Shishir -- HTGAA 2026 Profile
About me
I like Dinosaurs
I like Dinosaurs
Week 1 HW: Principles and Practices
BIO QR CODES My Synthetic Biology proposal is to create Biological QR codes that update based on the body's internal signals. (Think Biomarkers). They act as unique IDs. This is based on the dermal abyss study done by researchers at MIT and Harvard, back in 2017. "Living" tattoos take in the body's metabolic signals as inputs. (This image is from Fluxey, Deviant art: link)
Week 2 HW: DNA Read, Write & Edit
Professor Jacobson: The error rate of DNA polymerase differs in organisms and also because there are different types of polymerase enzymes. Usually this accuracy is 1 in 100,000 bp before proof reading and error correction. (I used an AI prompt to confirm, the prompt: What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?). The length of the human genome is around 3.2 billion bp. To compare, without error correction, this would cause around 32,000 errors if the entire genome is replicated .To correct for this, biology employs DNA repair mechanisms and proof reading that massively reduce the error rate.
Automation Art Now following from previous week’s idea. This gives me more freedom to make the chrome dino art work. I uploaded the chrome dino image into the automation art website. In the end I came up with this design. This is the link to the automation art page
Week 4 HW: Protein Design Part-i
Part A : Conceptual Questions 1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons) Some of the leanest cuts of meats have ~155gm of protein for 500gm of meat. The number of moles= Total weight/molecular weight = 155/100 – 1.55 moles .
Week 5 HW: Protein Design Part-ii
Part A: SOD1 Binder Peptide Design (From Pranam) Part 1: Generate Binders with PepMLM Before mutation sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2 MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ After A4V mutation MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ I used the pepMLM colab to generate 4 binders Part 2: Evaluate Binders with AlphaFold3 Each of the above binders are number 0 to 3 and the binder_ex is the known SOD1-binding peptide FLYRWLPSRRGG
HTGAA Committed Listener (CL) Agreement I am a HTGAA Committed Listener, my responsibilities are: Watching class lectures and recitations Participating in node reviews Developing and documenting my homework Actively communicating with other students and TAs on the forum Allowing HTGAA and BioClub to share my work (with attribution) Honestly reporting on my work, and appropriately attributing and citing the work of others (both human and non-human) Following locally applicable health and safety guidance Promoting a respectful environment free of harassment and discrimination Signed by committing this file to my documentation page/repository,

(This image is from Fluxey, Deviant art: link)
There are many goals to keep in mind to avoid or atleast minimize these risks. Some of them are listed below.
Goal: To ensure this organism does not cause physical harm to its users and the ecosystem.
Method : One such method is to enclose the organism in a hydrogel matrix and also make the organism a synthetic Auxotroph. This matrix maintains the shape of the QR, prevents spread, overlap and supplies the nutrients for growth and maintenance. If the organism escapes the matrix, it cannot survive. (Think Lysine Contingency from Jurassic Park.).
Governance Action: One way to enforce the method is to ensure large regulatory bodies (even labs and policy makers) follow strict guidelines, a ‘Death-on-escape’ standard where the survival rate of the organism is lesser than a set probability.
Goal: Providing users an easy option to opt out of this system, in case of distrust or other disadvantages. This gives the users a choice to stay away from potential constant monitoring and/or fears of data leaks.
Method : Users must have a way to remove the tattoo or deactivate it. The storage system should ensure complete data removal after the deactivation of the tattoo. This removal should not lead to restricted access to healthcare and other services.
Governance Action : Reliable methods to remove the tattoos should be provided when this is being supplied. Common OTCs or software methods of deactivation for temporary cases. Laws should prevent institutions from punishing induviduals who opt out of this system and ndependent audits can ensure that opting out truly removes users from data collection and tracking.
Goal: To prevent access to personal information from unauthorised personnel.
Method: Rather than making the QR code provide information directly, the QR code acts a pointer in a database. This allows a tiered access to information. For example, healthcare workers can access information about the patient’s medical history, blood type or medical insurance but an average civilian can only get limited information, this is left up to the user. Additional safeguards include access logging, authentication, and automatic alerts for unusual activity to prevent silent abuse.
Governance Action: Federal Health Agencies must mandate that any Bio-ID system intergrates with existing National Provider Identifier databases to verify the identity of healthcare workers
Purpose : Currently, Medical reports are protected by laws like HIPAA but this falls into a gray area. This proposal mandates that the QR codes follow a standardised API portocol.
