week-10-hw-imaging-and-measurement


Homework: Final Project
For your final project:
1.Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc.
In the final project the presence of specific AMR gene is detected using aptamer fluorescence.The fluorescence intensity is directly proportional to the amount of the gene present at the site.
The system relies on the COF acting as a quencher for a fluorophore-labeled aptamer. The efficiency of the “turn-on” signal upon target binding must be measured to calculate the detection limit.
2.Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements.
Measure the fluorescence recovery upon target binding (AMR genes)
Determine the lowest concentration of AMR Genes that can be detected.
Ensure the aptamer only binds to the targeted AMR gene/protein.
3.What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail.
Smartphone-based Handheld Readers: 3D-printed attachments for smartphones that integrate a violet light source (LED) and a specialized emission filter, allowing them to serve as portable, low-cost scanners for quantifying fluorescence intensity.
The COF material or a complementary strand acts as a quencher, keeping the fluorophore-tagged aptamer “off.” Upon binding to the target AMR gene, the aptamer undergoes a conformational change, separating the fluorophore from the quencher and initiating a bright “turn-on” fluorescent signal detectable by the handheld device.
Homework: Waters Part 1 — Molecular Weight
1.We will be analyzing an eGFP standard onto a BioAccord LC-MS system to determine the molecular weight of intact eGFP and observe its charge state distribution in the denatured (unfolded) state. The conditions for LC-MS analysis of intact protein cause it to unfold and be detected in its denatured form (due to the solvents and pH used for analysis).
1.Based only on the predicted amino acid sequence of eGFP (see below), what is the calculated molecular weight? You can use an online calculator like the one at https://web.expasy.org/compute_pi/
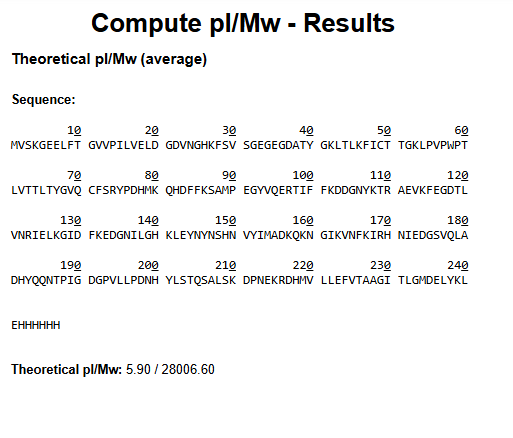
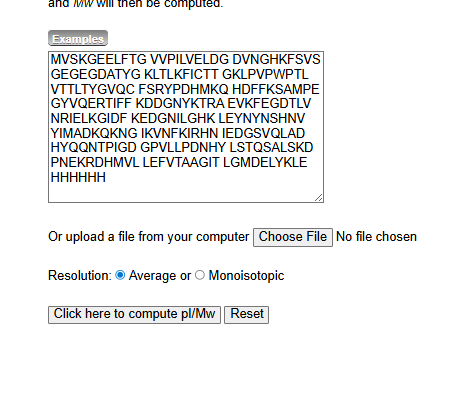



2.Calculate the molecular weight of the eGFP using the adjacent charge state approach described in the recitation.
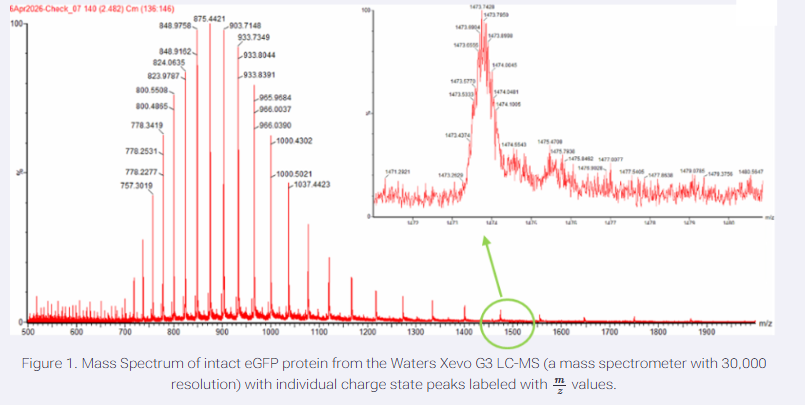

