week-11-hw-Bioproduction & Cloud Labs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
2.Make a note on your HTGAA webpages including:
a.What you contributed to the community bioart project (e.g., “I made part of the DNA on the bottom right plate”)
I made the upper triangle part before it was modified.8 pixels.
b.What you liked about the project
I liked the way the 1536 plate has been converted for bioart purpose instead of a test validation plate, i also liked the teamwork from people all around the world come together to get a unified artwork done. The best part was involving different fluorescent protein for colouring the pixel.
c.What about this collaborative art experiment could be made better for next year.
Increase the number of plates if possible, so that we can make a bigger design!! I contributed very little, missed the fun.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
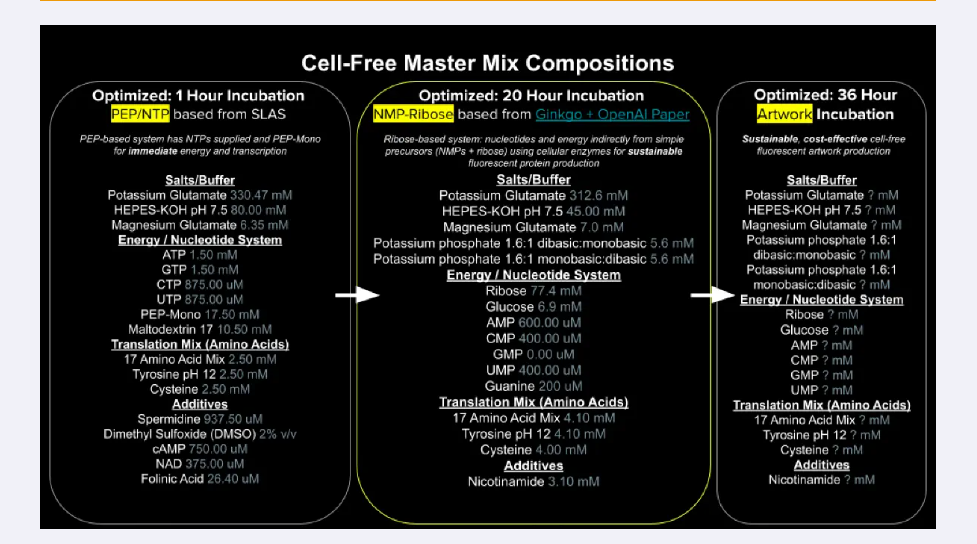

