Week 06 HW: Genetic Circuits Part I
DNA Assembly
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
- Phusion DNA Polymerase: synthesizes new DNA strands by adding nucleotides to the template strand during PCR
- dNTPs (deoxynucleotides): building blocks of DNA
- Buffer: provides the optimal conditions
2. What are some factors that determine primer annealing temperature during PCR?
- Tm (melting temperature) of the primer: the temperature at which half of the DNA duplex dissociates
- Primer length: longer primers generally require higher annealing temperatures
- GC content: higher GC content increases the Tm and may require higher annealing temperatures
- Salt concentration: higher salt concentrations can stabilize the DNA duplex and may require higher annealing temperatures
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR:
- Protocol: denaturation, annealing, and extension steps to amplify specific DNA sequences using primers and a DNA polymerase
- Advantages: can amplify specific DNA sequences from complex mixtures, does not require specific restriction sites, can introduce mutations or tags through primer design
- Disadvantages: may produce non-specific products, requires optimization of conditions, can be time-consuming
Restriction enzyme digests:
- Protocol: cutting DNA at specific sites using restriction enzymes
- Advantages: can generate specific fragments based on known restriction sites
- Disadvantages: requires specific restriction sites
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
Create overlapping sequences at the ends of the DNA fragments that are complementary to each other, which allows for efficient assembly during Gibson cloning.
5. How does the plasmid DNA enter the E. coli cells during transformation?