Design: Regulators must define a “Manifest File” for the ID. Developers must implement a Zero-Knowledge Proof handshake. For example, when scanned, the system should only tell you “This person has insurance” rather than handing out the insurance provider’s details unless the scanner has high-level credentials.
Assumptions: This assumes that federal agenecies have the expertise and trust the technology to keep up with it.
Risk of Failure: A “Bio-Data Breach” where a single hack links a person’s digital ID to their permanent biological tattoo, making the leak “un eraseable.”
Success: if successful, medical bracelets become obsolete. There is no risk of loss of ID. An uninteded consqeuence would be a class-divide, people without the tattoo would have slower access to healthcare and other services.
Purpose A Physical Deleteion Requirement where every Bio-ID must be paired with a “Deactivator”
Design: The bacteria should be engineered to a specific chemical that for example, could tigger a lysis circuit and kill the bacteria. This could range from really specific to really broad.
Assumption: This chemical would not be accidentally applied, usually from an unrealted agent (like soap or lotion) and that it is 100% effective at killing all cells.
Risk of Failure : If too broad, a malicious agent could spray the deactivator and erase the ID’s of many people. If too specific, it would be hard to erase the ID.
Success: Complete user autonomy. It ensures that no one is “permanently tracked” and gives people a way to “log out” of the biological world.
Purpose: Many optical sensors (like pulse oximeters) have historically failed for induviduals with different skin tones. For this, a mandatory Inclusion Standard for biological pigments should be introduced.
Design: Before the ID can be supplied, a test on the Fitzpactrick Skin Type scale must be performed and must be certified. Also the ID should be exposed to various environmental conditions for testing.
Assumptions: Assumes that a single set of pigments can be engineered to be high-contrast for everyone and can withstand various conditions.
Risk of Failure: The certification is expensive and makes it accessible to only large corporations which might lead to a centralised system.
Success The code becomes readable by everyone and wouldn’t discriminate.
Purpose: Prevent Bio-IDs from being repurposed for surveillance, policing, advertising, or social control.
Design: Legally restrict Bio-ID usage to clearly defined domains (e.g., emergency healthcare, patient identification, Transactions). Any expansion requires public review and legislative approval.
Assumptions:Assumes lawmakers can adapt quickly to technological change.
Risk of Failure:Governments or corporations may attempt to bypass limits in emergencies.
Success:Stops dystopian “function creep” where helpful tools become control systems.
| Does the option: | National Bio-digital standard | The right to deletion | Optical Scannability Certification | Purpose Limitation Law |
|---|---|---|---|---|
| Enhance Biosecurity | ||||
| • By preventing incidents | 1 | 1 | n/a | 1 |
| • By helping respond | 1 | 1 | n/a | 3 |
| Foster Lab Safety | ||||
| • By preventing incident | 3 | 1 | 3 | n/a |
| • By helping respond | 3 | 1 | n/a | n/a |
| Protect the environment | ||||
| • By preventing incidents | 2 | 1 | 3 | n/a |
| • By helping respond | 2 | 1 | n/a | n/a |
| Other considerations | ||||
| • Minimizing costs and burdens to stakeholders | 2 | 1 | 3 | 1 |
| • Feasibility? | 2 | 2 | 1 | 1 |
| • Not impede research | 3 | 1 | 3 | 1 |
| • Promote constructive applications | 1 | 2 | 1 | 1 |
Based on Everything, I would choose a combination of option 1, 2 and 4 as priority with 3 as an addon later for fairness and accessibility. National Bio-digital Standard provides a technical and legal backbone by enforcing least-privilege access, identity verification and privacy preservation. Wihtout this, the system becomes liable to misuse and leaks.
The Right to Deletion is the best framework for lab and environmental safety, by creating a way to opt out of this system, it supports ethical governance and protects users from long term tracking and limits harm if the system fails. This is essntial for human trust and prevents a dystopian outcome.
Purpose Limitation Law This block the function creep where a healthcare tool becomes a surveillance, policing or commerical control system. This limits the area which the system can creep into, crucial for long-term ethical stability.
Option 1 and Option 3 reduce innovation by prioritising secrurity.
Option 2 greatly enchnaces freedom and autonomy by providing a way to opt out based on the induvidual’s choice. This is an acceptable tradeoff to stability.
Obtaining certifications and audits are expensive, however not being inclusive cause public backlash and becomes far more disadvantageous in the long term. If cost is prioritised, there is an non-economic divide, if inclusiveness is a priority, costs might increase, causing an economic divide.