1.Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
E. coli Lysate
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) Salts/Buffer
Potassium Glutamate
HEPES-KOH pH 7.5
Magnesium Glutamate
Potassium phosphate monobasic
Potassium phosphate dibasic
Energy / Nucleotide System
Ribose
Glucose
AMP
CMP
GMP
UMP
Guanine
Translation Mix (Amino Acids)
17 Amino Acid Mix
Tyrosine
Cysteine
Additives
Nicotinamide
Backfill
Nuclease Free Water
Lysate and Buffer System
E. coli Lysate (BL21 (DE3) Star): Contains the essential cellular machinery, including ribosomes, tRNAs, and native enzymes, required to drive the translation of proteins from a DNA template.
Potassium Glutamate: Acts as the primary salt to maintain optimal osmotic pressure and ionic strength for efficient enzyme activity and protein folding.
HEPES-KOH pH 7.5: Functions as a buffering agent to maintain a stable physiological pH throughout the reaction.
Magnesium Glutamate: Provides essential Mg 2+ions, which are critical cofactors for ribosomes, RNA polymerase, and the stabilization of nucleic acid structures.
Potassium phosphate (monobasic/dibasic): Acts as a secondary buffer and a crucial source of inorganic phosphate to support metabolic energy regeneration.
Energy and Nucleotide System
Ribose and Glucose: Serve as sustainable metabolic substrates that are broken down by endogenous enzymes in the lysate to regenerate ATP and other required nucleotides.
AMP, CMP, GMP, UMP, and Guanine: Provide the essential nucleotide precursors that are phosphorylated or converted in situ into triphosphates (ATP, CTP, GTP, UTP) necessary for transcription and translation.
Translation Mix and Additives
17 Amino Acid Mix, Tyrosine, and Cysteine: These serve as the fundamental building blocks required by ribosomes to assemble polypeptide chains during translation.
Nicotinamide: Often acts as a modulator of cellular metabolic pathways and enzyme activity, which can help support extended reaction longevity and stability.
