week-06-hw-genetic-circuits-part-i

Assignment: DNA Assembly
Answer these questions about the protocol
1.What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The components of Phusion High-Fidelity PCR Master Mix contains Phusion DNA Polymerase (high-accuracy, 50x higher than Taq), dNTPs (nucleotide building blocks), and optimized reaction buffer with MgCl2(essential for enzyme activity).
2.What are some factors that determine primer annealing temperature during PCR?
- The annealing temperature depends on the melting temperatures of primers.It is generally 3–5°C lower than the lowest temperature of the primer-template pair.
-The GC content of primers and length of primers increase the melting temperature, require higher annealing temperature.
-Higher magnesium ion concentration tends to satbilize teh duplex tehreby increasing annealing temperature.
-Higher concentration of primers require higher annealing temperature.
-DMSO or glycerol decrease melting temperature of Dna so lower annealing temperatuer is required.
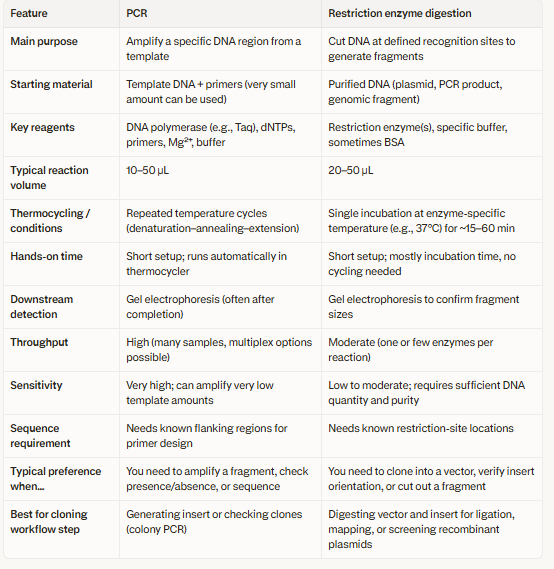
3.There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.

4.How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
The designed primers should be of 15-40 bp in length, terminal overlaps matching adjacent fragments, mutations can be minimised by using high fidelity polymerases and purify PCR products.This will remove templates and primers. Gel electrophoresis can be used to verify the PCR or restriction digest products by linearizing the vectors.
5.How does the plasmid DNA enter the E. coli cells during transformation?
The plasmid Dna can enter via the electroporation method where a brief electric pulse is applied to a cell. This creates pores in the membrane and allows the plasmid DNA to enter the cell.
The other method is to treat the cell to Heat shock method. The cells are treated with cold CaCl2,making it permeable and neutralizes charge on DNA. A sudden increase in tempertaure to 42°C helps create pores in the membrane and allows DNA to enter the cell.
6.Describe another assembly method in detail (such as Golden Gate Assembly)
1.Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Golden Gate Assembly is a precise, “one-pot” cloning method that utilizes Type IIS restriction enzymes (e.g., BsaI, BsmBI) and T4 DNA ligase to directionally assemble multiple DNA fragments simultaneously. It is “scarless,” meaning no extra, unwanted sequences remain between fragments, making it superior to traditional restriction cloning.
Type IIS enzymes recognize asymmetric DNA sequences but cleave outside of them, leaving unique 4-base pair overhangs. These overhangs are designed to be complementary, allowing multiple fragments to ligate in a specific order.
The restriction digestion and ligation occur in a single tube, typically using thermal cycling to alternate between optimal digestion and ligation temperatures.
The Type IIS recognition sites are positioned at the very ends of the DNA fragments, designed so they are removed from the final assembled product. Because the product lacks these sites, it cannot be re-digested, making the reaction irreversible and highly efficient. If internal Type IIS sites exist within the DNA fragments, they must be removed via silent mutations (a process called domestication) to prevent premature digestion.
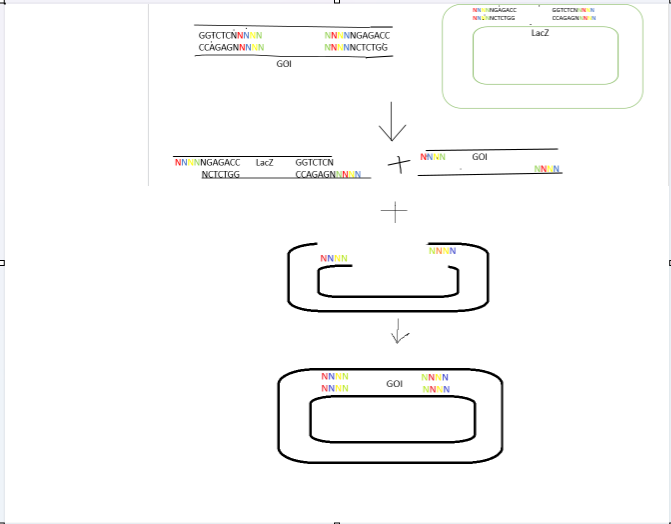