- Heat shock: Generate pores in bacterial cell wall with an abrupt temperature change
- Electroporation: Generate pores in bacterial cell wall with high electrical voltage
6. Describe another assembly method in detail (such as Golden Gate Assembly)
Golden Gate Assembly: assembly of multiple DNA fragments in a single reaction using type IIS restriction enzymes and DNA ligase. The type IIS restriction enzymes cut the recognition site and creates a overhang that is designed to be complementary to the overhang of the fragment to be inserted, allowing for seamless assembly. This method is efficient and allows for the assembly of multiple fragments in a single reaction.
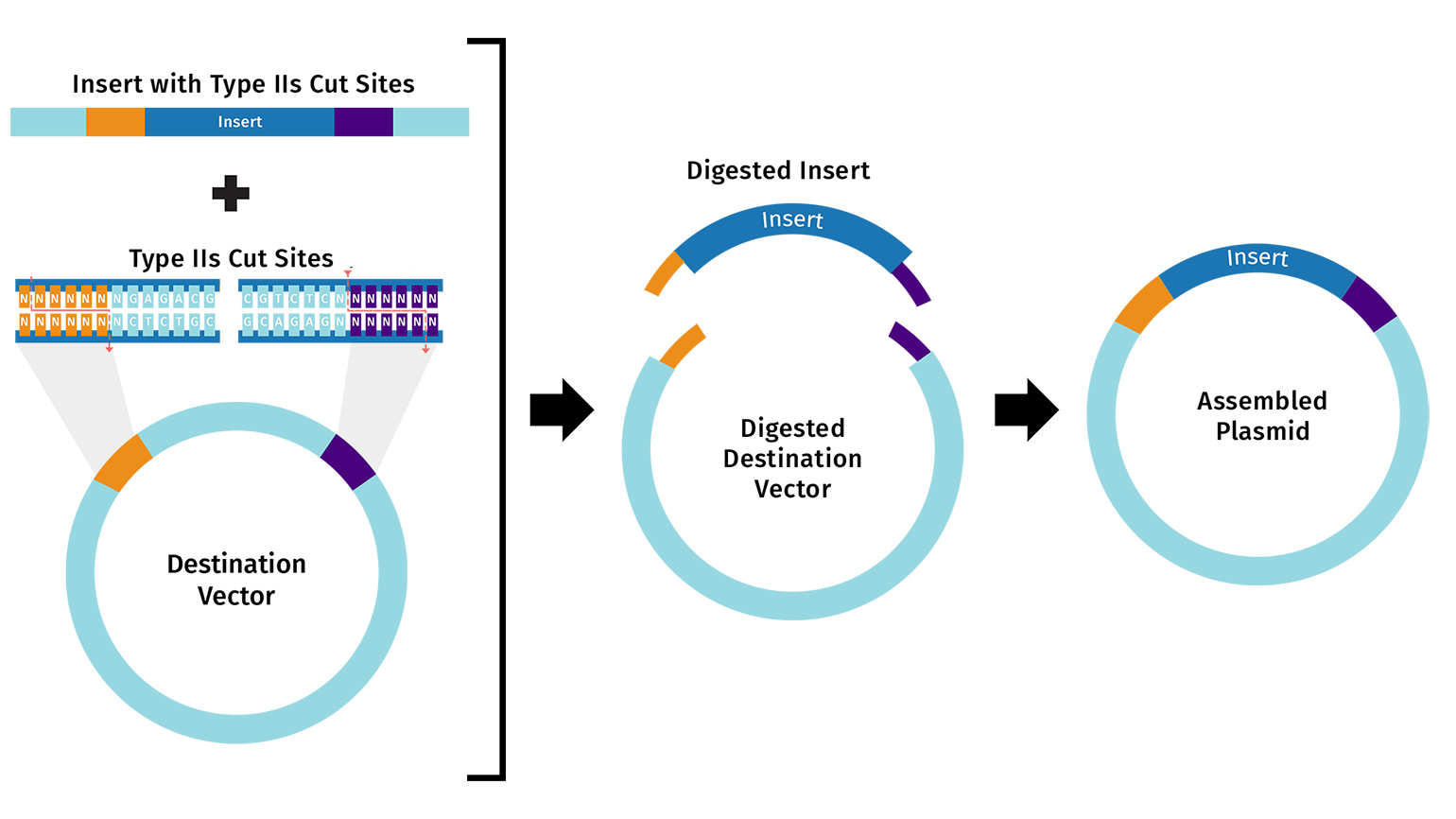
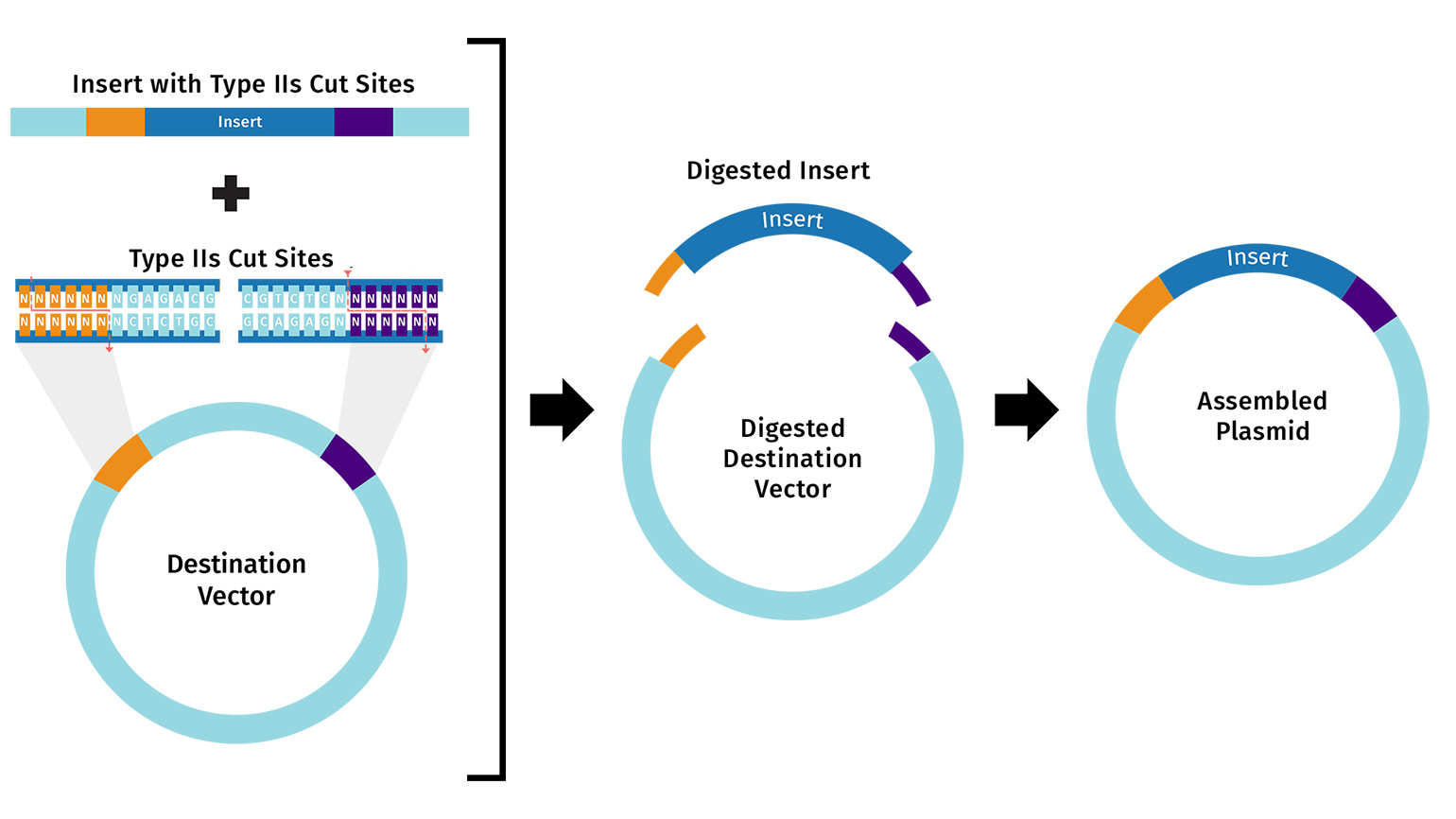
Asimov Kernel
Recreate Repressilator
I recreated the Repressilator by dragging the parts one by one from the Characterized Bacterial Parts repository to the editor.
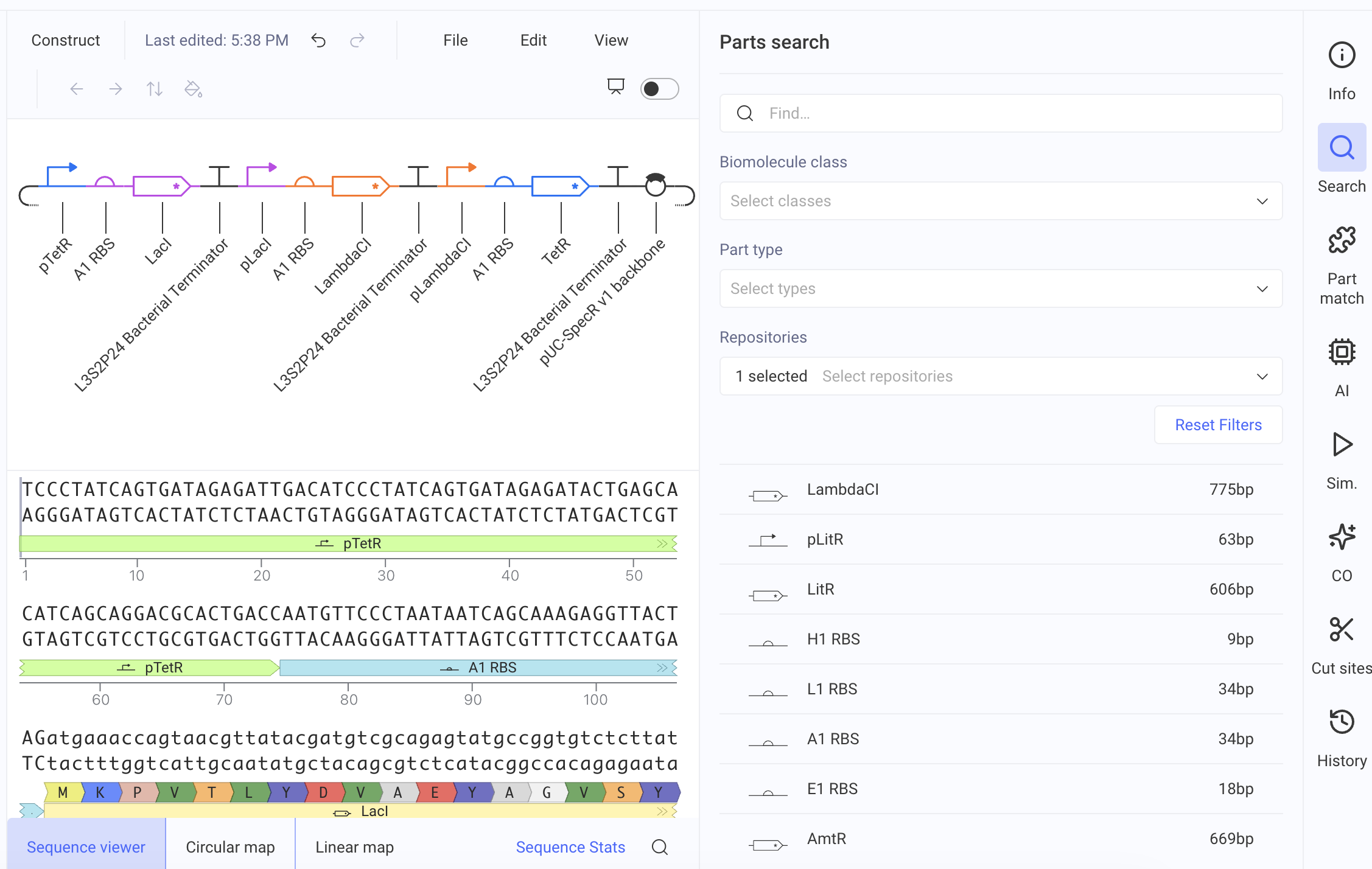

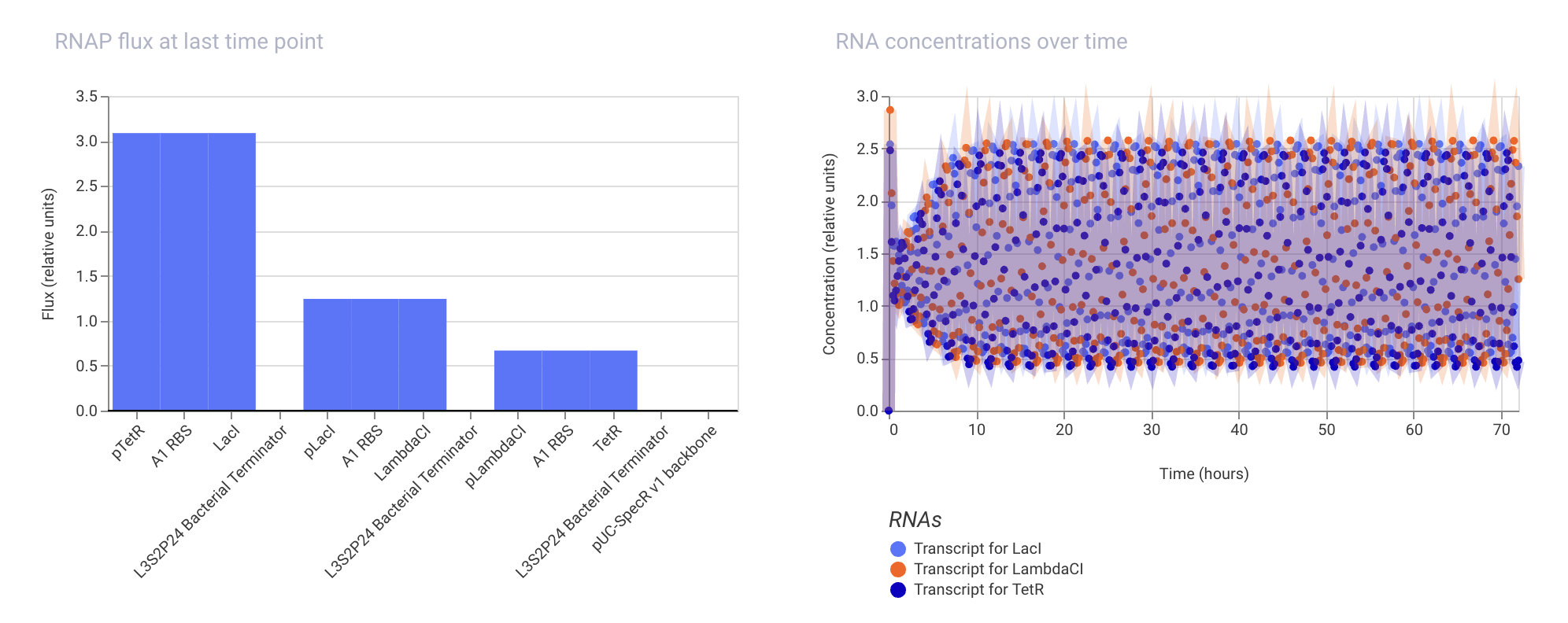
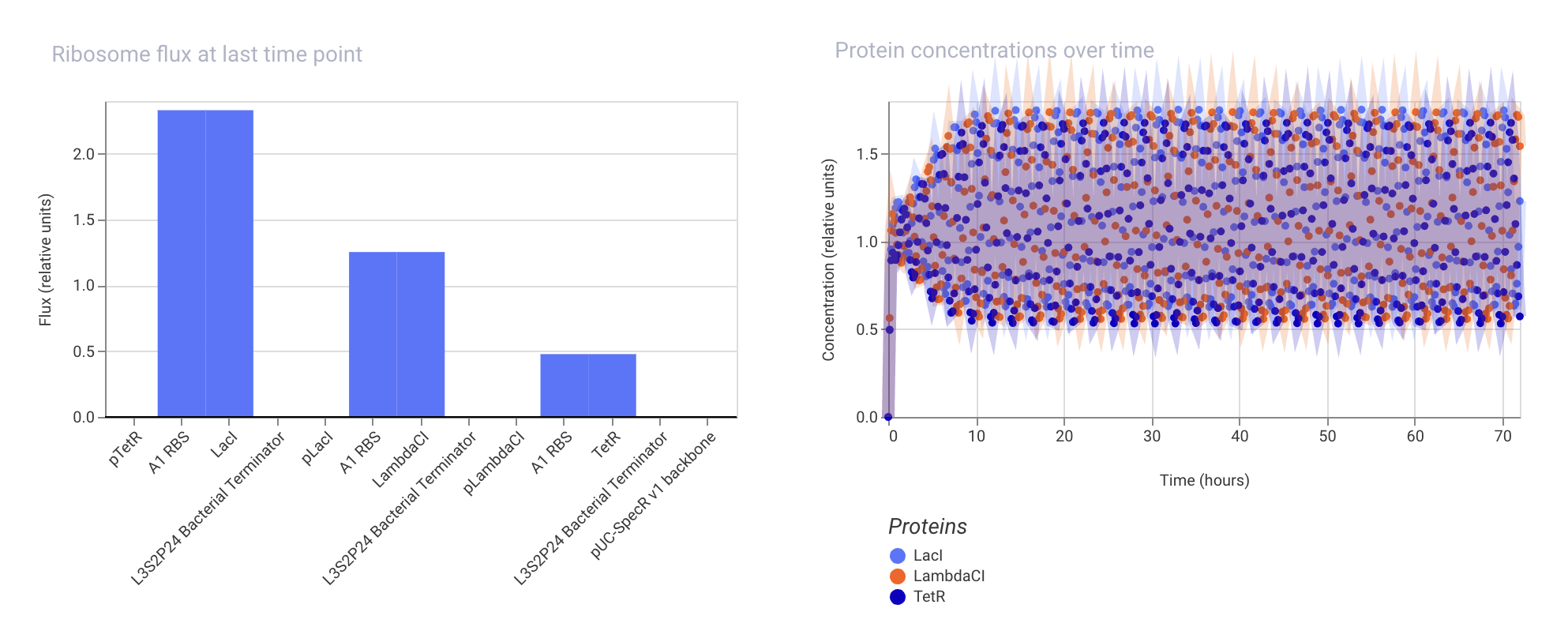
Below are the simulation results:
The results look similar to the original Repressilator construct simulation results, which shows oscillations in the expression levels of the three proteins.