Nuclease Free Water: Provides the solvent medium for the reaction while ensuring the absence of enzymes that would otherwise degrade the DNA or RNA templates.
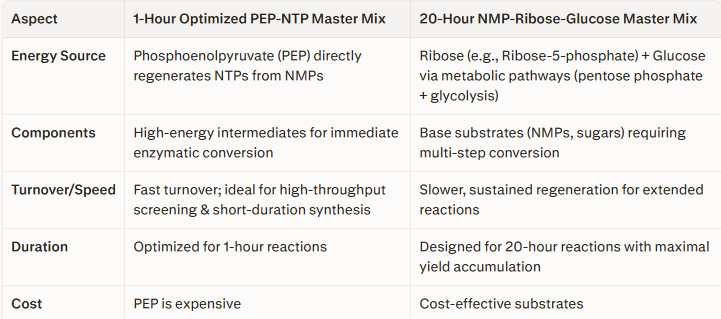
2.Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
1-Hour Optimized PEP-NTP Master Mix
Energy Source: Uses Phosphoenolpyruvate (PEP) to directly regenerate NTPs (Nucleoside Triphosphates) from Nucleoside Monophosphates (NMPs).
Components: Contains high-energy intermediates designed for immediate enzymatic conversion.
Fast turnover for high-throughput screening, short duration cell - free protein synthesis.
PEP is expensive
20-Hour NMP-Ribose-Glucose Master Mix
Energy Source: Relies on Ribose (likely Ribose-5-phosphate) and Glucose, which fuel slower, more complex metabolic pathways (like the pentose phosphate pathway and glycolysis) to generate precursors and energy.
Components: Contains base substrates (NMPs, sugars) that require enzymatic conversion to reach the final product, suitable for cell-free systems relying on metabolic regeneration.
Extended for maximal yield accumulation
Cost-effective substrates; sustained energy

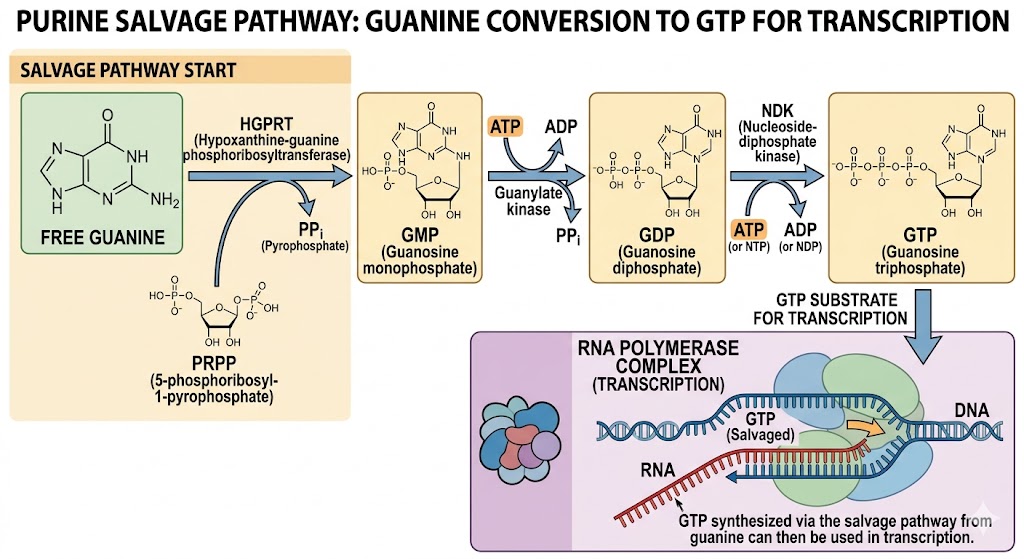
3.How can transcription occur if GMP is not included but Guanine is?
Transcription occurs if GMP is not included but Guanine is present via the salvage pathway.
In this pathway,cells can recycle free guanine or convert it into guanine nucleotides using enzymes like Hypoxanthine-guanine phosphoribosyltransferase (HGPRT). This enzyme converts Guanine and PRPP (5-phosphoribosyl-1-pyrophosphate) into GMP.GMP is then phosphorylated by guanylate kinase into GDP, which is subsequently converted into GTP by nucleoside-diphosphate kinase (NDK).
While GTP (not GMP) is the actual substrate required for RNA synthesis,the GTP synthesized via the salvage pathway from guanine can then be used in transcription.

Part C: Planning the Global Experiment | Cell-Free Master Mix Design
1.Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
sfGFP
mRFP1
mKO2
mTurquoise2
mScarlet_I
Electra2
The amino acid sequences are shown in the HTGAA Cell-Free Benchling folder.
sfGFP (Superfolder GFP):
Property: Extremely fast folding and enhanced solubility.
Effect: sfGFP folds correctly even when fused to poorly folding proteins, making it an ideal, reliable reporter for protein synthesis in E. coli cell-free lysates. It matures quickly, allowing for rapid readout of gene expression.
mRFP1 (Monomeric Red Fluorescent Protein 1):
Property: Rapid maturation compared to its parent protein (DsRed).
Effect: Rapid maturation allows for faster detection of protein synthesis. As a monomer, it does not form tetramers that could hinder the functional activity of fused proteins, allowing for more accurate, quantitative readouts of protein expression levels.
mKO2 (Monomeric Kusabira Orange 2):
Property: High brightness and fast maturation.
Effect: mKO2 offers a bright, orange readout that enables strong signals in cell-free systems, enabling high sensitivity in biosensor applications. Its high brightness is critical when measuring low-level synthesis.
mTurquoise2 (mTq2):
Property: High quantum yield (QY=0.93) and excellent photostability.
Effect: Because mTurquoise2 is one of the brightest cyan fluorescent proteins available, it is an excellent FRET donor. In cell-free systems, it provides a very high signal-to-noise ratio and, due to its mono-exponential lifetime, it is optimal for FLIM (Fluorescence Lifetime Imaging) readouts.
mScarlet_I:
Property: High molecular brightness (product of extinction coefficient and quantum yield).
Effect: As a bright red monomeric FP, mScarlet_I provides high-intensity, red-shifted fluorescence (good for avoiding auto-fluorescence) and is commonly used for quantifying protein production in eukaryotic cell-free systems, maturing rapidly to report on translation.
Electra2:
Property: Good solubility and stability with specific sensitivity to acidity.
Effect: Electra2 behaves as a stable blue fluorescent protein (BFP) suitable for tracking in cell-free systems. However, like other DsRed-derived proteins, it can form aggregates (puncta) in certain conditions and its readout can be affected by the pH of the cell-free reaction.
2.Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
sfGFP
Hypothesis: For sfGFP, increasing Mg²⁺ slightly and maintaining adequate amino-acid supply plus an energy-regeneration system will increase the time-integrated fluorescence over 36 hours because sfGFP benefits from efficient translation and robust folding into its fluorescent β-barrel.
Expected effect: More mature, functional sfGFP accumulates, so fluorescence rises faster and stays higher for longer.
mRFP1
Hypothesis: For mRFP1, adjusting the mastermix to favor a mildly oxidizing environment and sufficient tyrosine / precursor availability will improve chromophore maturation efficiency and increase fluorescence over 36 hours because red fluorescent proteins rely strongly on correct chromophore formation.
Expected effect: A larger fraction of synthesized mRFP1 becomes fluorescent rather than remaining non-matured protein.
mKO2
Hypothesis: For mKO2, increasing energy-regeneration components and maintaining stable pH buffering will improve peak brightness and sustained fluorescence because orange fluorescent proteins are sensitive to prolonged expression conditions and benefit from continued synthesis without pH-driven signal loss.
Expected effect: Higher and more stable orange fluorescence across the 36-hour period.
mTurquoise2
Hypothesis: For mTurquoise2, optimizing Mg²⁺ and potassium glutamate to support ribosome performance will improve folding efficiency and total fluorescent output over 36 hours because cyan proteins can be limited more by expression yield than by intrinsic chromophore formation.
Expected effect: More correctly folded cyan protein accumulates, raising fluorescence amplitude and duration.
mScarlet_I
Hypothesis: For mScarlet_I, supplementing the mastermix with robust energy-regeneration substrates and a slightly oxidizing buffer condition will improve chromophore maturation and ong-term signal retention over 36 hours because bright red proteins often depend on maintaining productive folding while avoiding late-stage signal decay.
Expected effect: Stronger red fluorescence appears earlier and remains detectable longer.
Electra2 Hypothesis: For Electra2, tuning redox balance and ionic strength in the mastermix will improve functional fluorescence yield over 36 hours because this protein’s mature fluorescent state is likely sensitive to folding environment and prolonged incubation stability.
Expected effect: A higher fraction of Electra2 reaches and maintains its fluorescent conformation, increasing total fluorescence.
3.The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here.
I have designed wells based on above hypotehsis for each fluorescent protein in each well.
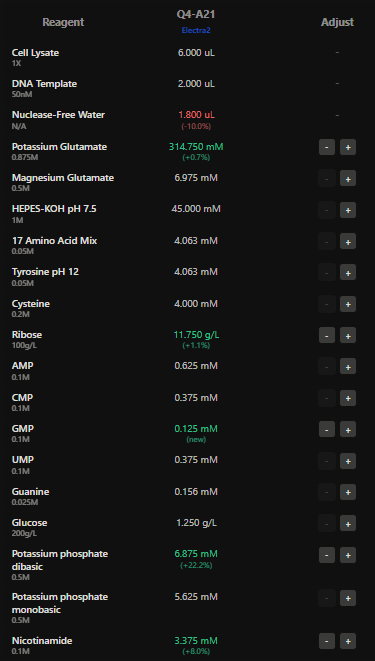

