Week 2 HW: DNA Read, Write and Edit
Part 0
Done.
Part 1
Since it’s passe to create “MIT” with the electrophoresis gel, I decided to reverse the order of the letters to spell “TIM” instead.
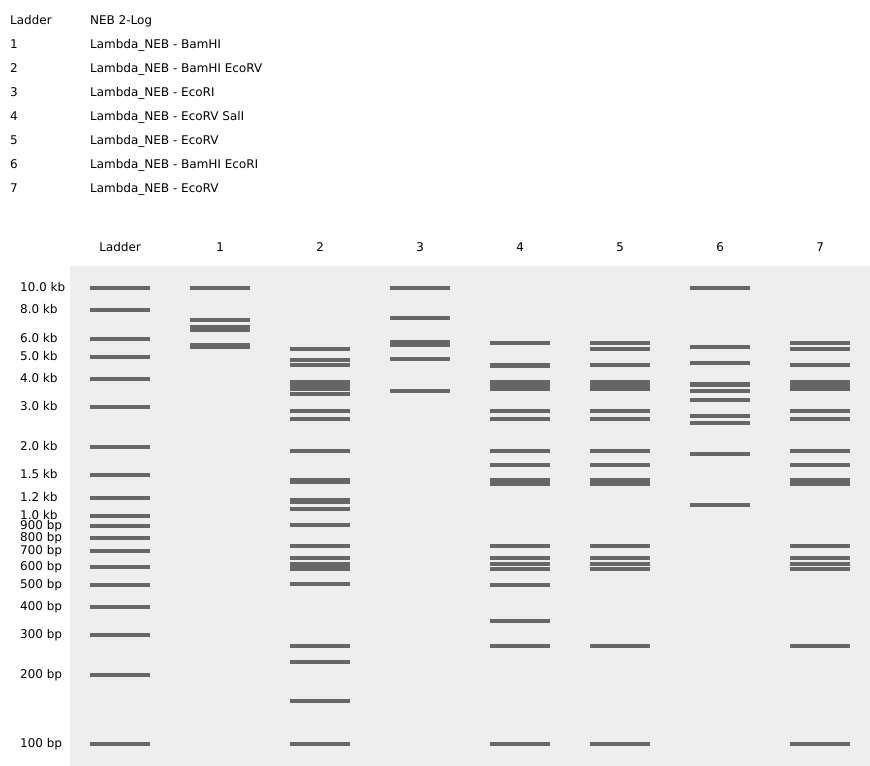


Part 2
I don’t have in-person access to a node, so can’t perform the wet lab component.
Part 3
Part 3.1
I chose to investigate an inorganic pyrophosphatase-driven proton pump because it’s the most relevant to the SoilBuddy MVP. Here are other candidates I considered and why I rejected them:
- Bacteriorhodopsin: +Prokaryote-native -Light-driven rxn is impractical for a soil-dwelling bacterium
- Membrane H+-ATPase: +Native cellular localization at membrane is favorable -High risk of cross-talk with native membrane proton pumps and may preclude bacterial survival
- Methanogenic proton pump: - Presupposing an electron acceptor for the correct functioning of the proton pump limits our choices of target organism and may hinder the systems integration of other effector means for SoilBuddy
The sequence is as follows, obtained from UniProt
Part 3.2
The reverse translated nucleotide sequence is as follows, obtained using bioinformatics.org’s reverse translation tool
Part 3.3
We need to optimize codons because different model organisms have aminoacyl-tRNAs in different abundances. Certain organisms may not have enough, or any, tRNAs with the anticodons corresponding to our nucleotide sequence, while for others, certain tRNAs may be more abundant, and thus facilitate more efficient translation of mature mRNA.
I’ll be optimizing my sequence for E. coli since it’s both a well-characterized transformation and plasmid amplification platform, and amenable to SoilBuddy’s target application. Given the length of the target sequence, I’ll hedge my bets on Golden Gate Assembly, and optimize my codons without the restriction sites for common type IIS enzymes such as BsaI, BbsI, BsmBI, and FokI. I used Twist Bioscience’s codon optimization tool:
Part 3.4
Once the target sequence is spliced into a cloning vector via golden gate assembly and a competent strain of E. coli is obtained, the bacteria could be transformed using heat shock treatment. After selection for expression of the cloning vector, transformed bacteria would be incubated in broth.
Cells would be extracted via centrifugation and lysis with detergent to release the membrane protein. The membrane protein would be purified using chromatography and verified using Western blot.
A cell-free protocol would use E. coli lysate to conduct the in vitro transcription then translation of the plasmid vector containing the target sequence. Purification and verification of the protein would similarly involve chromatography and Western blot.
Part 3.5
Upon transcription of a DNA sequence to pre-mRNA, post-transcriptional modification of the pre-mRNA involves alternative splicing, which allows for different combinations of exons to be present in the mature mRNA, and subsequently translated into protein isoforms.