2.Model this assembly method with Benchling or a similar tool!
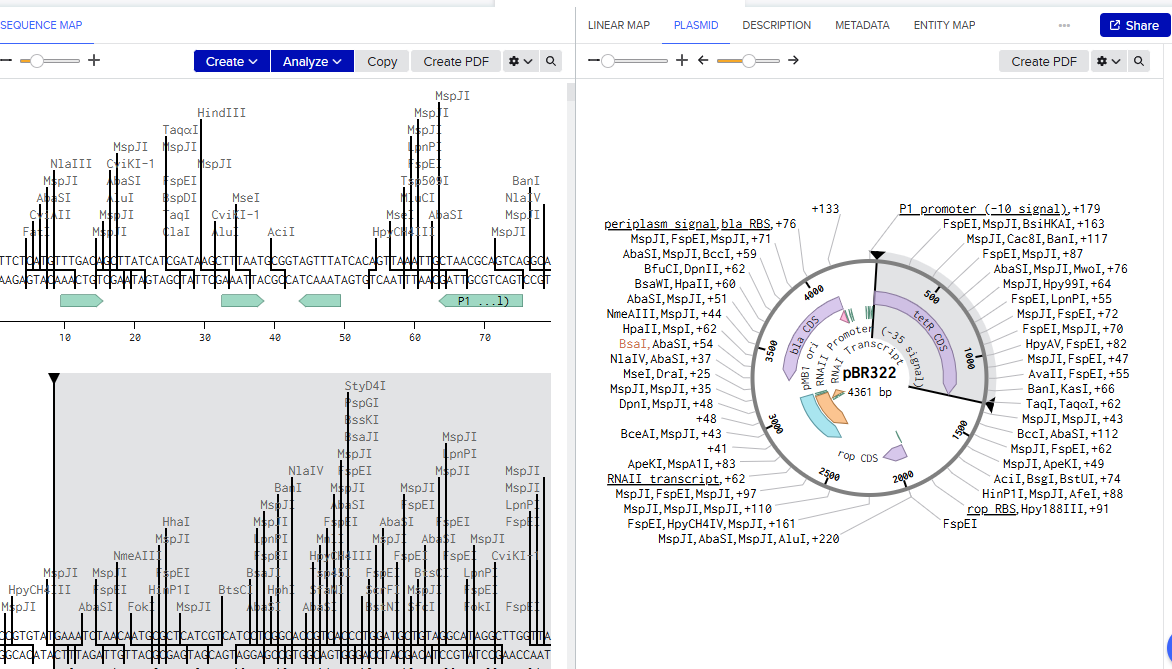