My Construct
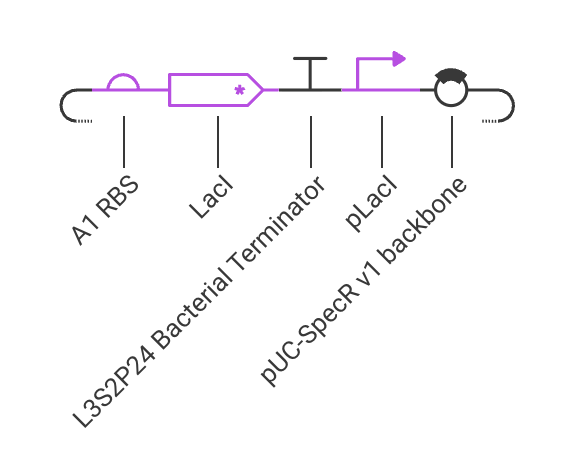
I created a construct consists of pLacI promoter, RBS, LacI coding sequence, and terminator. Below are the simulation results, which show that the LacI protein is expressed at a high level:
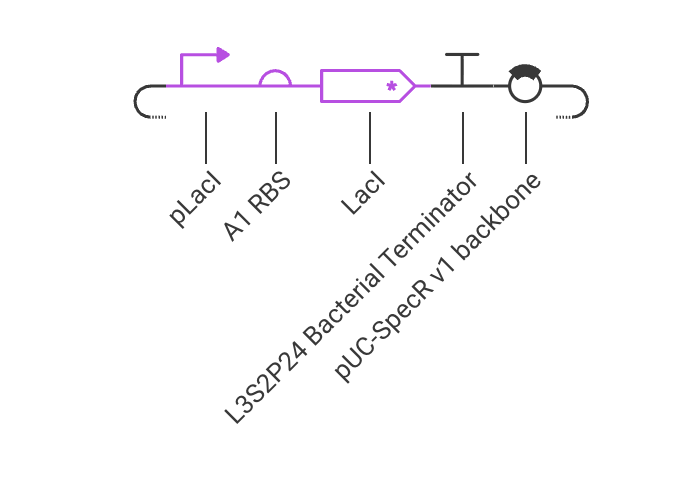

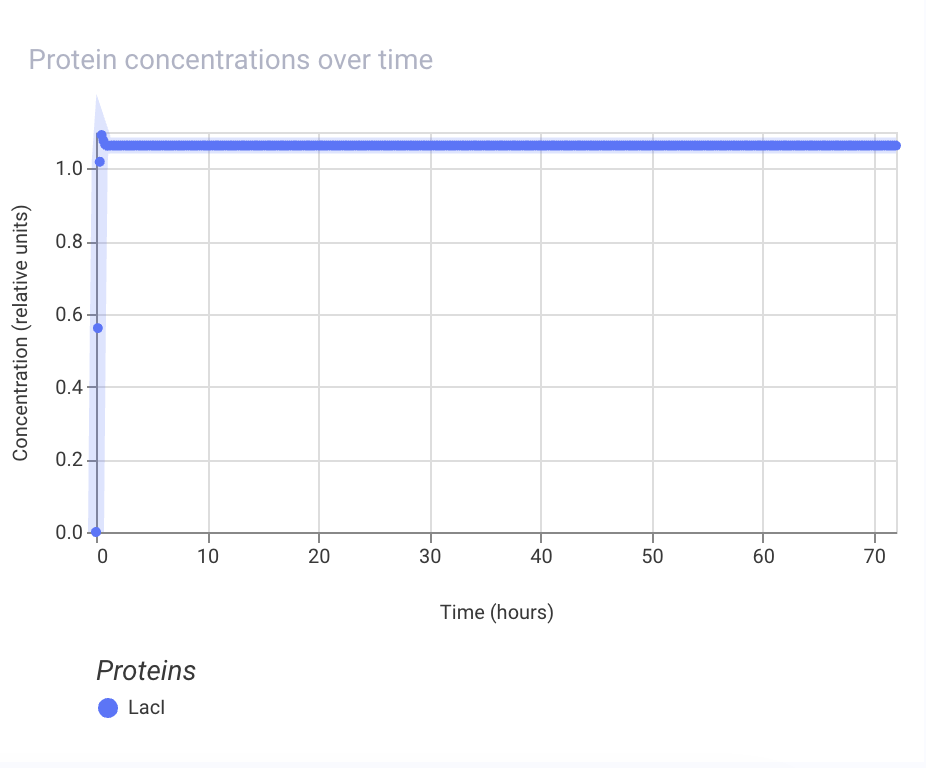