Select two charge states from intact LC-MS data (Figure 1) and:
The peaks around m/z 875.4421 and m/z 903.71
1.Determine z for each (n, n+1)
The formula
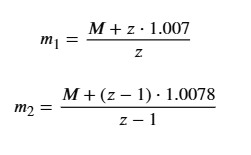
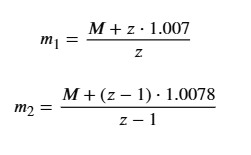
so, z = m2-1/m1-m2, M = z(m1-1)
For m/z m1 = 903.71 and m2 = 875.4421
so, z = m1-1.008/m2-m1
= 903.71-1.008/903.71-875.4421
= +32
Thus, z+1 = +33
2.Determine the MW of the protein using the relationship between m/z,MW and z
The formula derives from
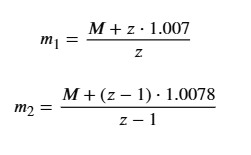
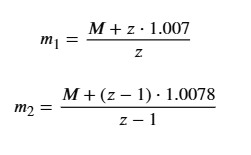
since the difference gives -(z_n / z_{n+1}) = -1/z_n).
so,z = m2-1/m1-m2,
M = z(m1-1)
The molecular weight (MW) of the protein is calculated from the adjacent peaks
using MW ≈ z × (m/z)-(z*1.oo8), where z = 31 for m/z = 903.71 and z = 32 for m/z = 875.4421.
MW ≈ 903.71 × 32-(32*1.008) = 28918.72 - 32.256 = 28886.464 Da from the first peak
and 875.4421× 33 -(33*1.008) = 28889.553 - 33.264 = 28856.289 Da from the second.
average MW ≈ 28871.3715 Da
3.Calculate the mass accuracy of the measurement using the deconvoluted MW from 2.2 and the predicted weight of the protein from 2.1 using these formulae:
Accuracy=[MW experiment-Mw theory]/MW theory
n = m/zn+1/m/zn-m/zn+1
The deconvoluted molecular weight from the adjacent charge state analysis is approximately (using H=1 Da approximation).
Theoretical MW = 28006.60 Da ( from https://web.expasy.org/compute_pi/ results.
MW experiment = 2871.3765 Da
Error = 2871.365 - 28006.60 = 864.7715
Mass accuracy = 864.7715/28006.60*10^6 = 3ppm
3.Can you observe the charge state for the zoomed-in peak in the mass spectrum for the intact eGFP? If yes, what is it? If no, why not?
Homework: Waters Part 2 — Secondary/Tertiary structure
We will analyze eGFP in its native, folded state and compare it to its denatured, unfolded state on a quadrupole time-of-flight MS. We will be doing MS-only analysis (no liquid chromatography, also known as “direct infusion” experiments) on the Waters Xevo G3-QToF MS.
Based on learnings in the lab, please explain the difference between native and denatured protein conformations. For example, what happens when a protein unfolds? How is that determined with a mass spectrometer? What changes do you see in the mass spectrum between the native and denatured protein analyses (Figure 2)?
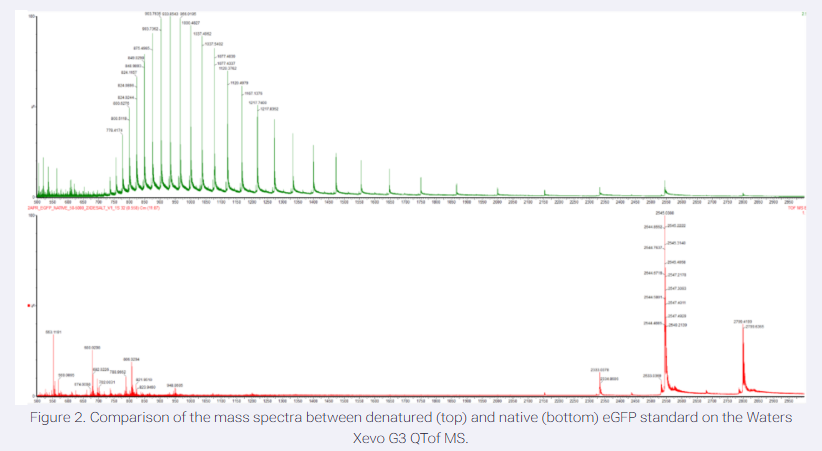

