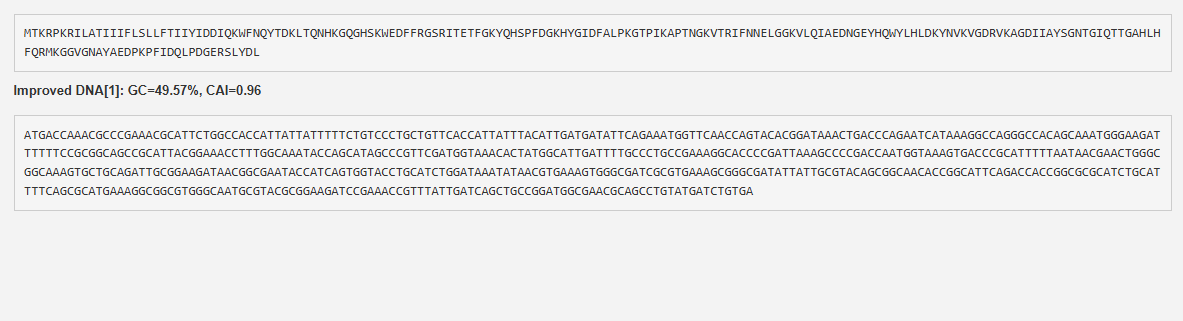
 This was done using Vector Builder {Link: https://www.vectorbuilder.com https://en.vectorbuilder.com/resources/cite.html}.
This was done using Vector Builder {Link: https://www.vectorbuilder.com https://en.vectorbuilder.com/resources/cite.html}.
Vector Construction and Expression Systems: The gene is cloned into various expression vectors (e.g., pET-22b(+), pWB980, pET32a) and transformed into hosts like E. coli BL21(DE3) or Bacillus subtilis WB600, a strain engineered to be deficient in six extracellular proteases, reducing protein degradation.
Constitutive and Inducible Promoters: While many systems use inducible promoters (e.g., IPTG) to control production, recent advances include using constitutive, non-inducible promoters (e.g., pemIK-Sa1 from staphylococcal toxin-antitoxin systems) to reduce costs for large-scale production.
Restriction Enzyme Cloning: The synthetic lysostaphin gene is digested with restriction enzymes (e.g., EcoRI, XhoI, NdeI) and ligated into expression vectors like pPIC9 or pET22b(+) using T4 DNA ligase.
Gibson Assembly: The NEBuilder Assembly Tool is used to design primers for amplifying target regions, which are then assembled into plasmids (e.g., pMAD).
Homologous Recombination: Homologous recombinase is used to ligate optimized lysostaphin fragments into vectors.
2. What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
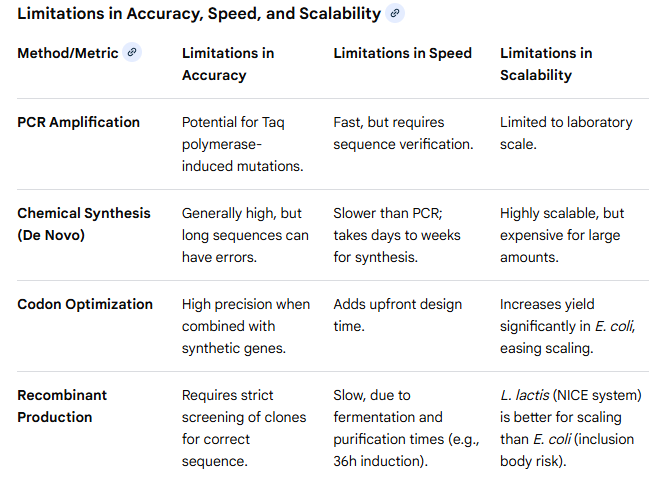

5.3 DNA Edit
i) What DNA would you want to edit and why?
I would want to edit, silence, or delete antimicrobial resistance (AMR) genes in bacteria, specifically targeting resistance plasmids or chromosomal genes to restore antibiotic susceptibility.
1.What technology or technologies would you use to perform these DNA edits and why?
I would want to use CRISPR technology.CRISPR-Cas9 is a genome-editing technology that uses a guide RNA (gRNA) to direct the Cas9 enzyme to a specific DNA sequence, where it acts as molecular scissors to create a double-strand break. The cell then repairs this cut using either NHEJ (resulting in gene knockouts) or HDR (enabling precise gene insertion/correction). It is faster, cheaper, and more accurate alternative to previous methods.
The other editing technologies are TALENs & ZFNs: Older, customizable nuclease technologies that bind to specific DNA sequences to induce breaks, though they are generally less flexible than CRISPR.
TALENs generally exhibit significantly lower off-target effects compared to both ZFNs and CRISPR, making them safer for certain applications.They are widely used for precise, large-scale, and stable genome engineering in plants and animals.
2. How does your technology of choice edit DNA? What are the essential steps?
CRISPR-Cas9 is a programmable gene-editing technology that uses a guide RNA (sgRNA) to direct the Cas9 enzyme to a specific DNA sequence, creating a targeted double-strand break (DSB). The system relies on a PAM sequence for binding, after which the cell repairs the cut using error-prone NHEJ (for gene disruption) or precise HDR (for gene correction).
Guide RNA (sgRNA): A synthetic RNA sequence designed to be complementary to the target DNA, directing the Cas9 enzyme to the precise location in the genome.
Cas9 Nuclease: An enzyme acting as molecular scissors that creates a double-strand break (DSB) in the DNA, specifically three bases upstream of a required
Protospacer Adjacent Motif (PAM) sequence.
Target Recognition: The CRISPR-Cas9 complex scans the genome for a PAM sequence (commonly 5’-NGG-3’). Once found, it checks if the sgRNA matches the adjacent DNA sequence.
Once the DNA is cut, the cell attempts to repair it, which allows for gene editing:
Non-Homologous End Joining (NHEJ): A fast,, error-prone repair mechanism that often introduces small deletions or insertions (indels), disrupting or “knocking out” the target gene.
Homology-Directed Repair (HDR): A precise repair mechanism used if a repair template is provided, allowing for the insertion of new, desired genetic information or correction of mutations.
3. What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
The preparation step involves the identification of the AMR gene to be targeted, then analyse the sequence for protospacer adjacent motif(PAM).
Design gRNAs complementary to AMR genes.Benchling tool can be used to reduce off-target effects and maximise specificity.
Construct plasmids for cloning gRNA and Cas9 gene inserts using restriction enzyme and ligase.
Transformation of E.coli cells with the plasmids, purication and verification of sequences using Sanger sequencing.
Preparation of delivery systems: chemical transformation, electroporation,bacteriophage particle for phage mediated delivery.
**What are the limitations of your editing methods (if any) in terms of efficiency or precision?*8
Inputs for AMR editing
Cas Nuclease: Cas9 (commonly S. pyogenes Cas9) is the primary protein, or Cas12a/Cpf1.
Guide RNA (gRNA/sgRNA): Specifically designed 20-nt guide sequence with a scaffold.
Plasmids: Expression vectors containing both the Cas9 gene and the gRNA sequence (e.g., pX330).
Primers: For PCR verification of the edit.
Enzymes: Restriction enzymes (e.g., BpiI, BsmBI) and T4 DNA ligase for cloning.
Delivery Vehicle: Phages (e.g., temperate/lytic phage), nanoparticles, or conjugated plasmids.
Limitations of CRISPR
Off-Target Effects: The CRISPR-Cas9 complex may bind to and modify genomic sites that are not the intended target, leading to potential, unintended, and sometimes harmful mutations.
Delivery Challenges: Delivering the large CRISPR-Cas9 components into specific cells or tissues is difficult, which limits its application in many clinical contexts.
Low Efficiency: The process is not 100% efficient, particularly with homology-directed repair (HDR), leading to cells that may not have the desired edit.
Mosaicism: In animal models, not all cells may be edited equally, resulting in mosaicism where only some cells carry the desired modification, making it difficult to identify, study, or rely on the desired edit.
PAM Sequence Requirements: The Cas9 protein must bind to a specific protospacer adjacent motif (PAM) sequence located next to the target DNA, which may not be present at the desired location.
Persistent Binding: In some instances, the Cas9 protein binds to the cut site persistently, preventing the DNA repair machinery from functioning, leading to editing failure.