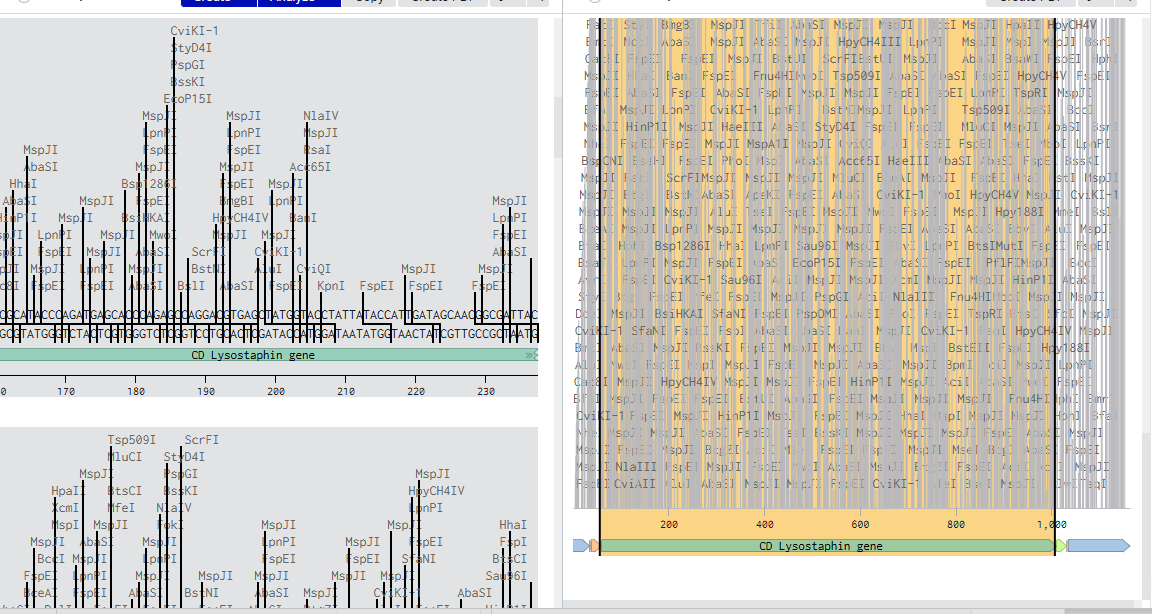

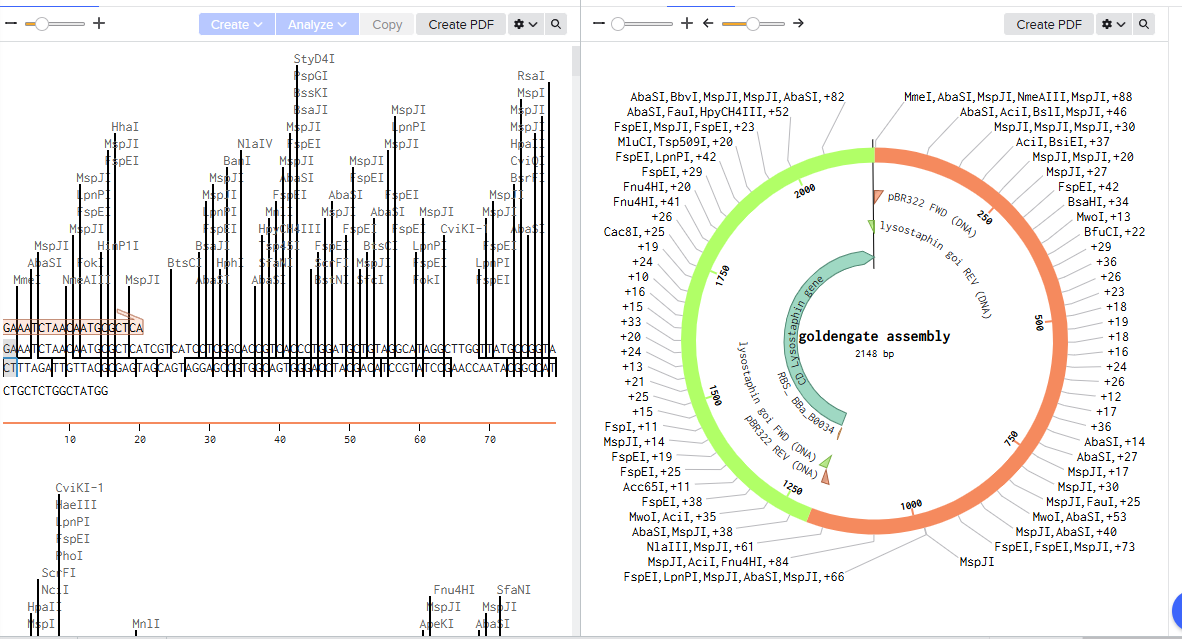

Assignment: Asimov Kernel
1.Create a Repository for your work
2.Create a blank Notebook entry to document the homework and save it to that Repository
4.Create a blank Construct and save it to your Repository
Explore the devices in the Bacterial Demos Repo to understand how the parts work together by running the Simulator on various examples, following the instructions for the simulator found in the “Info” panel (click the “i” icon on the right to open the Info panel)
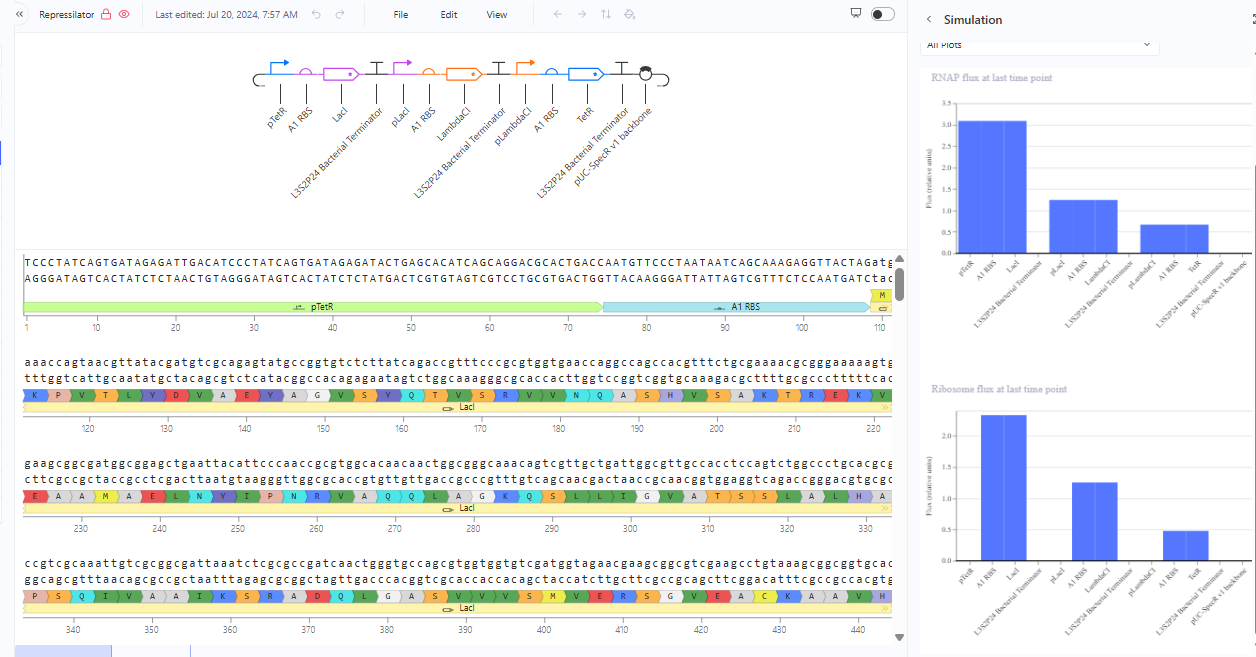