Native proteins maintain their folded, biologically active structures, while denatured proteins unfold into extended, disordered chains due to disruption of non-covalent interactions like hydrogen bonds and hydrophobic effects.Denaturation (e.g., via acid like formic or heat) breaks the highly ordered tertiary or quarternary structure, yielding random coils with exposed residues. Unfolding increases solvent-accessible surface area, often shifting charge statesNative proteins typically show narrow, high-charge distributions (e.g., +4 to +7 here), while denatured ones display broader, lower-charge envelopes (+3 to +5).
Mass spectrometers distinguish via charge state distributions (CSD) in ESI-TOF spectra: native MS yields narrow, symmetric peaks at higher m/z (lower z, intact fold limits adduction); denatured MS shows wider CSD at lower m/z (higher z, flexible chain binds more ions). Isotope resolution sharpens in native due to stable conformers.
Top Spectrum (Green, Denatured): This spectrum shows a broad distribution of many peaks across a wide range of mass-to-charge ratios (m/z), particularly at lower (m/z) values. This is characteristic of a highly charged species, as the charge (z) is high and the mass (m) is the same.
Bottom Spectrum (Red, Native): This spectrum shows fewer, more intense peaks clustered at higher (m/z) values. This indicates a lower charge state distribution, consistent with a compact, folded protein where fewer sites were available for protonation.
Zooming into the native mass spectrum of eGFP from the Waters Xevo G3 Q-Tof MS (see Figure 3), can you discern the charge state of the peak at ~2800 m/z? What is the charge state? How can you tell?
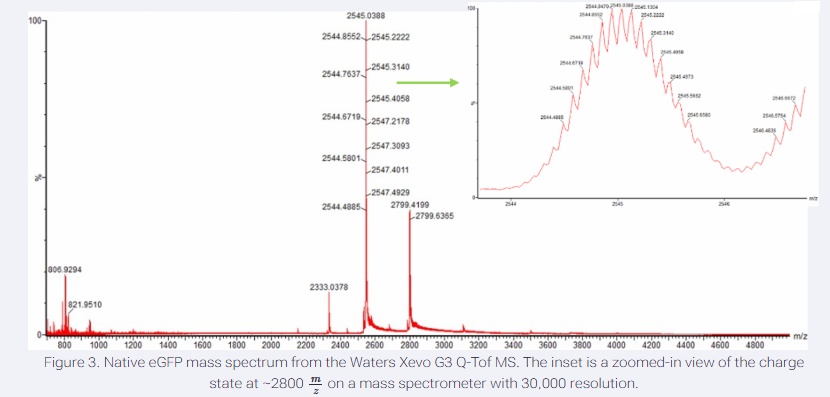

Identify adjacent isotopic peaks m/z1=2545.3140 m/z2=2545.4058
Calculate the difference in (m/z) values
m/z2 - m/z1
=2545.4058 - 2545.3140
= 0.0918
Calculate the charge state
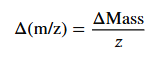
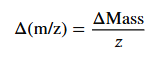
rearranging formula
z= 1.00335/0.0918 = 10.93 rounded to 11.
The charge state of the peak at approximately (2800,m/z) is approximately 11. This is determined by measuring the mass-to-charge difference between adjacent isotopic peaks in the high-resolution inset and dividing the known isotopic mass difference (approx. 1.00335 Da) by the observed (m/z) difference.
Homework: Waters Part 3 — Peptide Map Work - primary structure
We will digest the eGFP protein standard into peptides using trypsin (an enzyme that selectively cleaves the peptide bond after Lysine (K) and Arginine (R) residues. The resulting peptides will be analyzed on the Waters BioAccord LC-MS to measure their molecular weights and fragmented to confirm the amino acid sequence within each peptide – generating a “peptide map”. This process is used to confirm the primary structure of the protein.


