Week 4 HW: Protein Design Part-i
Part A : Conceptual Questions
1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
Some of the leanest cuts of meats have ~155gm of protein for 500gm of meat. The number of moles= Total weight/molecular weight = 155/100 – 1.55 moles .
Total molecules= no.of moles x avogadros constant
1.55X 6.023 x 1023
9.33 x 10^23 molecules of amino acids
2. Why do humans eat beef but do not become a cow, eat fish but do not become fish?
Digestion. The many proteins that make up a cow or fish or atleast the visible parts are actually composed of many smaller proteins, these smaller proteins do not form a whole structure, they instead form quaternary structures that are more complex and form things like bones and scales, etc.
When humans consume their meats, these quaternary structures are broken down into simpler amino acids through various enzymes (like proteases) in a process called digestion. These are then used as building blocks for body processes.
Prompt: in relation to protein biology, why can humans eat cows and fish but they dont turn in to one.
3. Why are there only 20 natural amino acids?
Codons. Starting from the LUCA (Last Universal Common Ancestor) , all organisms use the codon system to express genetic changes, with minimal changes among them. There are 64 possible codon combinations. 4 of them coding for each amino acids, however to reduce errors, redundancy is introduced. This redundancy ensures that incase there is an error in one of the nucleotides, the correct amino acid is still produced (synonymous mutation). There could be more amino acids but this would increase errors.
4. Can you make other non-natural amino acids? Design some new amino acids.
Yes, although there are only 20 possible amino acids in biology. Chemistry defines Amino acids as something with a central carbon atom, a NH2 group (amino group) a COOH (carboxyl group), Hydrogen and R (this R group is what differentiates every amino acid from each other).
Designed Amino acid: Electrine. Now previously, I had this idea of using eel elctrocytes to generate bioelectricty. Then it moved onto magnetic proteins that create flux. Electrine is simpler it transfers electrons. It takes glucose, squeezes it’s electrons and then uses quinone and naphthalene to transfer these electrons, via naphthalene rings.
Prompt: i wanna make an amino acid that can use to generate electricity
explain electrine’s mechansim in a simpler way
5.Where did amino acids come from before enzymes that make them, and before life started?
Theories suggest that either the first amino acids came from space, through asteroids that also carried water. There are other theories that lightning had a role to play, where it acted as a catalyst between methane, ammonia, hydrogen, carbon etc. Hydrothermal vents also may have had a role to play. These created the first amino acids that acted as the building blocks for life.
6. If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
The directionality of the α-helix depends on the chirality of the amino acids that compose it. D amino acids would produce a left handed α-helix, these are rare in nature as they are more stable and less reactive to biological enzymes and hence they have no particular use. But these dud to their high stability and less reactivity could be used for artificial purposes.
7. Can you discover additional helices in proteins?
Yes α-helices are the most common, other helices are based on the Residues per turn how many amino acids before it completes one full spiral 2. Rise per residue — how far it advances vertically per amino acid (Å) 3. Hydrogen bond pattern
Prompt: Can you discover additional helices in proteins?
8.Why are most molecular helices right-handed?
Most amino acids are L-amino acids and this causes the structure to be right handed. Theories suggest that L amino acids are more common because of their slightly lower stable energy state and /or their abundance in the first asteroids that came from space, creating life on earth.
10. Why do many amyloid diseases form β-sheets? Can you use amyloid β-sheets as materials?
Amyloid diseases form β-sheets because unfolded protein backbones inevitably fall into the lowest energy state available, a fiber like structure formed of H bonds. Due to their stability they can be used to make relatively strong and stable structures.
Part B : Protein Folding
Based on my hypothetical protein, electrin, Azurin does the same thing. Carries electrons. This was a good starting point.