Recreate the Repressilator in that empty Construct by using parts from the Characterized Bacterial Parts repository
Search the parts using the Search function in the right menu
Drag and drop the parts into the Construct
Confirm it works as expected by running the Simulator (“play” button) and compare your results with the Repressilator Construct found in the Bacterial Demos repository
Document all of this work in your Notebook entry - you can copy the glyph image and the simulator graphs, and paste them into your Notebook
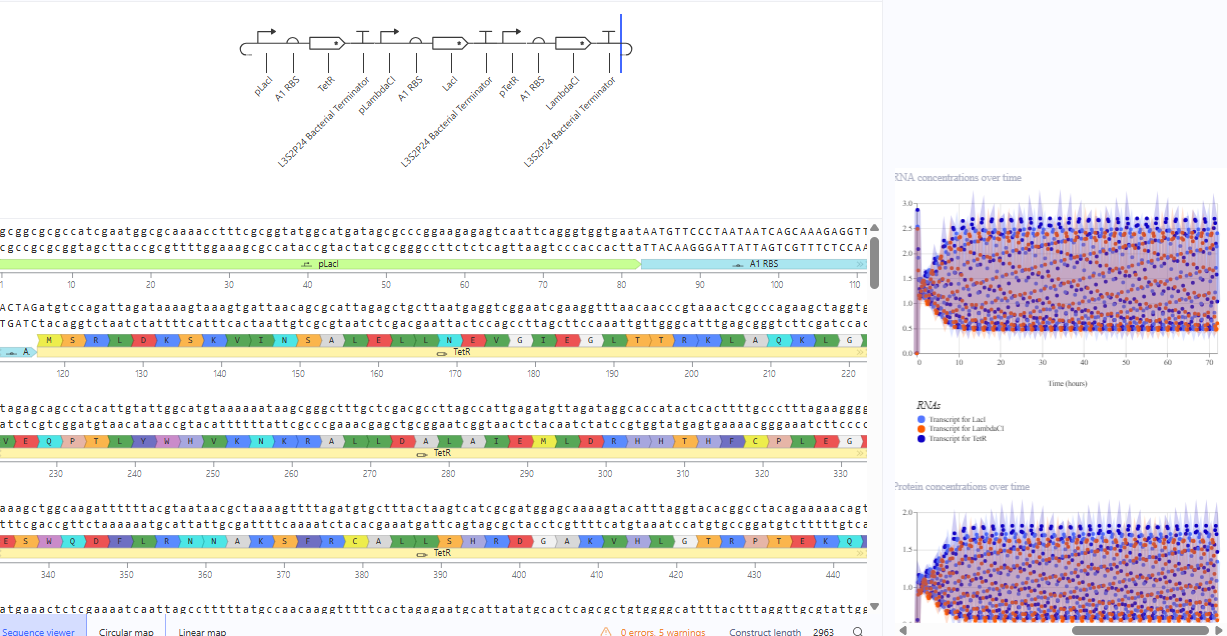