The lysine residues are marked in yellow and arginine residues are marked in light blue.
1.How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the in Waters Part I question 1 above. (Note: adding the sequence to Benchling as an amino acid file and clicking biochemical properties tab will show you a count for each amino acid).
There are 20 lysines(K) and 6 Arginines(R) in eGFP.
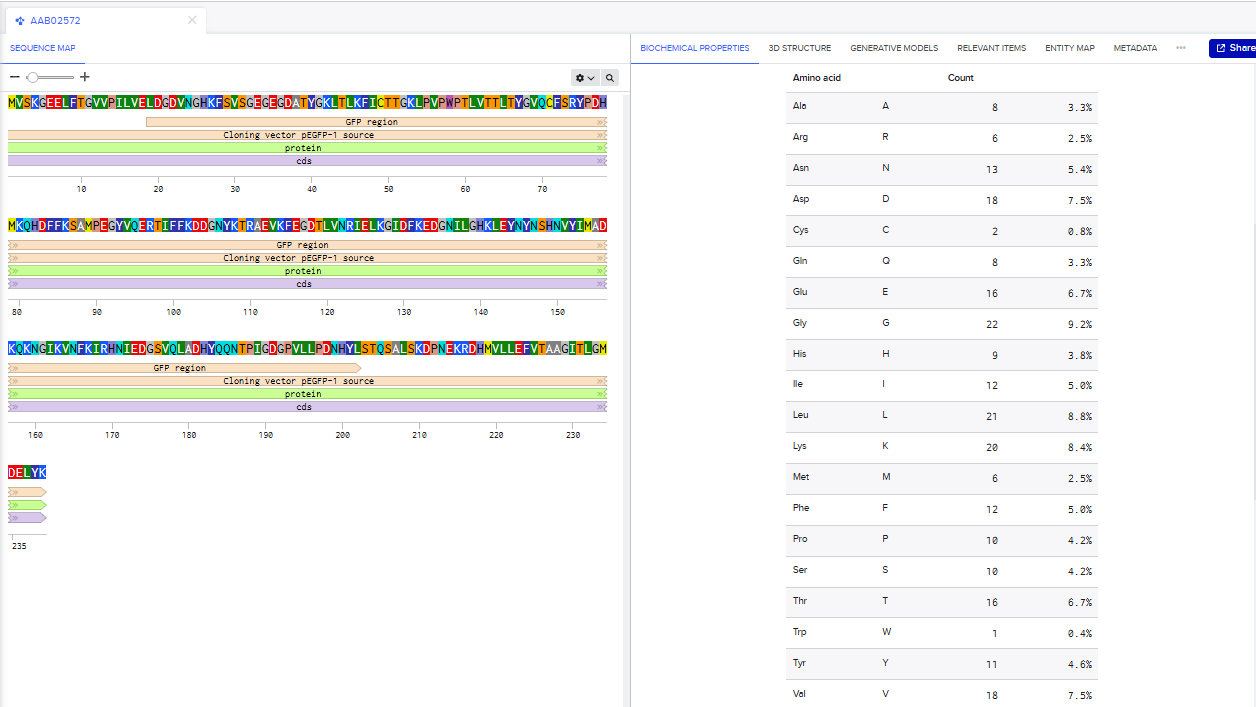

2.There are a variety of tools available online to calculate protein molecular weight and predict a list of peptides generated from a tryptic digest. We will be using tools within the online resource Expasy (bioinformatics resource portal of the SIB Swiss Institute of Bioinformatics) to predict a list of tryptic peptides from eGFP.
How many peptides will be generated from Tryptic digestion of eGFP?
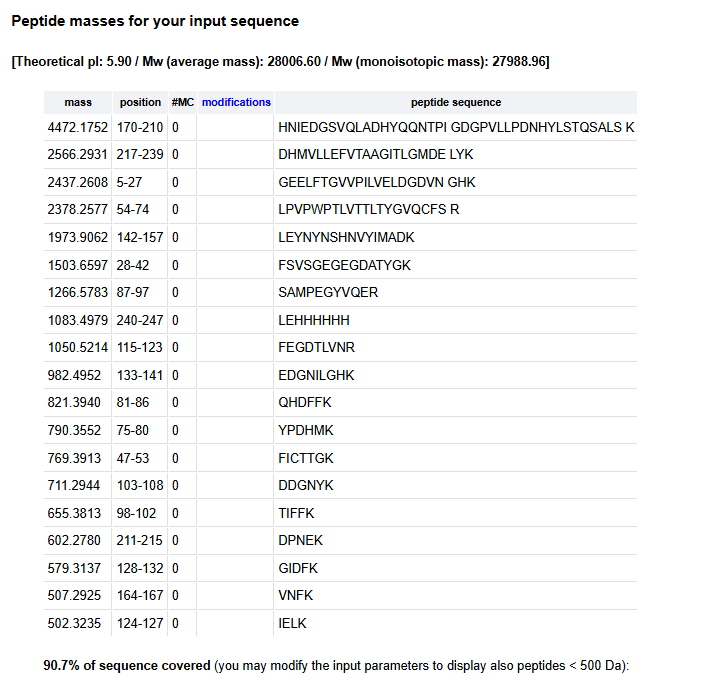

3.Based on the LC-MS data for the Peptide Map data generated in lab (please use Figure 5a as a reference) how many chromatographic peaks do you see in the eGFP peptide map between 0.5 and 6 minutes?You may count all peaks that are >10% relative abundance.
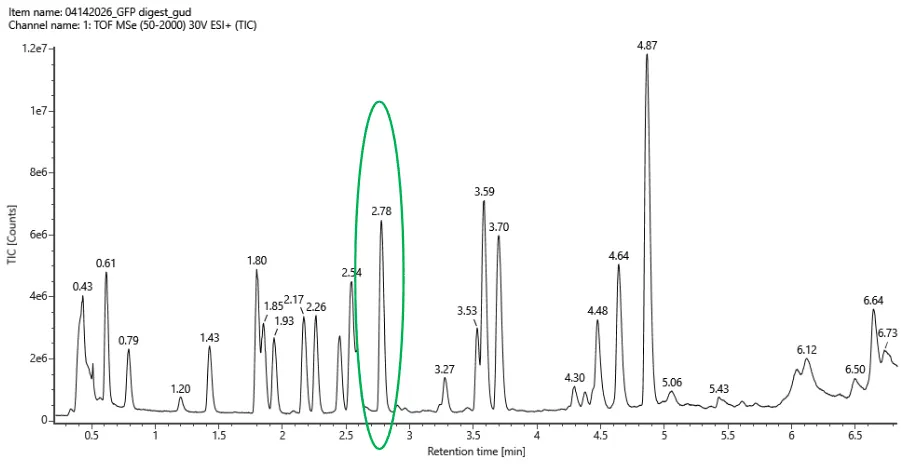