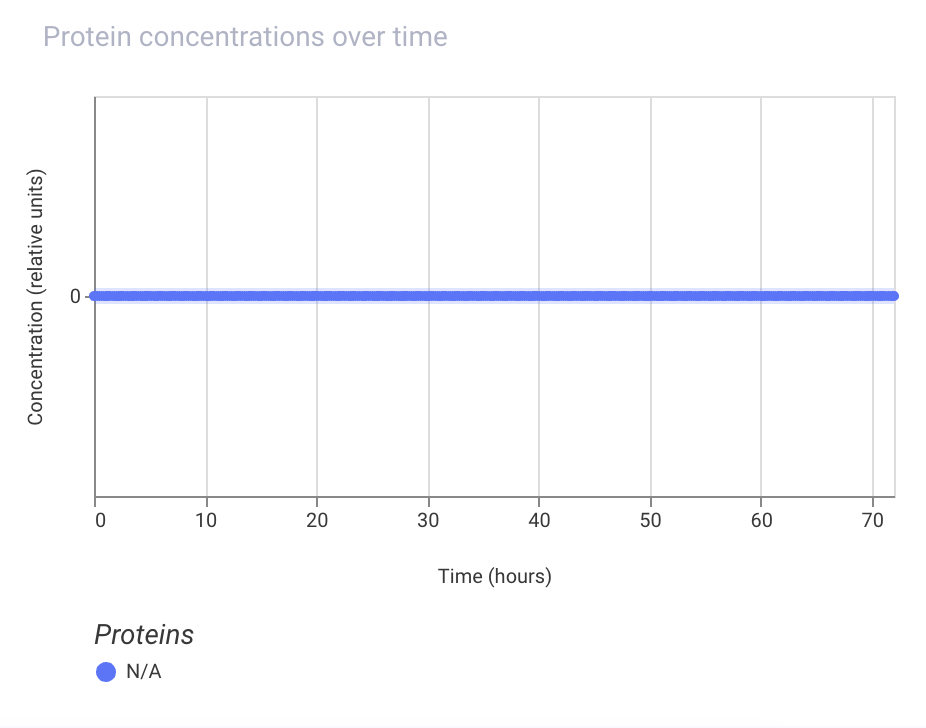
Next, I tried to remove the promoter, and the simulation results show that the LacI protein is not expressed at all:

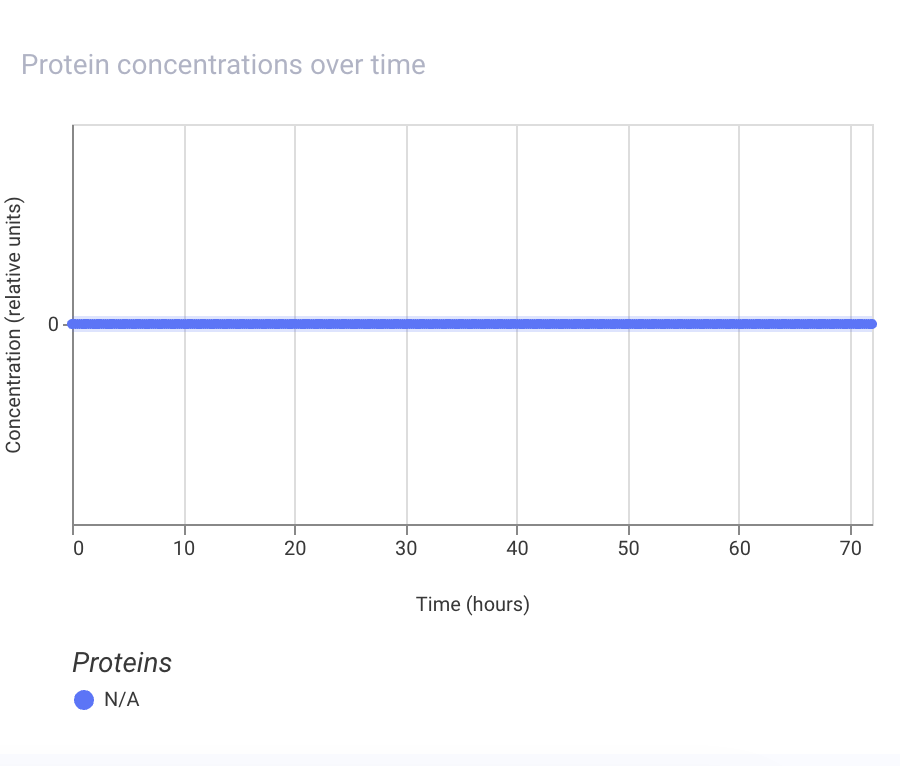

I also tried to put the promoter after the LacI coding sequence, and the simulation results show that the LacI protein is not expressed at all: