Week 5 HW: Protein Design Part-ii
Part A: SOD1 Binder Peptide Design (From Pranam)
Part 1: Generate Binders with PepMLM
Before mutation
sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2
MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
After A4V mutation
MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
I used the pepMLM colab to generate 4 binders

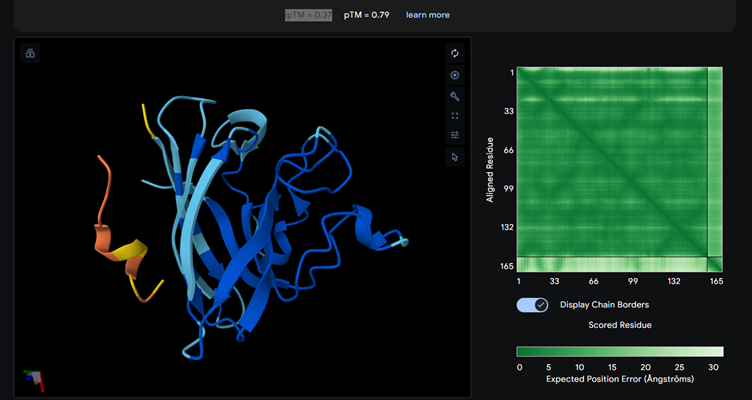
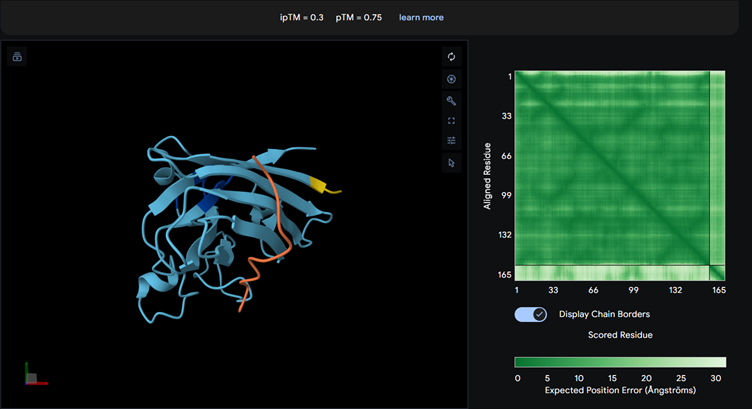
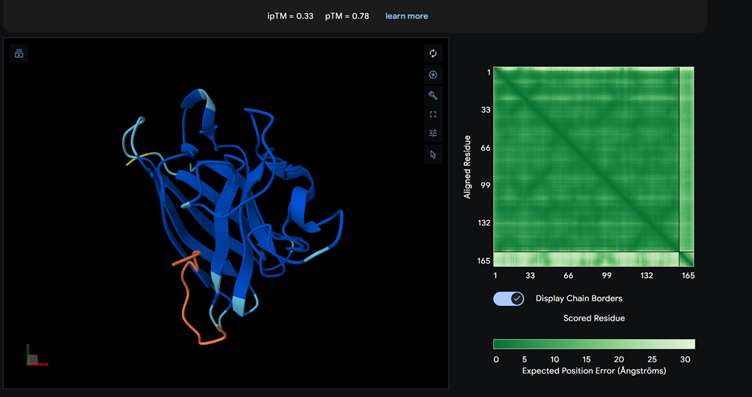
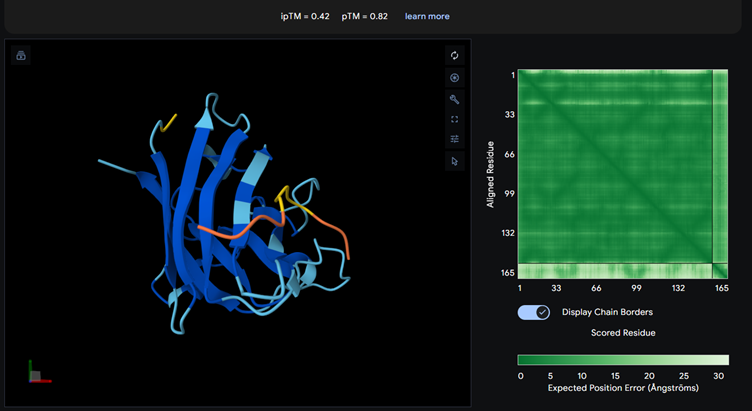
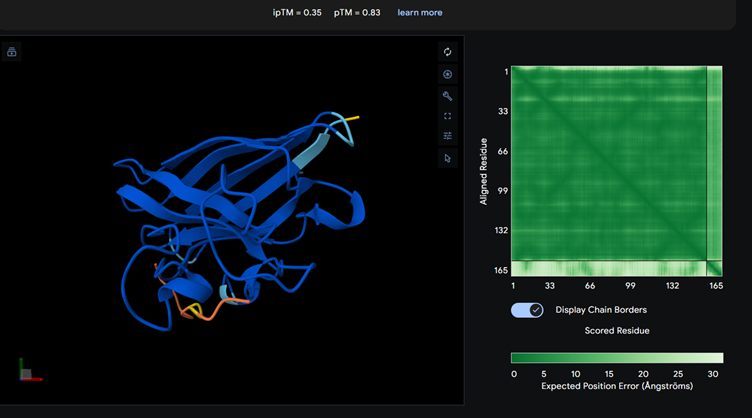
Part 2: Evaluate Binders with AlphaFold3
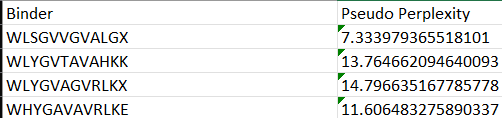
Each of the above binders are number 0 to 3 and the binder_ex is the known SOD1-binding peptide FLYRWLPSRRGG
Most of them appear to be binding near the B barrel structure
Binder_3 had the highest ipTM score this also had the highest perplexity score. This also exceeds the known binder. This measure means that this interaction/structure is more stable and probable than other structures.