Technical and Biologcial competence by the public and Federal agencies.
Ethical legitmacy is as important as technical performance.
The public is well informed and receptive to invasive procedures for long term ease and benefit.
Whether Synthetic Biology advances enough to reliably prevent mutations and reverse any global effects or any other effects.
Whether all nations would agree to use this technology in the same way and adhere to the same laws.
How quickly attackers adapt to security systems.
The error rate of DNA polymerase differs in organisms and also because there are different types of polymerase enzymes. Usually this accuracy is 1 in 100,000 bp before proof reading and error correction. (I used an AI prompt to confirm, the prompt: What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?).
The length of the human genome is around 3.2 billion bp. To compare, without error correction, this would cause around 32,000 errors if the entire genome is replicated .To correct for this, biology employs DNA repair mechanisms and proof reading that massively reduce the error rate.
An average protein in humans is around 375 aa long (Brocchieri, 2005). Each of those aa can be 1 of 20 types. Each of those Amino acids are coded by 3 codons on average. Codons are 3 nucleotide sequences that code for an amino acid and there are 64 of them. So, for an average protein 20375 different aa chains of length 375 . The number of different DNA encodings are 3375.
In practice most codons don’t code due to lack of specific tRNAs in the cell. These tRNAs match specific codons to specific amino acids before sending them to the ribosome for assembly. As a result if tRNAs do not recognise the specific codon, it stalls protein synthesis. (AI prompt: what are some of the reasons that all of these different codes don’t work to code for the protein of interest?).
Proteins also follow strict structural rules and if the amino acids change, the structure become unstable leading to protein destruction.
Solid phase phosphoramidite synthesis.
As the of an oligonucleotide increase, the coupling efficiency affects the base pair added, even with a 99.9% coupling efficiency after 200nt the probability of the correct bp being added is around 37% and this decreases further as the length increases. At longer lengths, purification becomes difficult. (Gene Synthesis: Methods and Applications, 2011) (AI prompt : what is coupling efficiency).
As mentioned due to coupling efficiency, long lengths of 2000bp would have a lot of errors and may even prevent the stabilisation of the nucleotide.
I made a benchling account and I followed the protocol for the lambda phage. I could not get the accession number. I just copied the entire sequence from the lambda phage fasta file. Lambda NEB.

And pasted it into benchling.Under the project named Test. Then created a new sequence by pasting the bases from the fasta file.
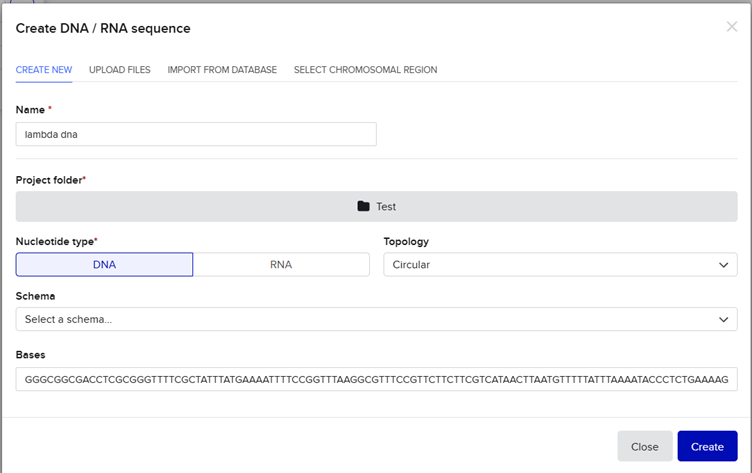

The DNA was then imported and the workspace was split into the entire genome view (on the right) and the individual bases view (on the left).