1AG0_1|Chains A, B|AZURIN|Pseudomonas aeruginosa
AAECSVDIQGNDQMQFNTNAITVDKSCKQFTVNLSHPGNLPKNVMGHNWVLSTAADMQGVVTDGMASGLDKDYLKPDDSRVIAHTKLIGSGEKDSVTFDVSKLKEGEQYMFFDTFPGHSALMKGTLTLK The length of the protein is: 129 aminoacids.
The most common amino acid is: D, which appears 12 times.
There were 250 homologs.
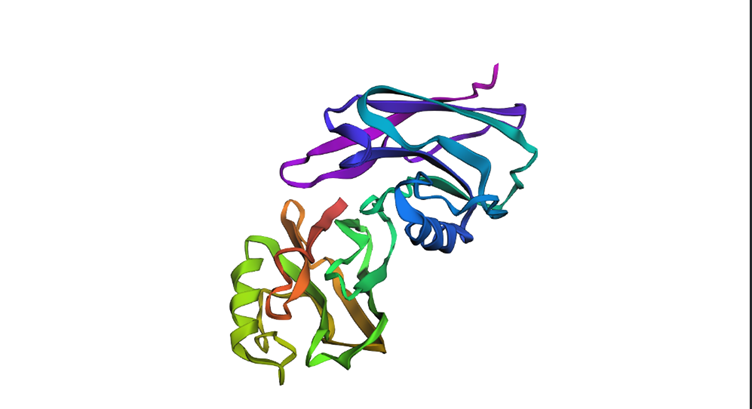
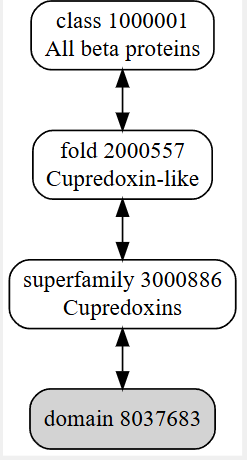
It belongs to a family called cupredoxin, redox proteins that bind to copper ionsThe structure was released on 1997-10-29. The structure seems to be of good quality 2.40 Å is the resolution. This belongs to the cupredoxin like family of structural proteins.

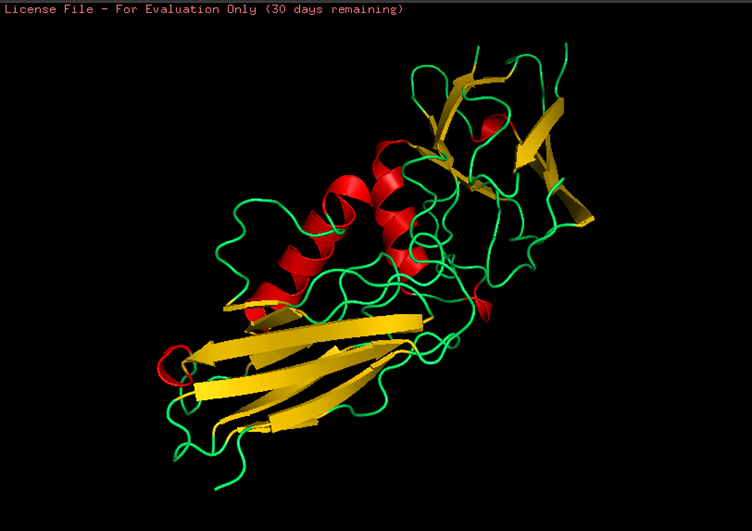
