week-09-hw-cell-free-systems

Homework Part A: General and Lecturer-Specific Questions
General homework questions
1.Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
Advantages: Proteins that are toxic to cells can be produced in cell free systems as there is no fear of damage to cell.
Metabolic pathways can be engineered to bypass unwanted steps and can increase the yield of proteins and allowing the removal of waste products.
The additives like chaperones or lipososmes or detergents can be added directly and conditions like pH and temperatutre optimized as the reactions are taking plavce outside the cell.
Cell-Free Protein Synthesis bypasses time-consuming cloning, cell transformation, and expression optimization steps, allowing for direct protein production from linear DNA (PCR products) in hours rather than days.
Proteins that are toxic to cell like antimicrobial peptides are produced because there is no cell system that can be damaged by these toxic proteins. The production of antimicrobial peptides (AMPs), such as human -defensin-2 and magainin 2, which are designed to kill bacteria, can be produced in high-concentration cell-free systems without damaging the production machinery.
Incorporation of Non-Canonical Amino Acids (NCAAs): Cell-free systems make it easy to incorporate unnatural amino acids site-specifically into proteins for drug discovery, as they do not have to cross a cell membrane.These are amino acids that are incorporated by genetic code expansion and modified tRNA synthetases that recognise amber codon TAG.

2.Describe the main components of a cell-free expression system and explain the role of each component.
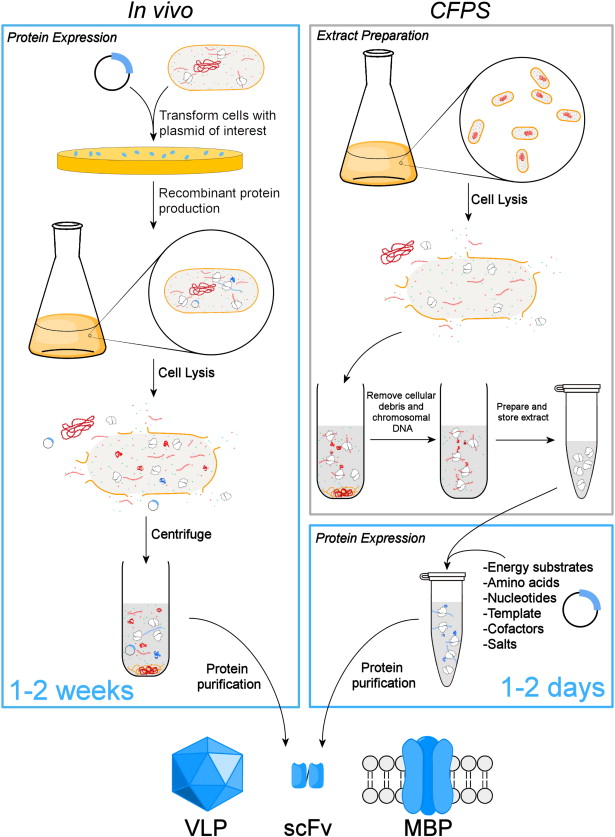
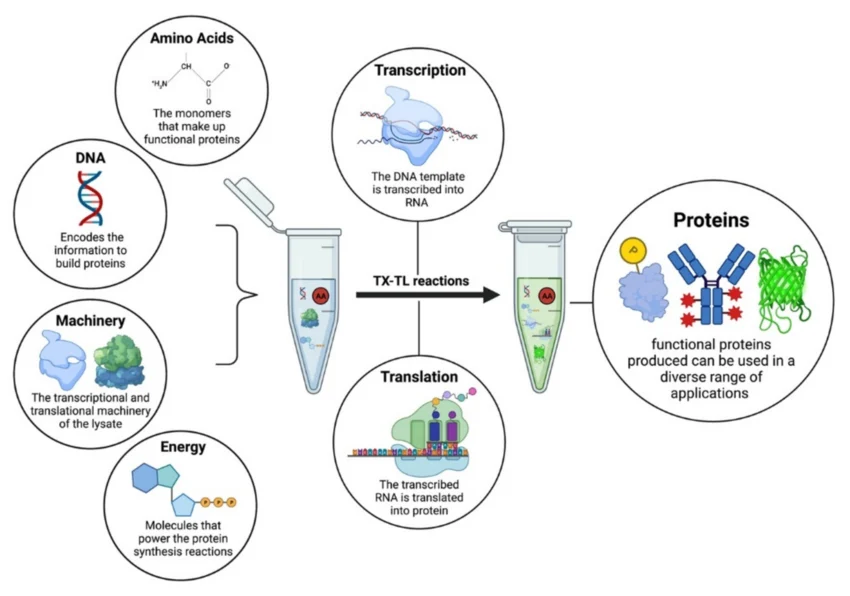
Cell Extract (Lysate or Reconstituted): Contains the machinery required for transcription and translation, including ribosomes, tRNAs, aminoacyl-tRNA synthetases, and initiation/elongation factors. Role: Provides the fundamental molecular machinery to transcribe DNA to mRNA and translate mRNA into protein.