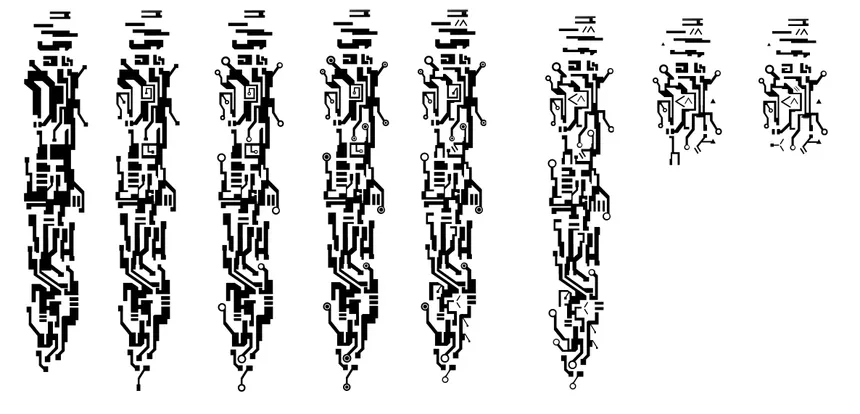
Then I made a list of restriction digestion enzymes that can be used to cut the DNA and named it HTGAA list. Every enzyme showed how many cuts they can produce on this particular genome. Before doing the cuts. I used Ronan’s Automation art website to look for a cool design.
I wanted to initially make the chrome dinosaur art, but that seemed too complex. Then I thought let me make the space invader alien, I failed. I then thought of using the barcoding like pattern to maybe convey a message. In the end I settled for this.
You would need a lot of assumptions for this, but this was the best I could come up with, a sauropod. If there were a lot more horizontal lanes, this would have come better.
The next step was to copy this to Benchling, and the automation art website gave the right enzymes to use. In benchling I made a slightly different verision. (Looks more like a camel)
I chose Phluorin2. Due to the relevance in my proposal. It shows a higher level of brightness changes and has mammalian codon optimization. It is a modified version of GFP and detects ph changes. Low broghtness low ph, high brightness- high Ph.
This is the AA sequence for the protein MSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLSYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KDDGNILGHK LEYNYNEHLV YIMADKQKNG IKVIFQVHHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LHTQSALSKD PNEKRDHMVF LEFVTAAGIT HGMDELYK
Using the Twist Biosciences website, I converted the AA sequence to their nucleotide sequence using the e coli codon usage table
ATGTCAAAGGGGGAGGAACTTTTTACCGGTGTAGTACCAATTCTGGTTGAGCTCGATGGCGATGT TAATGGCCATAAGTTCTCTGTGTCTGGCGAAGGCGAAGGCGATGCCACGTATGGAAAATTAACCC TGAAATTTATATGCACGACTGGGAAATTGCCTGTGCCATGGCCGACACTCGTGACCACTCTTAGT TATGGGGTTCAGTGTTTCAGTCGATATCCAGATCACATGAAGCAGCATGACTTCTTCAAGAGCGC CATGCCAGAAGGCTATGTCCAGGAGCGTACTATCTTTTTCAAAGATGATGGAAATTACAAAACTC GTGCGGAGGTGAAATTCGAAGGCGATACTCTTGTTAATCGGATCGAGTTGAAAGGAATTGACTTC AAGGATGACGGTAACATTTTAGGGCATAAACTGGAATATAATTACAACGAACATCTGGTCTATAT TATGGCTGACAAACAGAAAAACGGCATTAAAGTAATTTTCCAGGTACATCATAATATTGAAGATG GCAGCGTTCAACTGGCAGATCATTATCAACAGAATACGCCTATAGGCGACGGTCCGGTCCTGTTA CCAGATAACCATTACCTGCACACTCAATCAGCGCTTTCGAAGGATCCGAATGAAAAGAGAGACCA CATGGTATTTCTGGAGTTCGTCACAGCGGCGGGGATCACCCATGGCATGGATGAGCTGTATAAAT AA This is with the stop codon at the end.
I used the host organism as e coli since that is what I will probably use to test this gene. Now we use codon optimisation because of anticodon availability. Now, there are around 3 ways to code for a single amino acid but organisms prefer only of those 3 ways because the anticodons that detect that sequence is available in the organism. Hence for efficient translation and minimal errors, it is best to use codon sequences that are optimal for that organism.
For cell dependent systems, I would first insert the nucleotide sequence into a cloning vector. This will be then inserted into an e-coli cell, which will hopefully produce more copies and into more cells. This is then observed in the colonies, just like you would a regular GFP protein.
I followed the instructions and then annotated the inserted sequences.