Alternatively, transcription initiation complexes may form on different promoters within the gene, causing different transcripts to begin with.
Part 4
Part 4.1
Done
Part 4.2
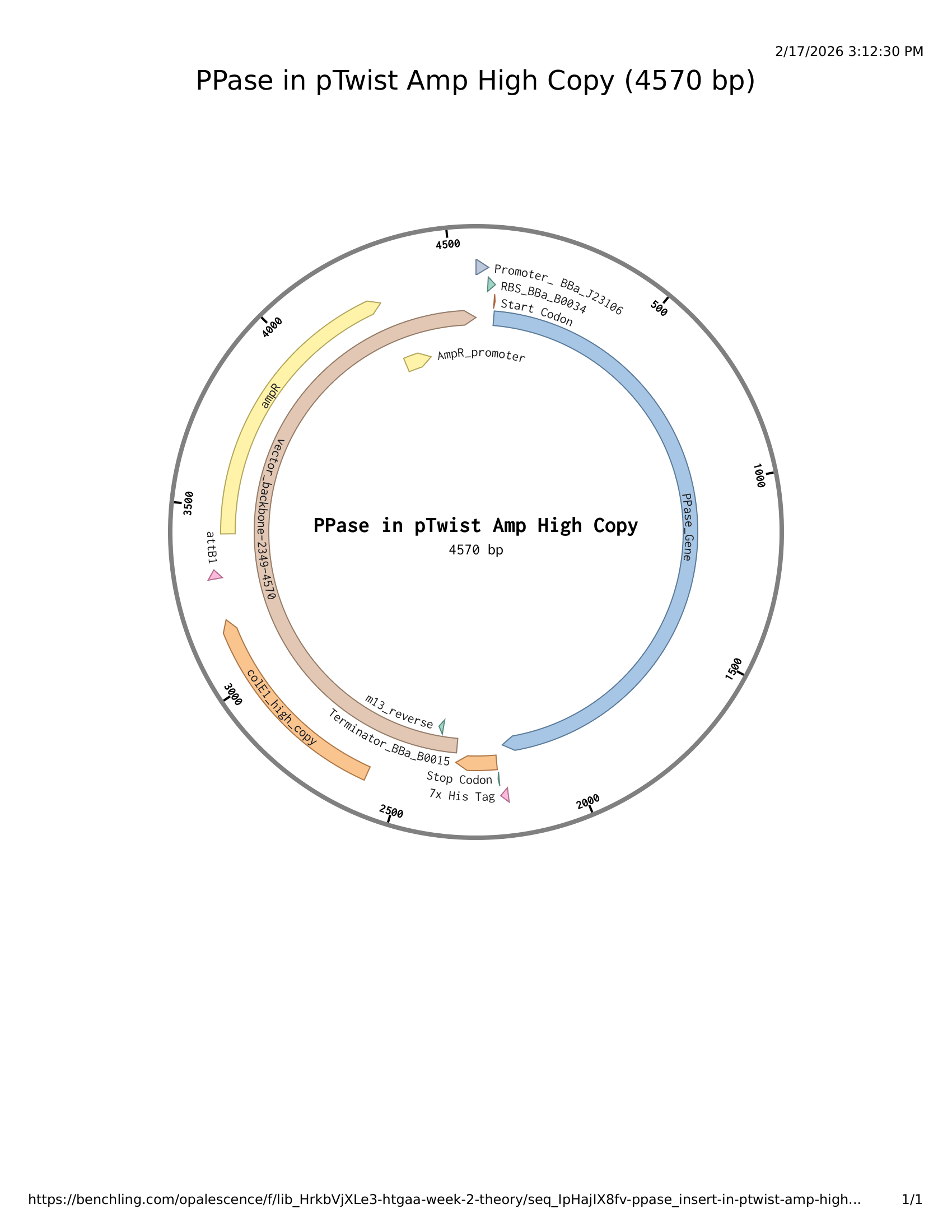

Part 5
Part 5.1: DNA Read
(i) I’d like to sequence the eDNA of the soil microbiome to gain an understanding of the organisms SoilBuddy would have to co-exist with.
(ii)
I’d use Illumina NGS sequencing because it balances read-length - more than sufficient for analyzing eDNA fragments - with cost and thoroughput. It is classified as a 2nd-generation sequencing technology since it utilizes sequencing-by-synthesis, albeit in a massively parallel architecture.
My input is purified DNA fragments obtained from soil solution. Depending on the distribution of fragment sizes, a coarse reading of which may be obtained with a pilot gel electrophoresis using a standard ladder, I will first use enzymatic fragmentation to produce suitable DNA fragments, then ligate Illumina NGS-specific adapters containing barcode sequences, and finally run a low number of PCR cycles to amplify my DNA fragments.
After PCR amplification, I’ll pool my samples to ensure equal representation in my library and verify their purity before loading them into the sequencer.