Plasmid DNA, PCR product, or linear DNA encoding the gene of interest. Role: Provides DNA to be transcribed and translated by the cell-free system.
Nucleoside triphosphates (NTPs: ATP, GTP, CTP, UTP), often supplemented with molecules like phosphoenolpyruvate (PEP) or creatine phosphate. Role: Powers the high energy demands of translation (peptide bond formation) and transcription.
Amino Acids: The 20 essential building blocks for protein synthesis. Role: Utilized by ribosomes to assemble polypeptide chains.
RNA polymerase: T7 polymerase added in the mix if transcription and translation has to be carried out. Role: Transcription of DNA to mRNA.
Reaction Buffer and Cofactors: Contains salts (e.g., Mg, K), pH buffers, and sometimes RNase/protease inhibitors. Role: Optimizes the environment for enzymatic activity (e.g., proper magnesium concentration for ribosome function) and protects the RNA and protein products.
Chaperones may be added for protein folding.

3.Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
The energy regenaration is critical as continuous transcription and translation require high ATP concentrations that has to be maintained. The protein synthesis stops in absence of ATP depletion, reducing yield and reaction.
Phosphoenolpyruvate (PEP) System
A highly effective method is using the PEP/pyruvate kinase (PK) system to continuously regenerate ATP from ADP.
Mechanism: Pyruvate kinase catalyzes the transfer of a high-energy phosphate group from phosphoenolpyruvate (PEP) to ADP, forming ATP and pyruvate.
Implementation: Supplement the reaction mixture with 10-30mM PEP and a catalytic amount of Pyruvate Kinase (>10U/ml) along with necessary magnesium and potassium ions.
Result: This system provides a high phosphate potential, significantly extending the duration of protein synthesis compared to simply adding ATP
4.Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
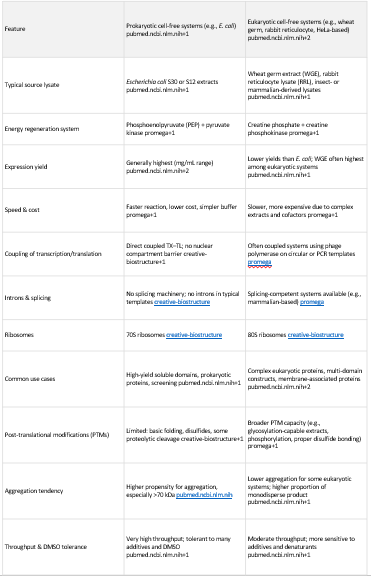

Prokaryotic Cell-Free System Production (e.g., E. coli lysate)
I choose the protein Renilla luciferase as it is used as reporter in many experiments.
Bacterial Cell-Free Protein Synthesis (CFPS) is a widely used, in vitro, open-system technique for producing luciferase proteins (such as Renilla or Firefly luciferase) by using crude cell extracts (typically E. coli) instead of living cells. It enables high-yield production, rapid prototyping of genes, and the ability to produce proteins that might be toxic to living cells.
Eukaryotic Cell-Free System Production
Protein Choice: Human Erythropoietin (EPO)
Why: Eukaryotic systems possess chaperones and machinery for complex folding and post-translational modifications (e.g., glycosylation). Eukaryotic, particularly mammalian or plant-based, systems are necessary to produce active, properly folded transmembrane or glycoproteins, which would otherwise become insoluble aggregates in prokaryotic systems.
5.How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
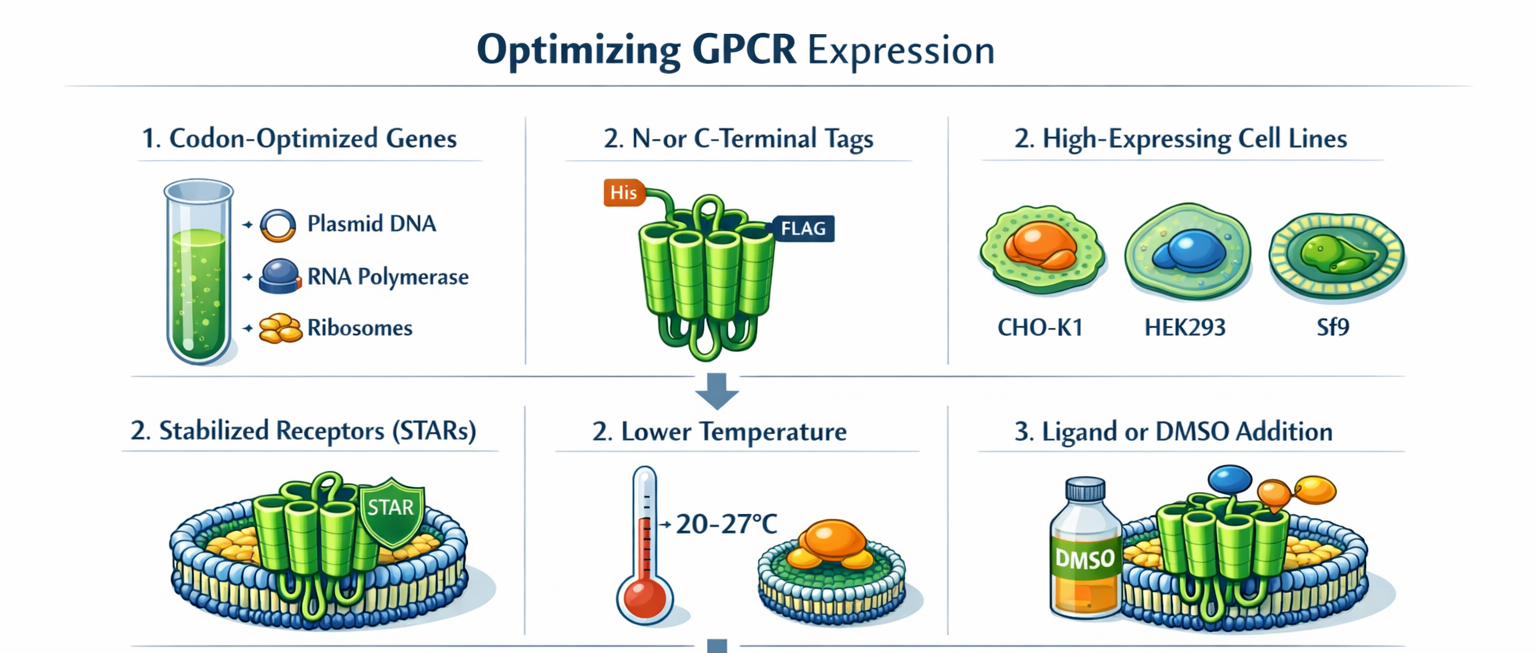
Optimizing Expression of a G-Protein Coupled Receptor (GPCR)
Optimizing G-protein coupled receptor (GPCR) expression involves using codon-optimized genes, N-terminal/C-terminal tagging, and selecting high-expressing mammalian (CHO-K1, HEK293) or insect (Sf9) cell clones. Techniques like using stabilized receptors (STARs), lower culture temperatures (20-27°C), adding ligand or DMSO to increase functional folding, and using detergent-screened membranes ensure high-yield, functional production.
Key Challenges in GPCR Expression
Low Expression Yields: Native GPCRs are present in low abundance; heterologous expression often results in poor yields.
Instability & Aggregation: GPCRs are highly dynamic and flexible, leading to rapid unfolding, degradation, or aggregation once extracted from the lipid bilayer as it is a 7-transmembrane protein and highly hydrophobic.
Toxicity: Overexpression of membrane proteins can overwhelm the host cell’s translocation machinery, leading to cell death.
Incorrect Folding/Post-translational Modifications (PTMs): Prokaryotic systems lack necessary PTMs (e.g., glycosylation), while eukaryotes may incorrectly fold the receptor.
Poor Membrane Targeting: Receptors may be trapped in the Endoplasmic Reticulum (ER) or mislocalized within the cell.
A successful setup combines codon-optimized synthetic genes, fusion partner engineering, and low-temperature, inducible expression in mammalian cells supplemented with pharmacological chaperones.



Homework question from Kate Adamala
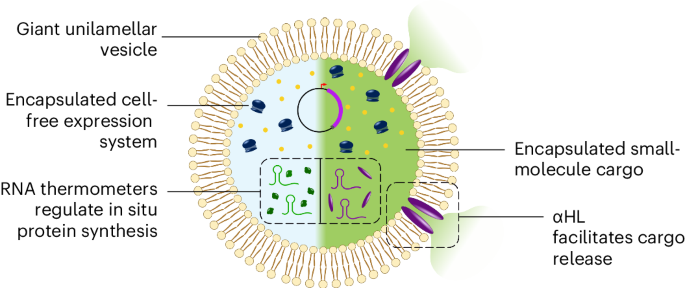
Design an example of a useful synthetic minimal cell as follows:

1.Pick a function and describe it.
a.What would your synthetic cell do? What is the input and what is the output?
The synthetic cell should sense the bacterial biofilm produced and then devour the film formation.Synthetic cells should be engineered with genetic circuits that sense chemical signals (autoinducers) produced by bacteria within a biofilm. Once a specific thtreshold is sensed, the synthetic cells activate, producing enzymes to break down the biofilm formed.
Input:Autoinducers like N-Acyl homoserine lactones (AHLs) or Autoinducer-2 (AI-2), which are produced by pathogenic bacteria to communicate and form protective biofilms.
Output:Lactonase or acylase to degrade the Quorum sensing signals, which halts the biofilm maturation.
b.Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
Yes it can be realized in cell free systems except that the devouring the biofilm cannot be performed in these systems instead these systems release enzymes like lactonases directly into the environment that degrades the extrapolymeric substances thet form the biofilm. Susceptible to sample interference (contamination) in complex environments, unlike compartmentalized synthetic cells.
c.Could this function be realized by genetically modified natural cell?
Yes this function can be realised in genetically modified natural cells. Natural cells like E.coli cells can be engineered to sense the presence of the biofilms and this can trigger the promoter to produce lactonases or other enzymes that degrade biofilm.
d.Describe the desired outcome of your synthetic cell operation. The use of synthetic cells helps to sense biofilm production and degrade the biofilm protecting the bacteria underneath. These bacteria are again susceptible to antibiotic treatment. The harmful bacteria is destroyed and the beneficical ones are not harmed thereby preventing off target effects.