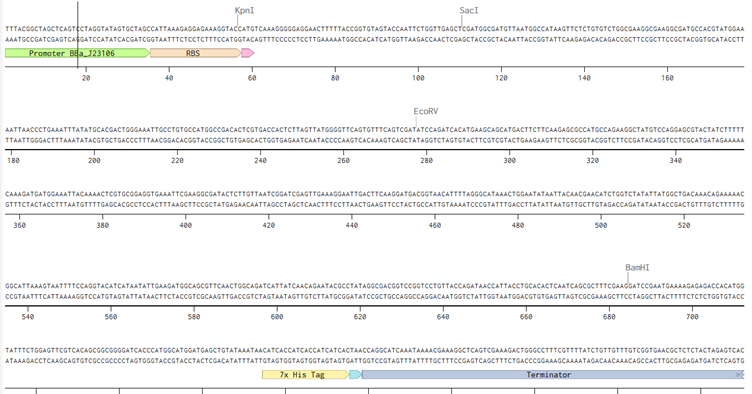

Now since my sequence is different, idk if this would change because through mere observation I noticed that the sequence I have inserted already had a stop codon, the sequence also had the promoter already built in. This is the link Link
After importing my sequence to Twist and using the pTwist high copy number as my cloning vector. I imported the entire genbank file to Benchling.
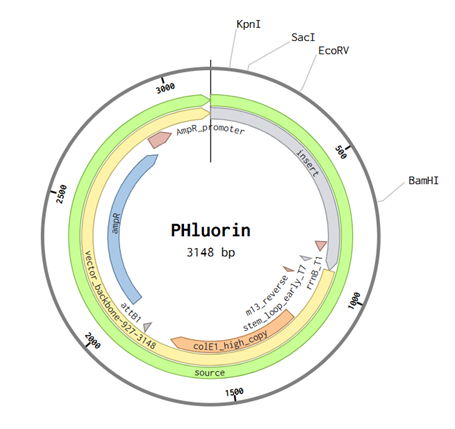

I would like to sequence a lung cell. Two types, one affected by cancer and the other a normal cell. This is to check for points of differences between the cells that could be used as biomarkers. Which can then be converted to a signal input for phluorin, maybe.
I would use Whole Genome Sequencing (WGS). Second Generation NGS.
The input would be DNA extracted from the lung cell and a cancerous lung cell.
Steps: Preparing the library: DNA Extraction Fragmentation End Repair & A-Tailing Adapter Ligation PCR Amplification Size Selection & Cleanup
How are the bases decoded? Step 1: Cluster Generation (Bridge Amplification) • Library fragments bind to complementary oligos on the flow cell. • DNA bends over and binds to nearby oligos. • PCR amplification forms clusters (many identical copies of each fragment). This increases signal strength.
Step 2: Sequencing by Synthesis (SBS) Illumina uses: • Fluorescently labeled nucleotides • Reversible terminator chemistry Process:
Base Calling • Each cluster emits fluorescence. • The color corresponds to A, T, C, or G. • Software converts fluorescence signals into nucleotide sequences. • This produces short DNA reads.
Output Illumina produces: • Millions to billions of short reads • Typically 100–150 bp paired-end reads • Stored in FASTQ format, containing: o DNA sequence o Quality score (Phred score)
The output would be the FASTQ file
The genome of a chicken to make a chicken-saurus. There are genes that are inactive in a chicken that were active in their ancestors (dinosaurs). By activating those genes we can potentially make them express and look more like a dinosaur. I would do this because Dinosaurs are cool and it would be harder to synthesize an entire Trex genome than to modify a chicken’s existing genome. Other reasons could be to understand better the evolution and the bifurcation of birds and reptiles.
I would use CRISPR to edit the DNA cause it is the easiest and most cost effective for these kind of things. The steps include making a guide RNA that detects the sequence in a chicken cell, usually in this case something that targets the repressors and/or promoters /enhancers. So the basic outline would be to identify the repressor that prevents the elongation of tail length for example and then make a guide RNA that detects that repressor and use CRISPR to cut out that part therby making it active
First would be to do a comparative analysis between the chicken and other closely related ancestors (like crocodiles) to identify Conserved developmental genes Regulatory DNA differences, Suppressed ancestral pathways.
Then you would make a genetic map identifying the relevant pathways and the promoters, enhancers and repressors associated with that pathway.
Next we either insert promoters in case of activating dormant genes or removing repressors using CRISPR and the guide RNA. This would be the input.
Limitations: Polygenic Traits, many of the pathways are controlled by many repressors/promoters and thereby editing one gene is not enough, this adds complexity.
Modifying one gene could disrupt other genes downstream potentially reducing viability.
Mosaicism only some cells could be modified and then other repair mechanisms may override the genetic modifications stopping the experiment prematurely.
