Week 2 HW: DNA Read, Write and Edit
Part 1: Benchling and In-Silico Gel Art
I made a Benchling accountand imported the Lambda DNA genbank sequence from https://www.ncbi.nlm.nih.gov/nuccore/J02459.1?report=genbank&log$=seqview
After importing everything I used Benchling’s Digest tool to stimulate the enzyme digestion of the Lambda DNA using the required enzymes. I first made all of the enzymes that were asked in the list.
EcoRI
HindIII
BamHI
KpnI
EcoRV
SacI
SalI
EcoRV made a lot of small fragments, which made very dense lower bands, BamHI produced less mid-size fragments.
At first I was experimenting with the fragments to make a “:D” smiley face. But later, while working through it I realized that I had created more of a wine glass / goblet looking shape and, hence, I pivoted to making just that.
For the final composition, I arranged the following enzyme digests across lanes:
Lane 1: BamHI Lane 2: EcoRI Lane 3: HindIII Lane 4: EcoRI Lane 5: BamHI
The symmetric outer lanes create the bowl width, the EcoRI lanes taper inward, and the HindIII digest forms the vertical stem due to its smaller fragment sizes. When cropped and viewed holistically, the gel bands form the silhouette of a wine glass.


Part 3: DNA Design Challenge
I picked Ovalbumin as my protein. The reason I picked this protein is becuase it is the main protein that is used in egg whites, and it’s historically important in immunology. (and i like eggs)
Protein Sequence:
tr|A0A411G5W6|A0A411G5W6_CHICK Ovalbumin OS=Gallus gallus OX=9031 PE=2 SV=1 MGSIGAASMEFCFDVFKELKVHHANENIFYCPIAIMSALAMVYLGAKDSTRTQINKVVRF DKLPGFGDSIEAQCGTSVNVHSSLRDILNQITKPNDVYSFSLASRLYAEERYPILPEYLQ CVKELYRGGLEPINFQTAADQARELINSWVESQTNGIIRNVLQPSSVDSQTAMVLVNAIV FKGLWEKTFKDEDTQAMPFRVTEQESKPVQMMYQIGLFRVASMASEKMKILELPFASGTM SMLVLLPDEVSGLEQLESIINFEKLTEWTSSNVMEERKIKVYLPRMKMEEKYNLTSVLMA MGITDVFSSSANLSGISSAESLKISQAVHAAHAEINEAGREVVGSAEAGVDAASVSEEFR ADHPFLFCIKHIATNAVLFFGRCVSP
Reverse Translation: (by bioinformatics.org)
reverse translation of tr|A0A411G5W6|A0A411G5W6_CHICK Ovalbumin OS=Gallus gallus OX=9031 PE=2 SV=1 to a 1158 base sequence of most likely codons. atgggcagcattggcgcggcgagcatggaattttgctttgatgtgtttaaagaactgaaa gtgcatcatgcgaacgaaaacattttttattgcccgattgcgattatgagcgcgctggcg atggtgtatctgggcgcgaaagatagcacccgcacccagattaacaaagtggtgcgcttt gataaactgccgggctttggcgatagcattgaagcgcagtgcggcaccagcgtgaacgtg catagcagcctgcgcgatattctgaaccagattaccaaaccgaacgatgtgtatagcttt agcctggcgagccgcctgtatgcggaagaacgctatccgattctgccggaatatctgcag tgcgtgaaagaactgtatcgcggcggcctggaaccgattaactttcagaccgcggcggat