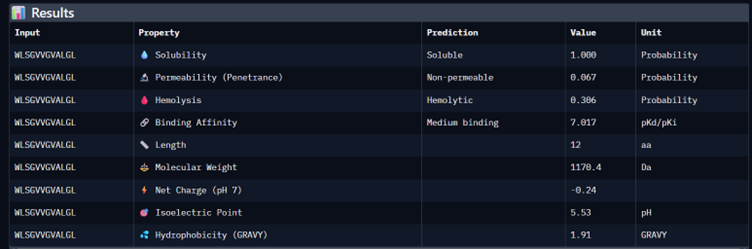
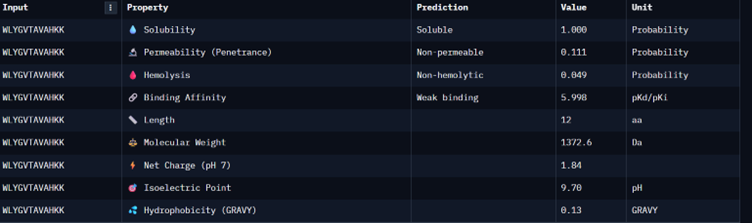
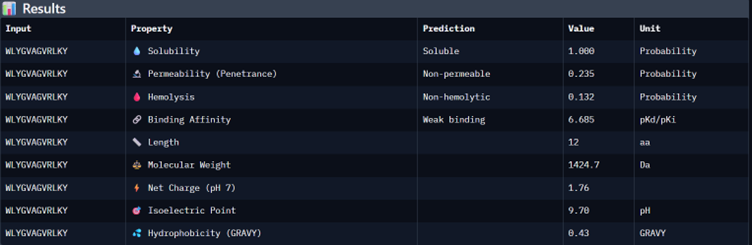
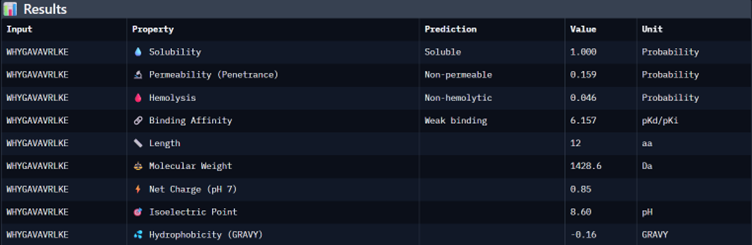
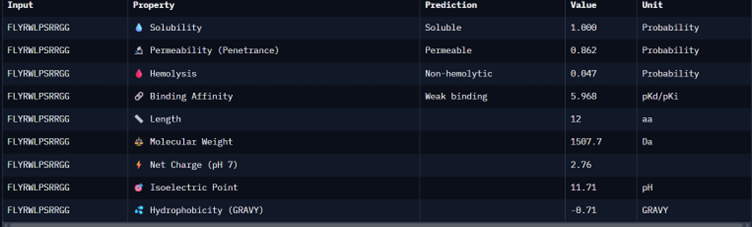
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
I would choose binder_ex i.e FLYRWLPSRRGG because it causes no hemolysis, it is soluble and permeable and although it has a weak binding affinity (most others have it too), it had the 2nd highest iPTM score making it probable structure. This is the best candidate. All other pepetides cause non permeability which makes them a bad candidate for therapy.