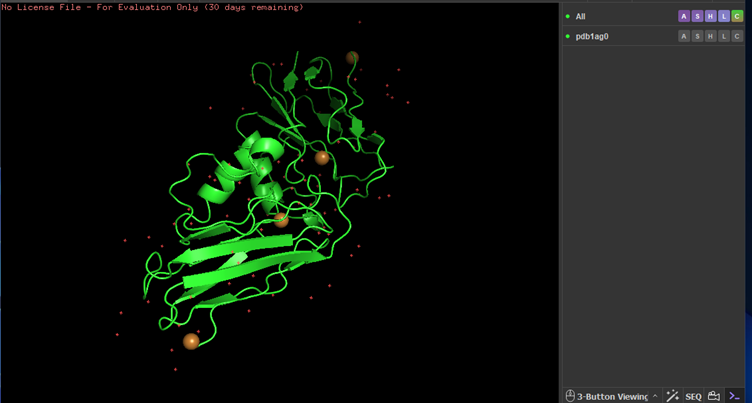
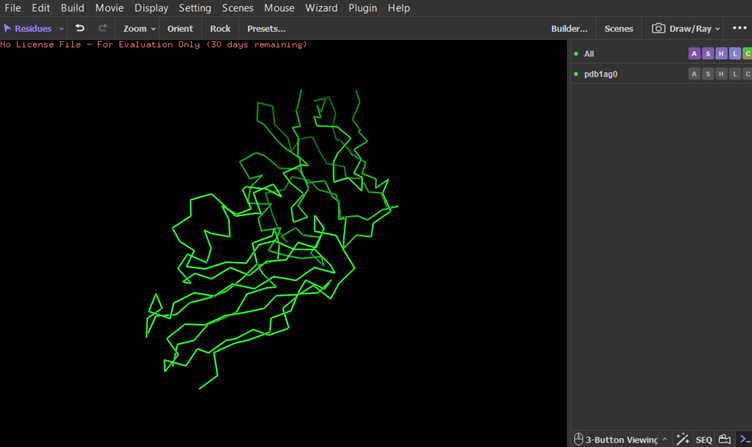
 Colored by secondary structures. The red are alpha helices, the yellow are beta sheets and the green are loops. There seems to be more beta sheets in the protein.
Colored by secondary structures. The red are alpha helices, the yellow are beta sheets and the green are loops. There seems to be more beta sheets in the protein.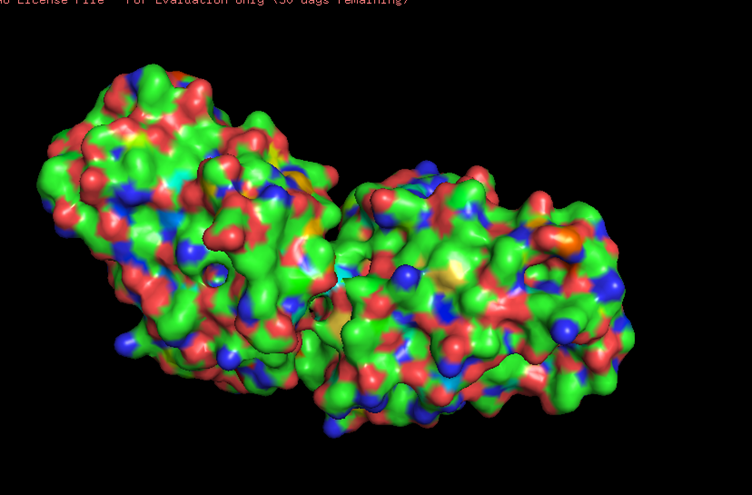
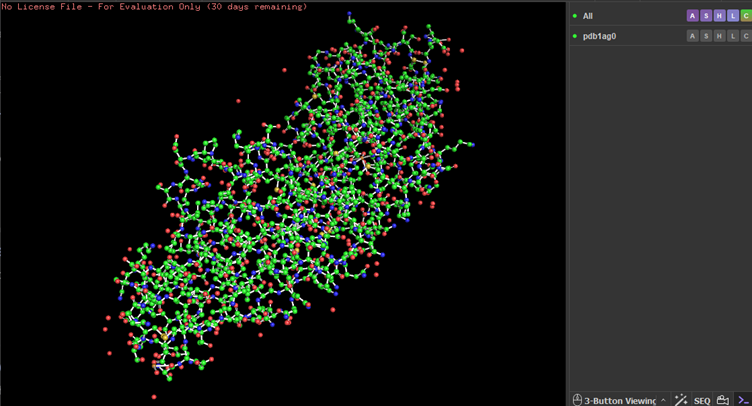
 Looking at the surface structure, there seems to be small holes in the middle of the protein,no major holes. The depression in the middle seems to be a good binding site.