Here I have adjusted the well with potassium glutamate as it is responsible for maintaing ionic strength,and the reagents related to redox balance like glucose, ribose as that improves the fluorescence of Electra2 fluorescence in a 36 hour incubation.
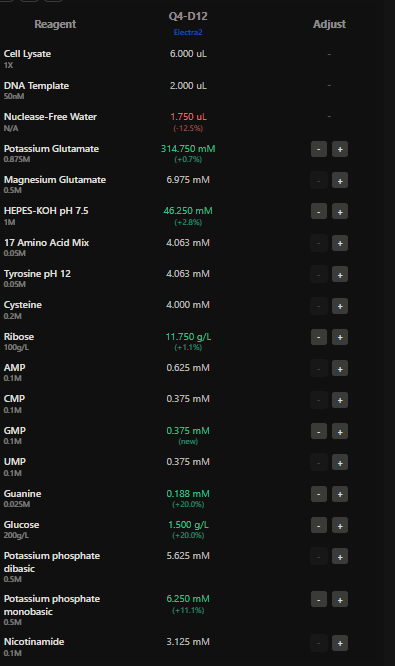

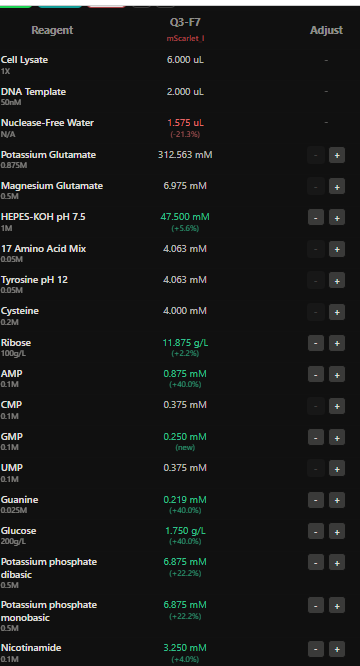

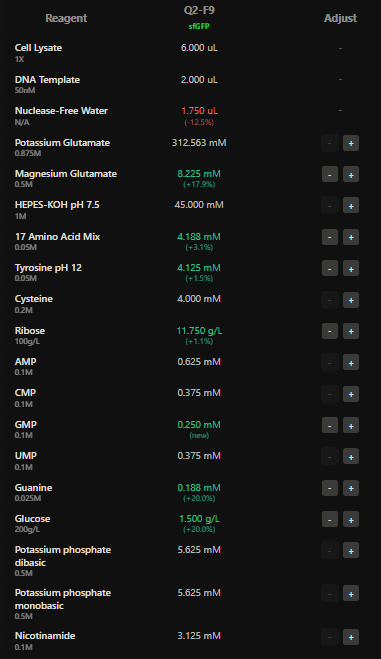

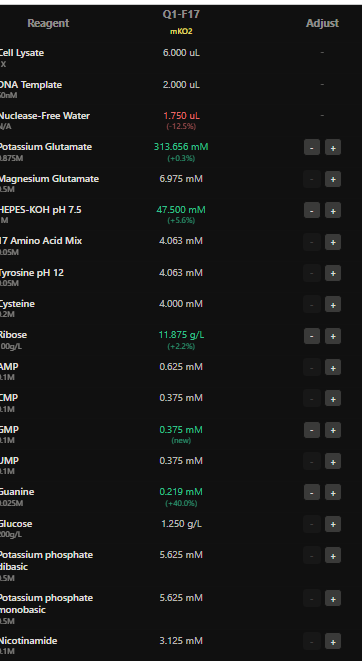

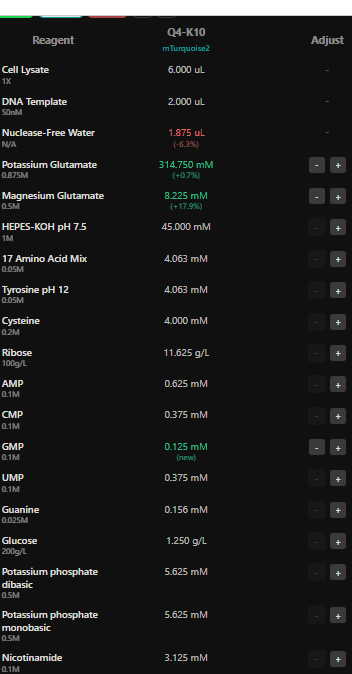

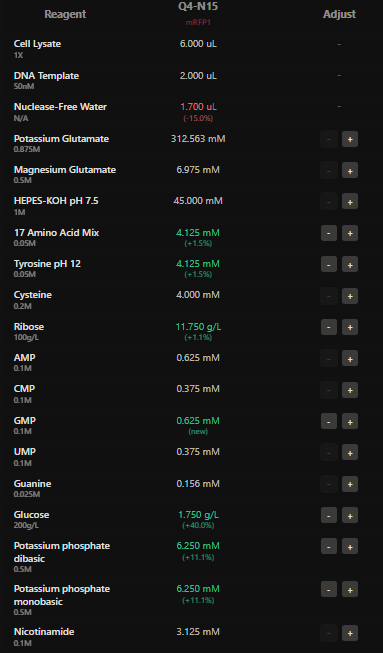

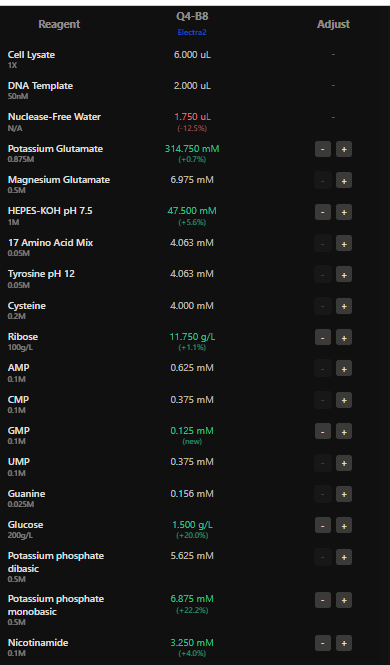

4.The final phase of this lab will be analyzing the fluorescence data we collect to determine whether we can draw any