The maximum peak height is at 1.2e7 so all peaks that are >10% relative abundance, should have 0.1*1.2e7 = 1.2e6. There are 21 such peaks.
4.Assuming all the peaks are peptides, does the number of peaks match the number of peptides predicted from Step 2.3 and 2.4? Are there more peaks in the chromatogram or fewer?
There are only 19 peptides predicted. So there are more peaks in the chromatogram. These may be due to impurities in the sample or sample degradation.
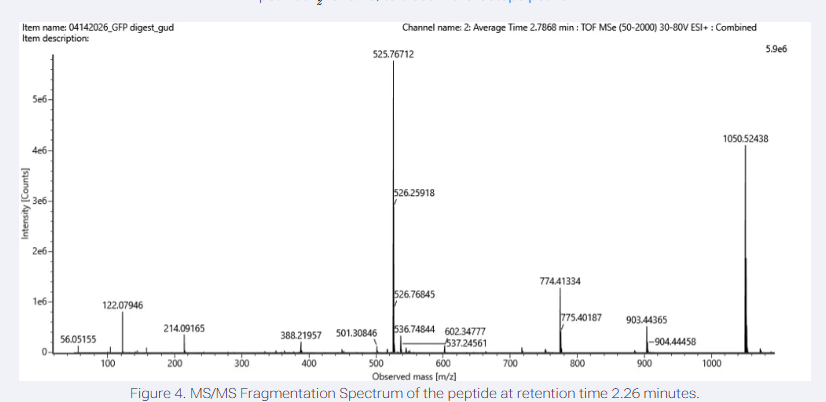
5.Identify the mass-to-charge (m/z) of the peptide shown in Figure 5b. What is the charge (z) of the most abundant charge state of the peptide (use the separation of the isotopes to determine the charge state). Calculate the mass of the singly charged form of the peptide based on its m/z and z ([M+H]+).
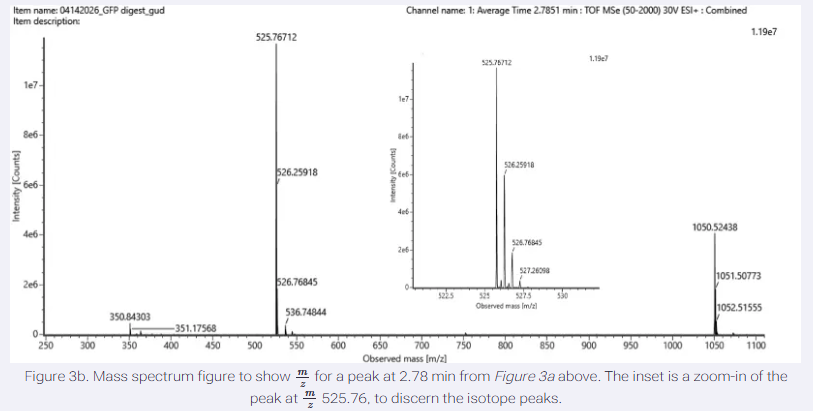

The most abundant peak in Figure 5b (GFP digest spectrum) is at m/z 525.76712, representing the monoisotopic m/z of the peptide’s most abundant charge state.
From the zoomed-in inset in the image, the m/z values of two adjacent major isotopic peaks are approximately 525.76712 and 526.25918.
The difference between these values is 526.25918-525.76712 = 0.492
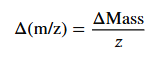
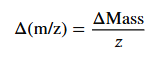
Rearranging the above formula,
z= 1/0.492 = 2.03
Rounded to nearest whole number the charge state (z) is 2+.
m/z = M + zH/z
in the formula replacing m/z = 525.767 and z =2
525.767= M + 2(1.0078)/2
M +2.0156 = 2(525.767)
M + 2.0156 = 1051.534
M = 1051.534 -2.0156
M = 1049.5184 Da
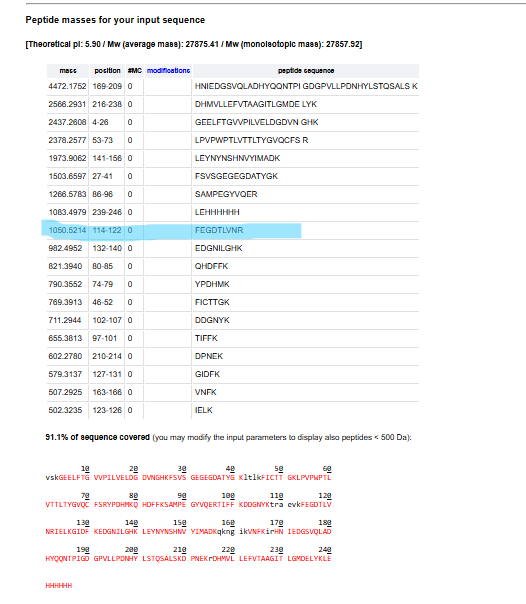
6.Identify the peptide based on comparison to expected masses in the PeptideMass tool. What is mass accuracy of measurement?