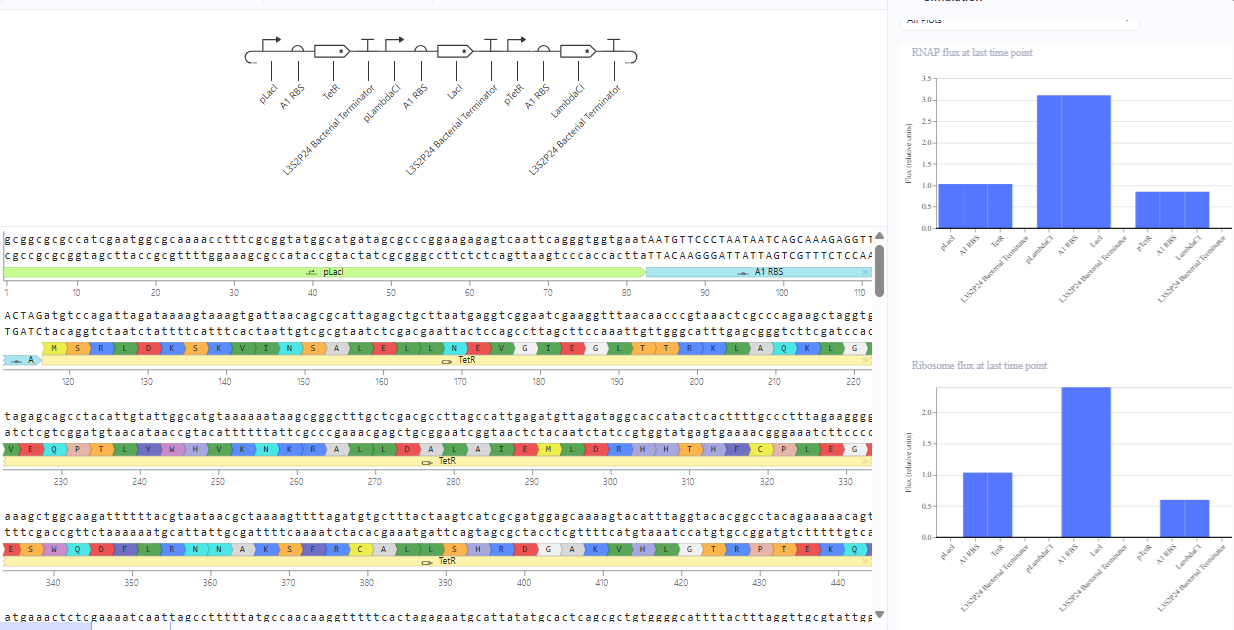

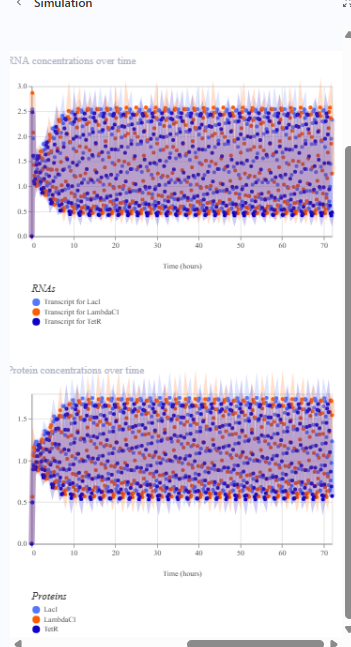
The repressilator cretaed by me does work as expected. In comparison to the construct found in Bacterial demos, the graph obtained is similar.
The RNA flux (rate of RNA production) for the plac promoter (often ) in the original repressilator is generally higher than that of pTet and pLambda due to higher basal or un-repressed promoter strength.The pLacI promoter, which regulates lacI, is a stronger promoter compared to other two when not fully repressed.The lacI system is inherently leakier than tetR or cI, allowing higher basal expression and faster overall accumulation of its mRNA when the repressive factor (TetR) is not present at high concentrations.
5.Build three of your own Constructs using the parts in the Characterized Bacterials Parts Repo
Explain in the Notebook Entry how you think each of the Constructs should function
Run the simulator and share your results in the Notebook Entry
Construct 1 : Expressing yeGFP gene
The construct should express the protein yeGFP.
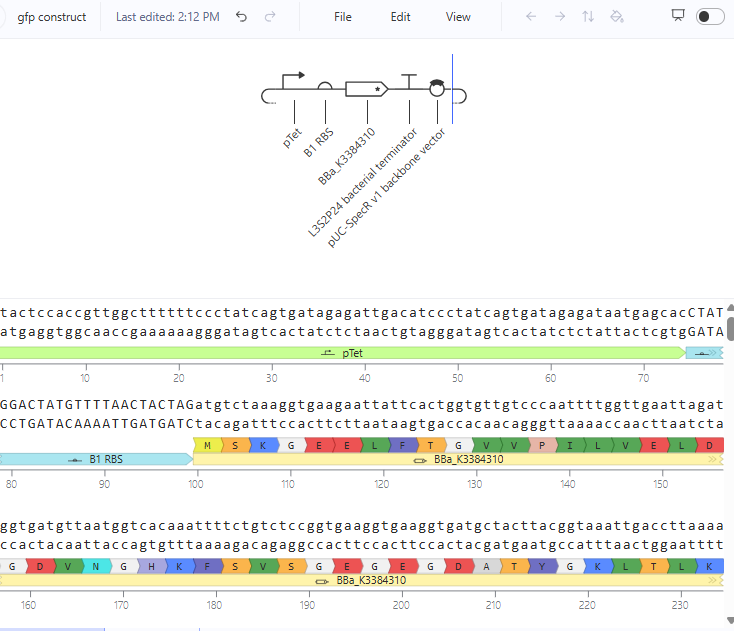