2.Design all components that would need to be part of your synthetic cell.
a.What would be the membrane made of?
Membrane would be made of phospholipids and cholesterol. The cell meembrane is also modified to express receptors specific to Acylhomoserine lactones(AHL)), signal molecules produced during biofilm formation.
b.What would you encapsulate inside? Enzymes, small molecules.
The cell encapsulates a cell-free transcription-translation (TX-TL) system, which contains necessary enzymes (like T7 RNA polymerase, ribosomes), a DNA circuit activated by AHL signalling molecules, bound to the transcriptional regulator Lux R and expresses the enzyme encoded by DNA and a-haemolysin for pore formation.
c.Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
Bacterial Tx/Tl systems are best because the input is AHL that can be sensed by Lux R and initiate gene expression in synthetic cells.
d.How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?)
The membrane is permeable to AHL, synthetic cell expresses a-haemolysin to create pores in membrane for release of lactonases to degrade the biofilm.
3.Experimental details
a.List all lipids and genes. (bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.)
Lipids: Phospholipids, cholesterol
Enzymes: bacterial cell-free Tx/Tl, lactonases, Lux R
Genes: a-hemolysin (aHL)
b.How will you measure the function of your system?
ATP bioluminescence is a highly sensitive method to measure the efficiency of biofilm degradation. Since living cells contain ATP, the degradation of the biofilm directly correlates with a reduction in ATP levels.
Homework question from Peter Nguyen
Freeze-dried cell-free systems can be incorporated into all kinds of materials as biological sensors or as inducible enzymes to modify the material itself or the surrounding environment. Choose one application field — Architecture, Textiles/Fashion, or Robotics — and propose an application using cell-free systems that are functionally integrated into the material. Answer each of these key questions for your proposal pitch:

1.Write a one-sentence summary pitch sentence describing your concept.
A wall hanging made using freeze dried cell free systems that can sense the quality of air within the house.
2.How will the idea work, in more detail? Write 3-4 sentences or more.
The freeze dried cell free systems expressing B-galactosidase is embedded in the paint.A DNA-based sensor is designed using specific transcription factors (TFs) or toehold switches that respond to target indoor pollutants. When a specific pollutant (e.g., formaldehyde from furniture) reaches a certain concentration in the room, it acts as an environmental signal. The Output: The biosensor activates and produces a colorimetric change (e.g., changing from white to blue).
The lyophilized cell free reaction powder is mixed with The powder is mixed with a bio-compatible, permeable binder (like specialized gels or alginate) and applied directly to the canvas or paper using screen printing or inkjet printing, allowing air to reach the sensor. The system requires moisture to function. To initiate the sensor, the user sprays a fine mist of water onto the painted area. Reaction: If a pollutant is present, it diffuses into the spot, triggering the genetic circuit to produce the blue dye, thus changing the color.
3.What societal challenge or market need will this address?
These are programmable materials that contribute to sensing pollutants on rehydation with water and do not contain living, genetically modified organisms, making them safe for indoor environments.These kinds of products are made accesible to general public making tehm aware of the scientific advancements.
These meet the current market needs for a smart, interactive, and sustainable home decor. Cell-free systems allow for the creation of biodegradable smart materials, reducing reliance on conventional synthetic electronics and plastics.
4.How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
The sensors have a limited lifespan (hours to days after rehydration) and need to be replaced, meaning the art piece must allow for new sensor spots to be applied over time.
This can be mitigated by using art designs in such a way that the lyophilized cell free systems are placed in certain pieces of the design that are removable probably that can be replaced with another new set of button of same shape in the design so that the wall hanging and its function is retained for years.
Homework question from Ally Huang
Freeze-dried cell-free reactions have great potential in space, where resources are constrained. As described in my talk, the Genes in Space competition challenges students to consider how biotechnology, including cell-free reactions, can be used to solve biological problems encountered in space. While the competition is limited to only high school students, your assignment will be to develop your own mock Genes in Space proposal to practice thinking about biotech applications in space!