Brocchieri, L. (2005). Protein length in eukaryotic and prokaryotic proteomes. Nucleic Acids Research, 33(10), pp.3390–3400. doi:https://doi.org/10.1093/nar/gki615.
Gene Synthesis: Methods and Applications. (2011). Methods in Enzymology, [online] 498, pp.277–309. doi:https://doi.org/10.1016/B978-0-12-385120-8.00012-7.
Lopez, M.J. and Mohiuddin, S.S. (2024). Biochemistry, Essential Amino Acids. [online] PubMed. Available at: https://www.ncbi.nlm.nih.gov/books/NBK557845/.
AI used: OpenAI’s ChatGPT 4.1
Now following from previous week’s idea. This gives me more freedom to make the chrome dino art work. I uploaded the chrome dino image into the automation art website. In the end I came up with this design.
Some of the leanest cuts of meats have ~155gm of protein for 500gm of meat. The number of moles= Total weight/molecular weight = 155/100 – 1.55 moles .
Total molecules= no.of moles x avogadros constant
1.55X 6.023 x 1023
9.33 x 10^23 molecules of amino acids
Digestion. The many proteins that make up a cow or fish or atleast the visible parts are actually composed of many smaller proteins, these smaller proteins do not form a whole structure, they instead form quaternary structures that are more complex and form things like bones and scales, etc.
When humans consume their meats, these quaternary structures are broken down into simpler amino acids through various enzymes (like proteases) in a process called digestion. These are then used as building blocks for body processes.
Prompt: in relation to protein biology, why can humans eat cows and fish but they dont turn in to one.
Codons. Starting from the LUCA (Last Universal Common Ancestor) , all organisms use the codon system to express genetic changes, with minimal changes among them. There are 64 possible codon combinations. 4 of them coding for each amino acids, however to reduce errors, redundancy is introduced. This redundancy ensures that incase there is an error in one of the nucleotides, the correct amino acid is still produced (synonymous mutation). There could be more amino acids but this would increase errors.
Yes, although there are only 20 possible amino acids in biology. Chemistry defines Amino acids as something with a central carbon atom, a NH2 group (amino group) a COOH (carboxyl group), Hydrogen and R (this R group is what differentiates every amino acid from each other).
Designed Amino acid: Electrine. Now previously, I had this idea of using eel elctrocytes to generate bioelectricty. Then it moved onto magnetic proteins that create flux. Electrine is simpler it transfers electrons. It takes glucose, squeezes it’s electrons and then uses quinone and naphthalene to transfer these electrons, via naphthalene rings.
Prompt: i wanna make an amino acid that can use to generate electricity
explain electrine’s mechansim in a simpler way
Theories suggest that either the first amino acids came from space, through asteroids that also carried water. There are other theories that lightning had a role to play, where it acted as a catalyst between methane, ammonia, hydrogen, carbon etc. Hydrothermal vents also may have had a role to play. These created the first amino acids that acted as the building blocks for life.
The directionality of the α-helix depends on the chirality of the amino acids that compose it. D amino acids would produce a left handed α-helix, these are rare in nature as they are more stable and less reactive to biological enzymes and hence they have no particular use. But these dud to their high stability and less reactivity could be used for artificial purposes.
Yes α-helices are the most common, other helices are based on the Residues per turn how many amino acids before it completes one full spiral 2. Rise per residue — how far it advances vertically per amino acid (Å) 3. Hydrogen bond pattern
Prompt: Can you discover additional helices in proteins?
Most amino acids are L-amino acids and this causes the structure to be right handed. Theories suggest that L amino acids are more common because of their slightly lower stable energy state and /or their abundance in the first asteroids that came from space, creating life on earth.
Amyloid diseases form β-sheets because unfolded protein backbones inevitably fall into the lowest energy state available, a fiber like structure formed of H bonds. Due to their stability they can be used to make relatively strong and stable structures.
Based on my hypothetical protein, electrin, Azurin does the same thing. Carries electrons. This was a good starting point.
1AG0_1|Chains A, B|AZURIN|Pseudomonas aeruginosa
AAECSVDIQGNDQMQFNTNAITVDKSCKQFTVNLSHPGNLPKNVMGHNWVLSTAADMQGVVTDGMASGLDKDYLKPDDSRVIAHTKLIGSGEKDSVTFDVSKLKEGEQYMFFDTFPGHSALMKGTLTLK
The length of the protein is: 129 aminoacids.
The most common amino acid is: D, which appears 12 times.
There were 250 homologs.