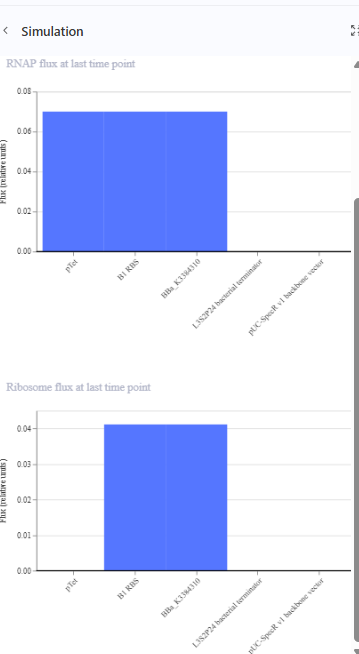

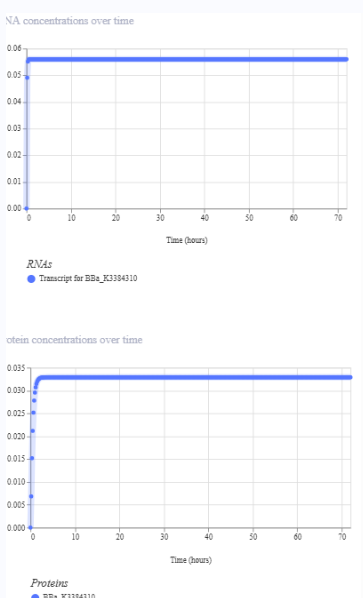

Construct 2: expression of eYFP and PcbA gene
The two genes should be expressed independently. In a two-gene construct, particularly if they are in close proximity, the activation of the strong promoter pLacI might cause the RNA polymerase to interfere with the transcription of the weaker promoter pTetR, resulting in no mRNA for eYFP.
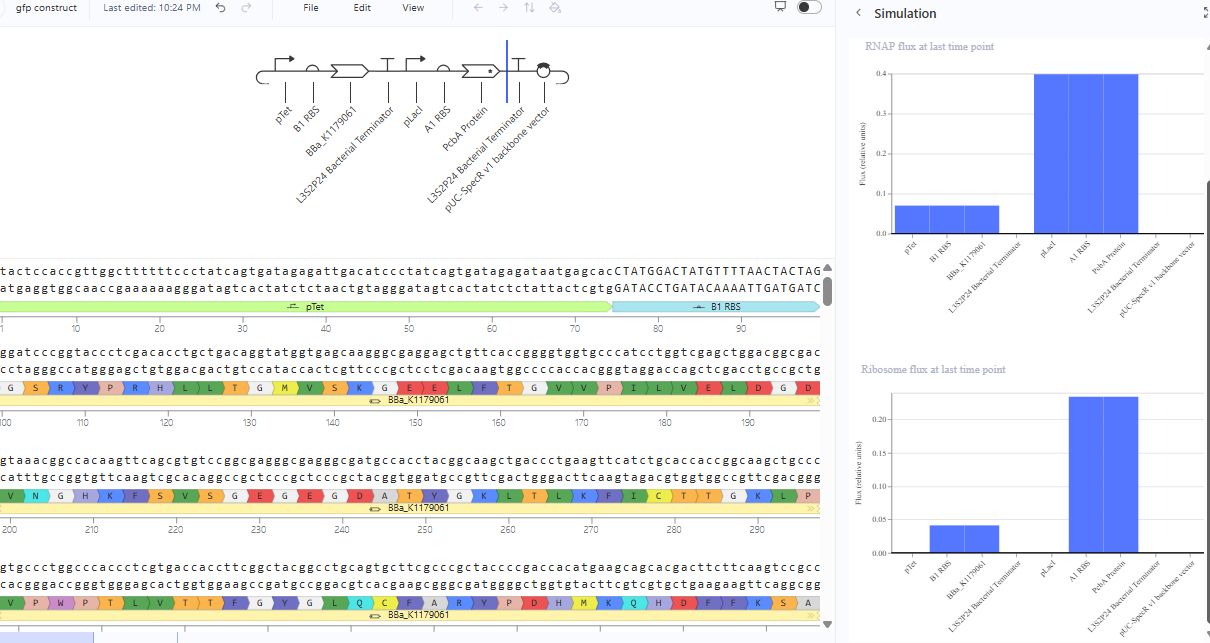