For this particular assignment, your proposal is required to incorporate the BioBits® cell-free protein expression system, but you may also use the other tools in the Genes in Space toolkit (the miniPCR® thermal cycler and the P51 Molecular Fluorescence Viewer). For more inspiration, check out https://www.genesinspace.org/ .
1.Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. (Maximum 100 words)
Musculoskeletal decline in microgravity is a rapid, serious consequence of spaceflight, causing astronauts to lose 1–2% bone mineral density per month and substantial muscle mass, particularly in weight-bearing legs and back.The slow-twitch muscle fibres atrophying the most in the lower limb.The bones become brittle as the microarchitecture degrades. The absence of gravity relates to no work required by bones and muscles to support the body so there is no bone formation and increased degradation.Mitochondrial stress and oxidative stress is more in space that leads to muscle degradation.
These help us understand better serious musculoskeletal diseaes on earth and how bone and muscle cells undergo damage on earth. This deterioration is a major health risk for long-term missions to Mars, as it can cause significant loss of function, increased fracture risk, and kidney stones from high calcium levels.
2.Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
Pax7(Paired box 7)is a crucial transcription factor and the primary marker for muscle stem cells, known as satellite cells (SCs).It is involved in maintenance, self-renewal, and regenerative capacity of skeletal muscle.
3.Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
There is a decrease in expression of Pax7 in muscle tissues in spaceflight. This leads to reduced myogenic potential, decreased cell growth, and overall sarcopenia-like muscle weakening.Microgravity, leads to impairment of the normal functioning of these cells, directly impacting muscle regenerative capacity.The reduction in Pax7 is accompanied by an increase in muscle-wasting markers, such as myostatin, which further drives the degradation of muscle tissue in the absence of gravity.Microgravity causes muscles to experience significantly reduced mechanical load, leading to a decrease in Pax7+ cells and a corresponding decrease in muscle fiber size.