conclusions about favorable reagent compositions for our fluorescent proteins. This will be due a week after the data is returned (date TBD!). The reaction composition for each well will be as follows:
6 μL of Lysate
10 μL of 2X Optimized Master Mix from above
2 μL of assigned fluorescent protein DNA template
2 μL of your custom reagent supplements
Total: 20 μL reaction
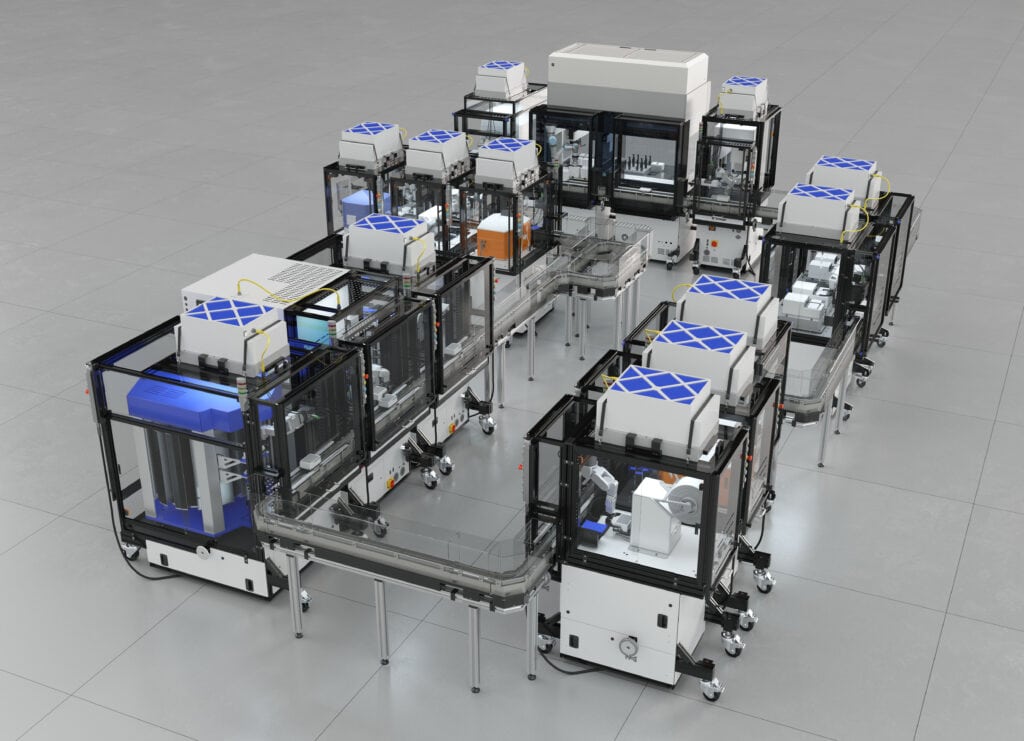
Part D: Build-A-Cloud-Lab | (optional) Bonus Assignment
Use this simulation tool to create an interesting looking cloud lab out of the Ginkgo Reconfigurable Automation Carts. This is just a minimal implementation so far, but I would love to see some fun designs!