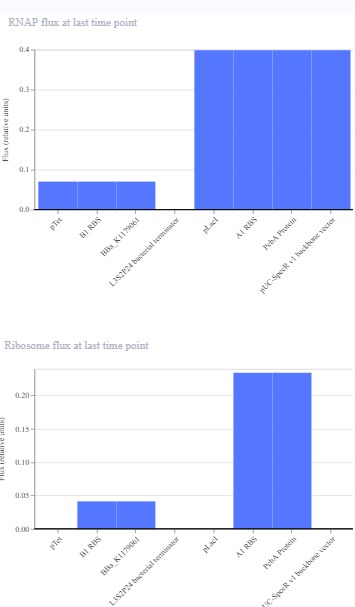

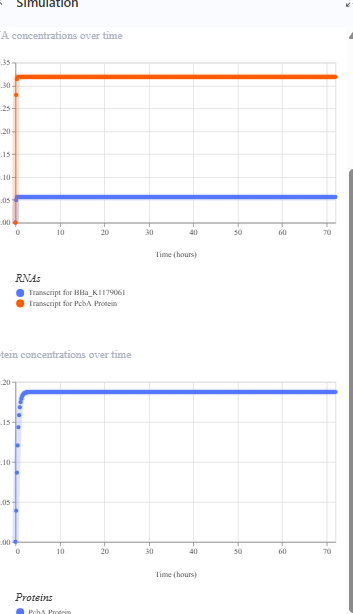

The same construct when the terminator for the eYFP was removed gave the following result.The transcription continues into the next downstream gene.
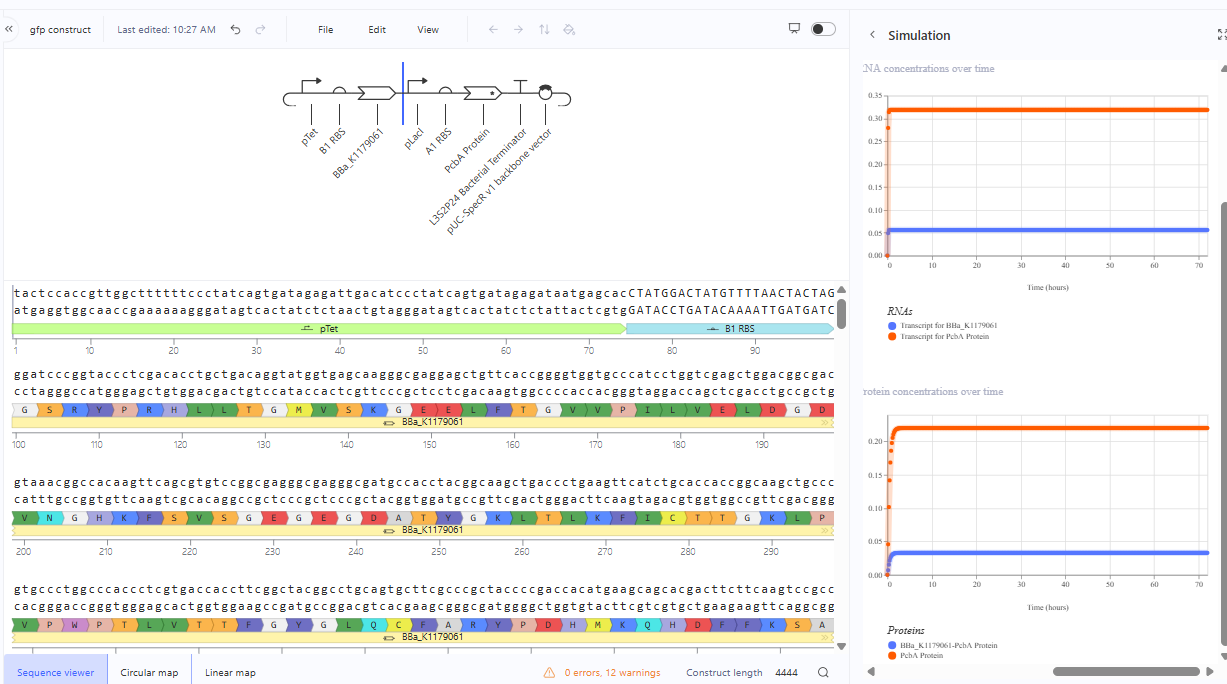