4.Clearly state your hypothesis or research goal and explain the reasoning behind it. (Maximum 150 words)
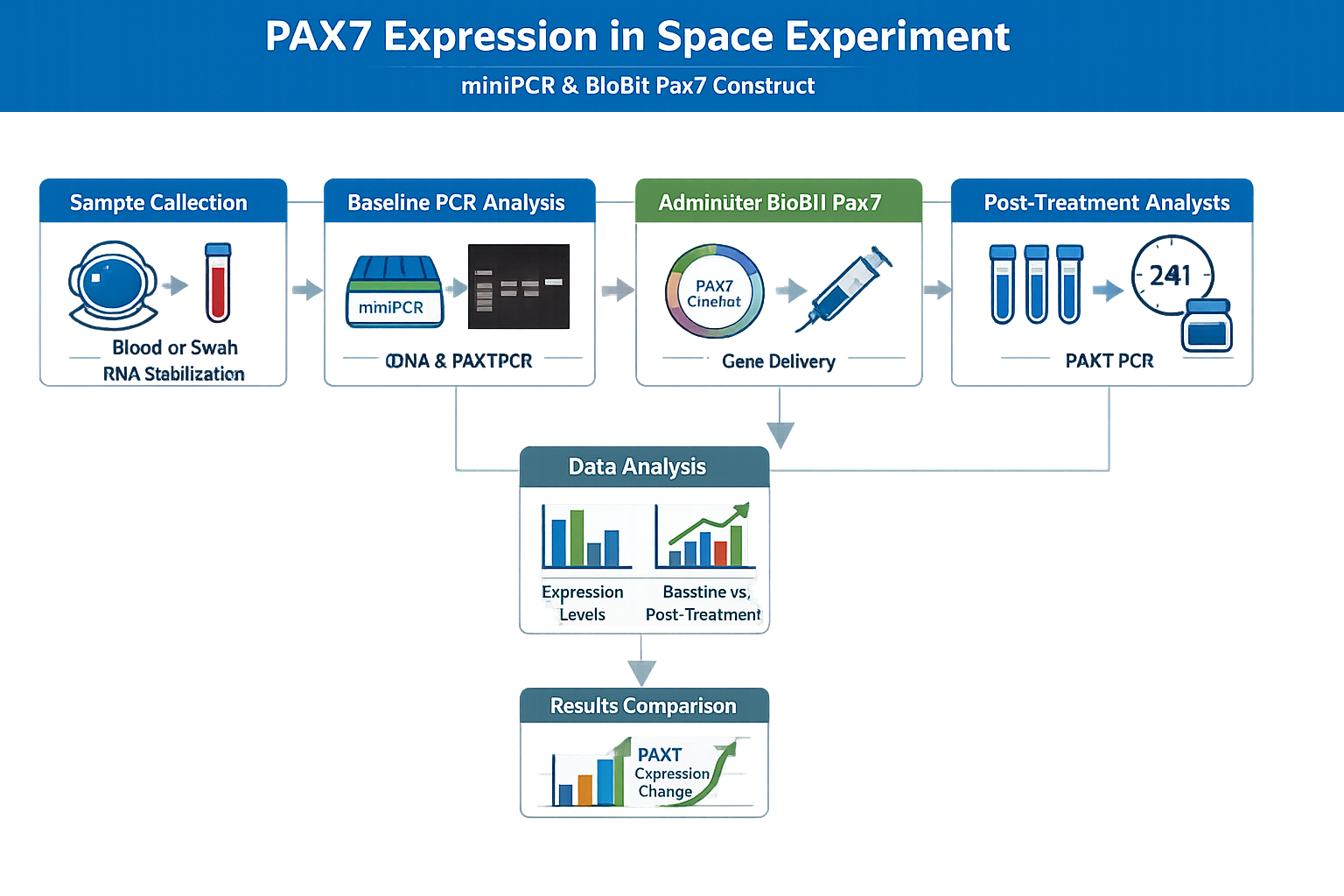
The research goal is to analyze the Pax7 expression in samples from crew members by amplifying it in miniPCR cycler and then administer the pax7 expression sytems lyophilized as BioBits, rehydrated, injected into teh muscle of interest.
Pax 7 expression is crucial for muscle regeneration. Space microgravity and the infrequent usage of muscles in space leads to musculoskeletal decline that can be tried to combat by expressing the Pax7 in cell free systems and injecting them in affected area to regenerate the muscle mass.The DNA sampling using miniPCR cycler can enable a specific dosage of a Pax7-based therapy is effective in promoting regeneration or mitigating atrophy in space.
BioBits® pellets are freeze-dried and contain the essential cellular machinery (enzymes, ribosomes) needed for protein synthesis, activated simply by adding water and the appropriate DNA, making them ideal for space-based, refrigerated-free, on-demand drug synthesis.
The dosage of BioBits and number of times to be administered will depend on the muscle mass loss and time spent in space.These would require further experimental validation.
5.Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)
miniPCR Analysis (Space-Based)
Blood samples are collected and analysed for muscle loss biomarkers before, during, and after spaceflight.
DNA is extracted using a commercial blood DNA extraction kit.
Amplify DNA using mini PCr cycler to analyse the markers for muscle loss.
Controls: Include positive control (known high-expression sample) and negative control (water).
BioBits Pax7 expression
Lyophilized cell-free extract (BioBits) containing E. coli transcription/translation machinery.
The BioBit pellets are rehydrated using DNA from samples of crew members.
Pax7 promoter with GFP is used as construct.
Incubate at 30-37°C for 4-12 hours in the miniPCR machine or heat block.
Visualize fluorescence on smartphone, indicating expression of the construct,to confirm that the synthetic construct is functional in microgravity.
Controls: Negative control (no cDNA), positive control (known muscle tissue sample).
This can be then administered via injections to affected areas.