It belongs to a family called cupredoxin, redox proteins that bind to copper ions
The structure was released on 1997-10-29. The structure seems to be of good quality 2.40 Å is the resolution. This belongs to the cupredoxin like family of structural proteins.
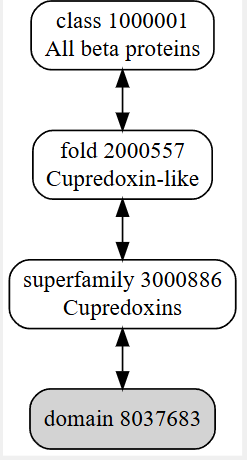

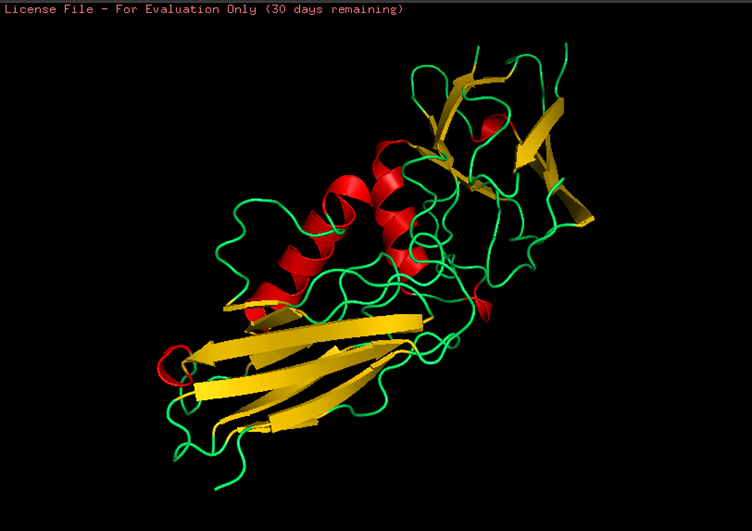
 Colored by secondary structures. The red are alpha helices, the yellow are beta sheets and the green are loops. There seems to be more beta sheets in the protein.
Colored by secondary structures. The red are alpha helices, the yellow are beta sheets and the green are loops. There seems to be more beta sheets in the protein.
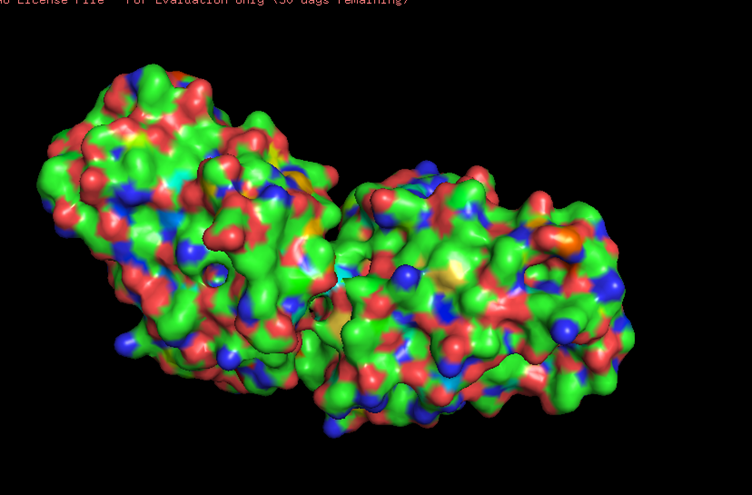
 Looking at the surface structure, there seems to be small holes in the middle of the protein,no major holes. The depression in the middle seems to be a good binding site.
Looking at the surface structure, there seems to be small holes in the middle of the protein,no major holes. The depression in the middle seems to be a good binding site.
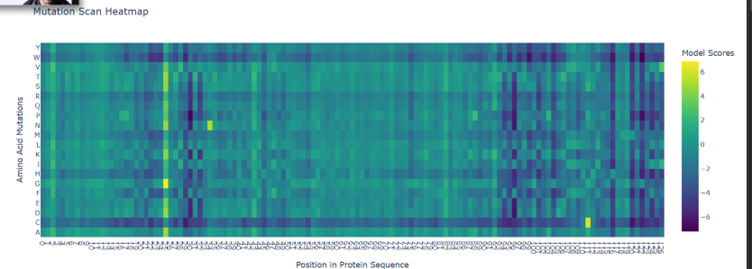

Here the green spots are regions that tolerate mutations pretty well. Ie. If they change doesn’t really affect the functionality much.