Looking at the surface structure, there seems to be small holes in the middle of the protein,no major holes. The depression in the middle seems to be a good binding site.
Part C
Mutational Scans
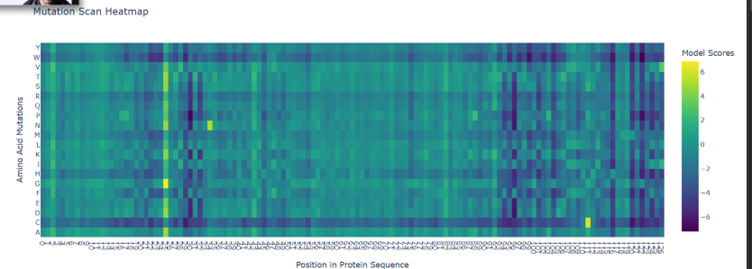

Here the green spots are regions that tolerate mutations pretty well. Ie. If they change doesn’t really affect the functionality much.
Purple are critical regions if there is a mutation here, there is a massive change in the overall structure and functionality.
Yellow regions are pretty sparse here, these regions are good to mutate
There seems to be good mutation candidates for position 25 (Glutamine). At the ends of the protein is where mutations seems to be really bad.
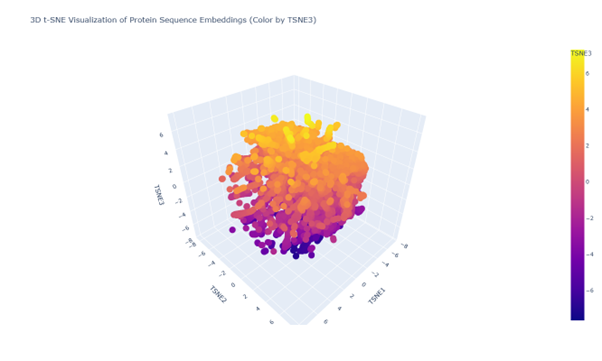
Latent Space Analysis
b. Looking at the 3d PCA plot there seems to be no distinct formation of clusters or neighbourhood. All seem to be similar proteins
C2
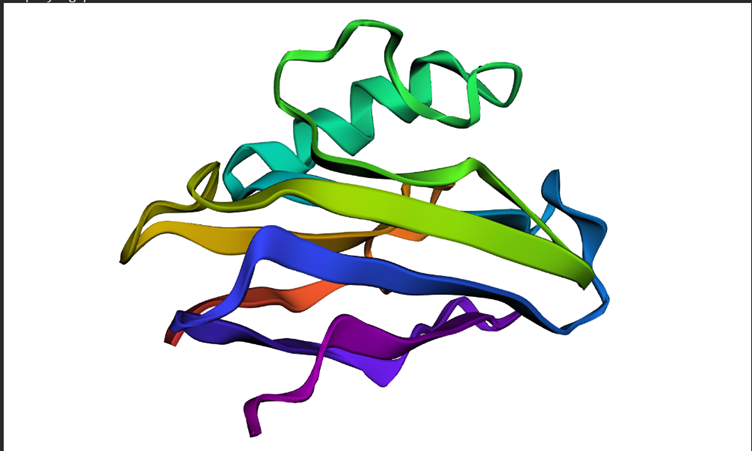
 Overall the structure seems to be similar to the first but looking in depth there seems to be some variations between the predictions and the original structure
Overall the structure seems to be similar to the first but looking in depth there seems to be some variations between the predictions and the original structureSmaller mutations did not create much difference but the larger mutations did. For example I replicated a chunk of the protein within the protein