- Under Illumina NGS, the fragments of DNA in my library hybridize sparsely over the surface of the sequencer’s flow cell, thanks to the adapters ligated to the eDNA samples. The fragments first undergo bridge amplification PCR wherein multiple copies of the fragments (both forward and reverse strands corresponding to the fragment sequence) are generated in small clusters within the flow cell’s nanowells. Reverse strands are enzymatically cleaved and washed off the flow cell.
Once clusters are produced, the cell is flooded with a primer, DNA polymerase and modified fluorescently-tagged nucleotides which can only be incorporated by the DNA polymerase one base at a time to the oligomers surrounding the clusters. Unbound nucleotides are washed away, then a picture of the flow cell is taken, which reveals the specific base incorporated in each cluster by virtue of the unique wavelength of fluorescent light given off by each modified nucleotide (ie. A, T, C or G). The fluorescent tag blocking DNA synthesis is chemically inactivated, then the chip is again flooded with fluorescently-tagged nucleotides. This process repeats till the full length of DNA fragments has been replicated.
Specifically, Illumina NGS identifies the specific bases using the unique fluorescent signature emitted by the fluorescent tag attached to each of the four possible nucleotides.
- After demultiplexing, the output is short read data. I’d subsequently process the raw read data using a metagenomic analysis pipeline to identify specific taxa and species present in the original soil solution sample.
Part 5.2: DNA Write
(i) I’d like to synthesize DNA for a genetic circuit that regulates bacterial membrane protein pumps depending on external pH as part of my SoilBuddy MVP. I’d choose a set point optimized for food crops such as corn.
(ii)
- I’d use Gibson Assembly to put my genetic circuit together given the simplicity and reliability of the protocol. First, I’d identify appropriate restriction enzyme combinations for the BioBricks parts I’ll require. Next, I’ll pick two BioBricks parts at a time for insertion into the target plasmid.
Each BioBricks part and the vector will be digested in respective reaction chambers with the appropriate restriction enzymes, followed by thermal denaturation (and thus inactivation) of the restriction enzymes. Thereafter, the digested BioBricks parts and vector will be incubated together in molar amounts designed to optimize formation of the target construct along with a DNA ligase. After transformation into a bacterial platform, selection, amplification and extraction, repeat the process till all the BioBricks parts are incorporated within the recombinant plasmid vector.
- Needless to say, the process is extremely time-intensive due to the multiple, repetitive transformations and amplifications involved. While laboratory automation could allow the process to scale, it would be very resource-intensive. Lastly, while each particular step might be lossy, regular purification, selection and verification steps would ensure the desired vector is formed at the end of the day.
Part 5.3: DNA Edit
(i) One possible development of SoilBuddy would be a process rather than an organism that transforms native soil microbiota to enhance their salutary functions (eg nitrogen-fixing, pH buffering). To that end, I would like to edit prokaryotic bacteria and eukaryotic fungal DNA considering the particular niche of commensal microbiota near plant roots. This would involve, mainly, gene insertion to introduce novel gene products or base edits to enhance existing regulatory mechanisms in the microorganisms.
(ii)
- Given the range of target systems (both prokaryotic and eukaryotic) as well as payloads desired (ie. novel genes and edits), I would leverage the versatility of CRISPR. Taking a CRISPR-based gene knock-in of yeast as an example, we first design a DNA sequence encoding a gRNA that recognizes our target insertion site and has minimal off-target effects. Then, we select a yeast plasmid with an antibiotic selection marker and which contains a sequence encoding Cas9 endonuclease. Using Gibson assembly, we first digest the vector and gRNA-encoding sequence with appropriate restriction enzymes, then ligate the sequence into the plasmid using DNA ligase, transform the recombinant vector into competent yeast cells, select and amplify the plasmid through incubation.
We prepare a donor template containing the gene to be inserted and transform the exogenous donor template into yeast. Now, the yeast contains Cas9 endonuclease from expression of the plasmid, which hybridizes with gRNA (also expressed from transcrption of the plasmid DNA) to form ribonucleoprotein complex. The crRNA in the gRNA binds to the recognition site before Cas9 domain activity causes a double-stranded break in the yeast DNA. Thereafter, the donor template provides for Homology-Directed Repair that knocks in the intended gene.
The preperatory work involves designing the gRNA, selecting yeast cells that are competent and do not suffer from damage to their Homology-Directed Repair mechanism, and selecting a donor template incorporating a reporter (eg GFP) such that unintended edit products are minimized. At the same time, edited yeast cells have to be selected due to the low efficiency of gene knock-in through fluorescence-based flow-cytometry for example.
While a well-designed gRNA and donor template will minimize indels, off-target edits and maximize the chances of successful gene knock-in, statistically speaking, the odds of a cell being successfully edited are low. Hence, care needs to be taken in selecting for edited cells, which can then be expanded to counteract the low efficiency of knock-in