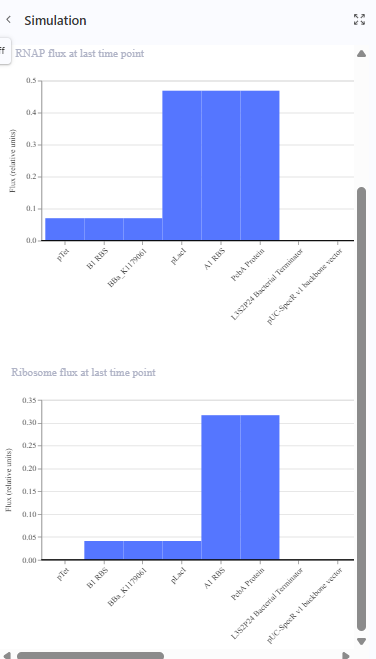

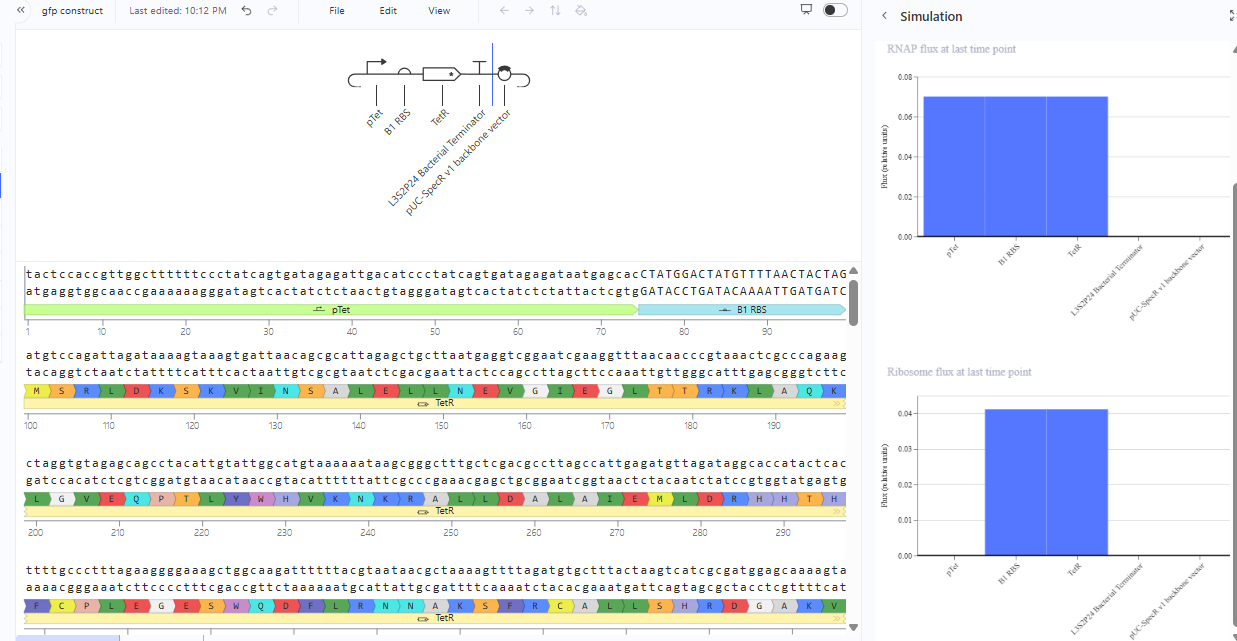
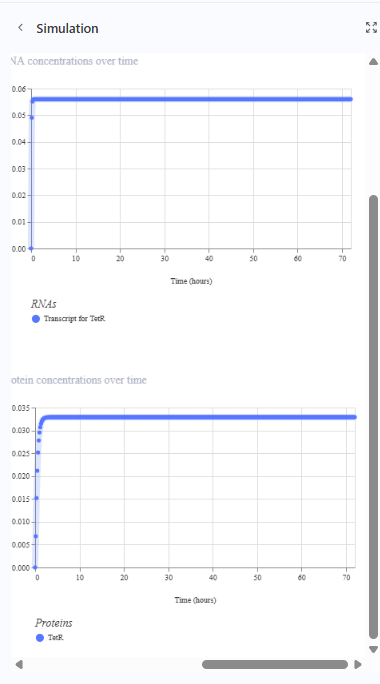
Construction 3: expression of autoregulatory TetR
The production of TetR by its promoter pTetR is autoregulated meaning it represses its own production. The amount of TetR controls the construct.