Accuracy=[MW experiment-Mw theory]/MW theory
Accuracy = 1050.52438-1050.5214/1050.5214
= 2.83ppm
7.What is the percentage of the sequence that is confirmed by peptide mapping (Figure 6)?
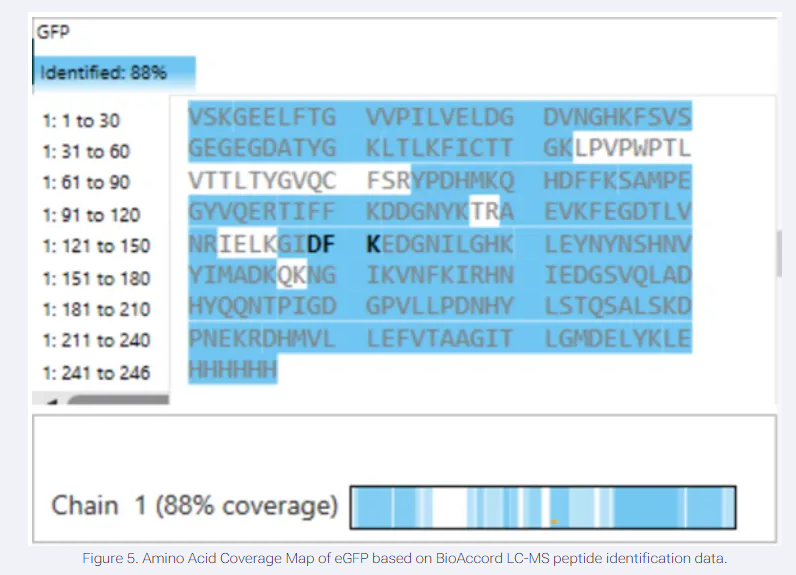

The image shows a peptide mapping result for GFP (Chain 1), explicitly labeling the identified sequence coverage as 88%.
Bonus Questions
8.Can you determine the peptide sequence for the peptide fragmentation spectrum shown in Figure 5c? (HINT: Use your results from Question 2 above to match the peptide molecular weight that is closest to that shown in Figure 5b. Copy and paste its sequence into this tool online to predict the fragmentation pattern based on its amino acid sequence: http://db.systemsbiology.net/proteomicsToolkit/FragIonServlet.html.
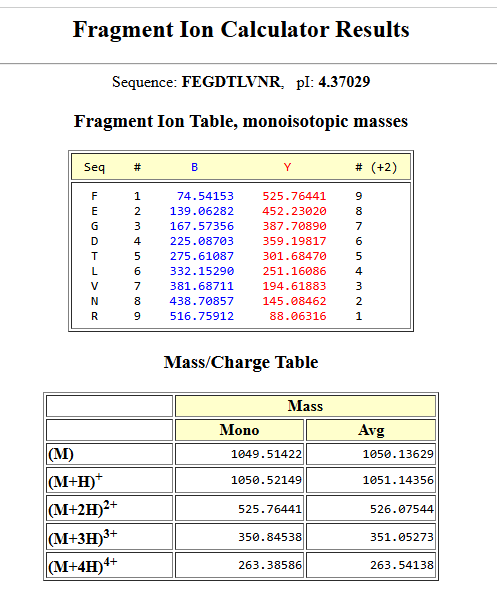

What is the sequence of the eGFP peptide that best matches the MS/MS fragmentation spectrum in Figure 5c?