Purple are critical regions if there is a mutation here, there is a massive change in the overall structure and functionality.
Yellow regions are pretty sparse here, these regions are good to mutate
There seems to be good mutation candidates for position 25 (Glutamine). At the ends of the protein is where mutations seems to be really bad.
b. Looking at the 3d PCA plot there seems to be no distinct formation of clusters or neighbourhood. All seem to be similar proteins
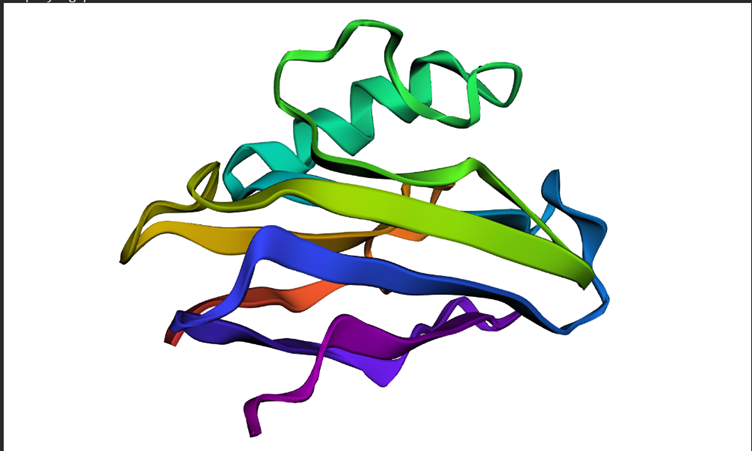
 Overall the structure seems to be similar to the first but looking in depth there seems to be some variations between the predictions and the original structure
Overall the structure seems to be similar to the first but looking in depth there seems to be some variations between the predictions and the original structure
Smaller mutations did not create much difference but the larger mutations did. For example I replicated a chunk of the protein within the protein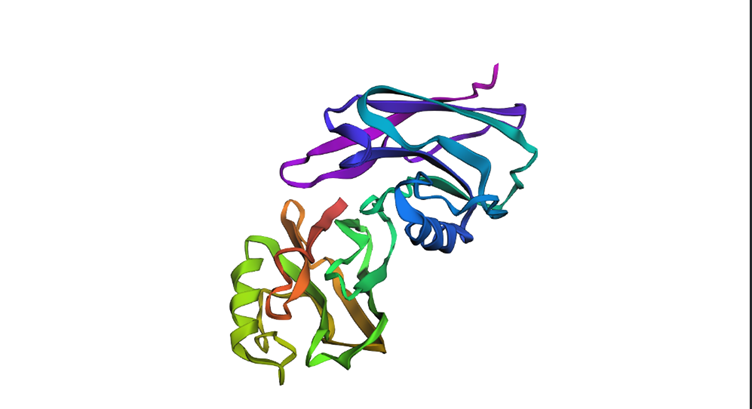

Before mutation
sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2
MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
After A4V mutation
MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
I used the pepMLM colab to generate 4 binders
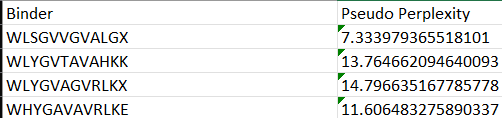

Each of the above binders are number 0 to 3 and the binder_ex is the known SOD1-binding peptide FLYRWLPSRRGG
Most of them appear to be binding near the B barrel structure
Binder_3 had the highest ipTM score this also had the highest perplexity score. This also exceeds the known binder. This measure means that this interaction/structure is more stable and probable than other structures.
I would choose binder_ex i.e FLYRWLPSRRGG because it causes no hemolysis, it is soluble and permeable and although it has a weak binding affinity (most others have it too), it had the 2nd highest iPTM score making it probable structure. This is the best candidate. All other pepetides cause non permeability which makes them a bad candidate for therapy.
HTGAA Committed Listener (CL) Agreement I am a HTGAA Committed Listener, my responsibilities are:
Watching class lectures and recitations Participating in node reviews Developing and documenting my homework Actively communicating with other students and TAs on the forum Allowing HTGAA and BioClub to share my work (with attribution) Honestly reporting on my work, and appropriately attributing and citing the work of others (both human and non-human) Following locally applicable health and safety guidance Promoting a respectful environment free of harassment and discrimination Signed by committing this file to my documentation page/repository,
{{ Shishir S Nair }}
{{ 08-03-2026}}