9.Do the Peptide Map data make sense and do the results indicate the protein is the eGFP standard? Why or why not? Consult with Figure 6, showing the % amino acid coverage of peptides positively identified by their calculated mass and fragmentation pattern.
The results make sense because they are from the fragments of GFP protein as they map to different protions of GFP sequence and it identified 88%.The rest of the sequences were not properly digested probably. There could be presence of post translational modifications that interfered with the expected mass and could not be matched with theoretical sequence.
Homework: Waters Part 4 — Oligomers
We will determine Keyhole Limpet Hemocyanin (KLH)’s oligomeric states using charge detection mass spectrometry (CDMS). CDMS single-particle measurements of KLH allow us to make direct mass measurements to determine what oligomeric states (that is, how many protein subunits combine) are present in solution. Using the known masses of the polypeptide subunits (Table 1) for KLH, identify where the following oligomeric species are on the spectrum shown below from the CDMS (Figure 7):
7FU Decamer 8FU Didecamer 8FU 3-Decamer 8FU 4-Decamer
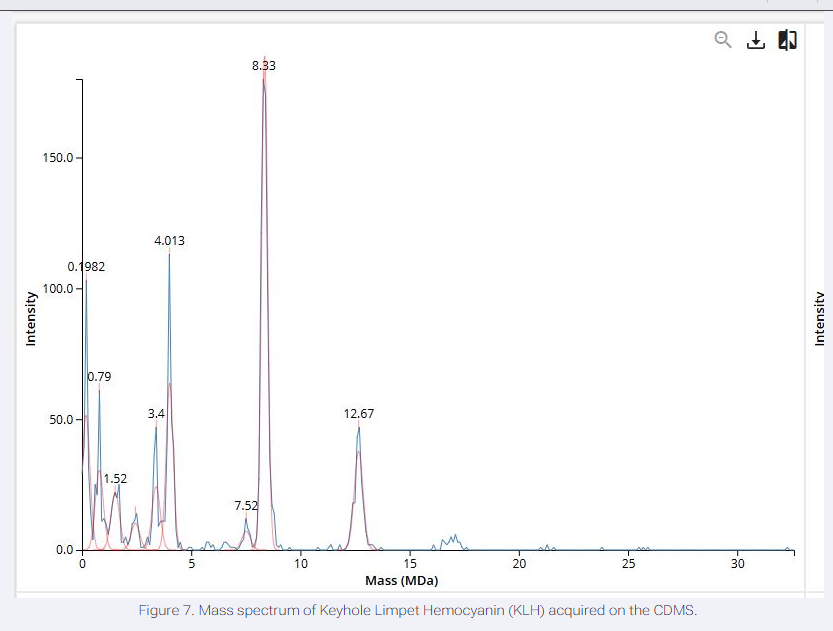

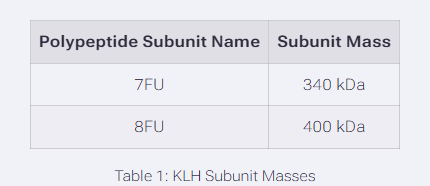

Oligomer Identifications The peak at ~4.01 MDa corresponds to a decamer of 7FU subunits (10 × 340 kDa = 3.4 MDa; observed shift due to glycosylation/adducts common in CDMS).
The peak at ~7.52 MDa aligns with a decamer of 8FU subunits (10 × 400 kDa = 4 MDa; higher oligomers or mixtures may shift observed mass).
The peak at ~8.33 MDa likely represents a di-decamer (20mer) of 7FU (20 × 340 kDa ≈ 6.8 MDa; adducted/glycosylated).
The peak at ~12.67 MDa corresponds to a di-decamer of 8FU (20 × 400 kDa = 8 MDa; further adducted) or larger assemblies.
Smaller peaks (e.g., ~3.4 MDa, ~0.98 MDa) may indicate monomers or fragments, but primary oligomeric species match decameric and didecameric forms typical of KLH
Homework: Waters Part 5 — Did I make GFP?