caggcgcgcgaactgattaacagctgggtggaaagccagaccaacggcattattcgcaac gtgctgcagccgagcagcgtggatagccagaccgcgatggtgctggtgaacgcgattgtg tttaaaggcctgtgggaaaaaacctttaaagatgaagatacccaggcgatgccgtttcgc gtgaccgaacaggaaagcaaaccggtgcagatgatgtatcagattggcctgtttcgcgtg gcgagcatggcgagcgaaaaaatgaaaattctggaactgccgtttgcgagcggcaccatg agcatgctggtgctgctgccggatgaagtgagcggcctggaacagctggaaagcattatt aactttgaaaaactgaccgaatggaccagcagcaacgtgatggaagaacgcaaaattaaa gtgtatctgccgcgcatgaaaatggaagaaaaatataacctgaccagcgtgctgatggcg atgggcattaccgatgtgtttagcagcagcgcgaacctgagcggcattagcagcgcggaa agcctgaaaattagccaggcggtgcatgcggcgcatgcggaaattaacgaagcgggccgc gaagtggtgggcagcgcggaagcgggcgtggatgcggcgagcgtgagcgaagaatttcgc gcggatcatccgtttctgttttgcattaaacatattgcgaccaacgcggtgctgtttttt ggccgctgcgtgagcccg
I codon optimized this using Twist Biosciences
Ovalbumin_Ecoli_Optimized_Insert_1158bp ATGGGCAGCATTGGCGCGGCGAGCATGGAATTTTGCTTTGATGTGTTTAAAGAACTGAAAGTGCATCATGCGAACGAAAACATTTTTTATTGCCCGATTGCGATTATGAGCGCGCTGGCGATGGTGTATCTGGGCGCGAAAGATAGCACCCGCACCCAGATTAACAAAGTGGTGCGCTTTGATAAACTGCCGGGCTTTGGCGATAGCATTGAAGCGCAGTGCGGCACCAGCGTGAACGTGCATAGCAGCCTGCGCGATATTCTGAACCAGATTACCAAACCGAACGATGTGTATAGCTTTAGCCTGGCGAGCCGCCTGTATGCGGAAGAACGCTATCCGATTCTGCCGGAATATCTGCAGTGCGTGAAAGAACTGTATCGCGGCGGCCTGGAACCGATTAACTTTCAGACCGCGGCGGATCAGGCGCGCGAACTGATTAACAGCTGGGTGGAAAGCCAGACCAACGGCATTATTCGCAACGTGCTGCAGCCGAGCAGCGTGGATAGCCAGACCGCGATGGTGCTGGTGAACGCGATTGTGTTTAAAGGCCTGTGGGAAAAAACCTTTAAAGATGAAGATACCCAGGCGATGCCGTTTCGCGTGACCGAACAGGAAAGCAAACCGGTGCAGATGATGTATCAGATTGGCCTGTTTCGCGTGGCGAGCATGGCGAGCGAAAAAATGAAAATTCTGGAACTGCCGTTTGCGAGCGGCACCATGAGCATGCTGGTGCTGCTGCCGGATGAAGTGAGCGGCCTGGAACAGCTGGAAAGCATTATTAACTTTGAAAAACTGACCGAA TGGACCAGCAGCAACGTGATGGAAGAACGCAAAATTAAAGTGTATCTGCCGCGCATGAAAATGGAAGAAAAATATAACCTGACCAGCGTGCTGATGGCGATGGGCATTACCGATGTGTTTAGCAGCAGCGCGAACCTGAGCGGCATTAGCAGCGCGGAAAGCCTGAAAATTAGCCAGGCGGTGCATGCGGCGCATGCGGAAATTAACGAAGCGGGCCGCGAAGTGG TGGGCAGCGCGGAAGCGGGCGTGGATGCGGCGAGCGTGAGCGAAGAATTTGCGCGGATCATCCGTTTCTGTTTTGCATTAAACATATTGCGACCAACGCGGTGCTGTTTTTTGGCCGCTGCGTGAGCCCG
The optimized ovalbumin gene was cloned into a pET plasmid for expression in E. coli. After transformation into bacterial cells, the T7 promoter drives transcription of the DNA into mRNA. Ribosomes translate the mRNA into ovalbumin protein. The protein can then be purified from bacterial lysate. This process follows the central dogma: DNA is transcribed into RNA, which is translated into protein.
Part 4: Prepare a DNA Synthesis Order
4.2
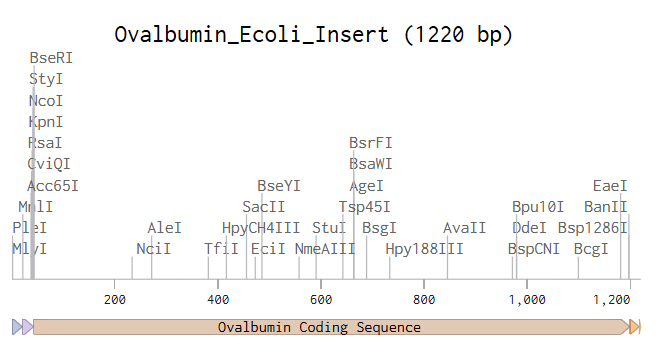

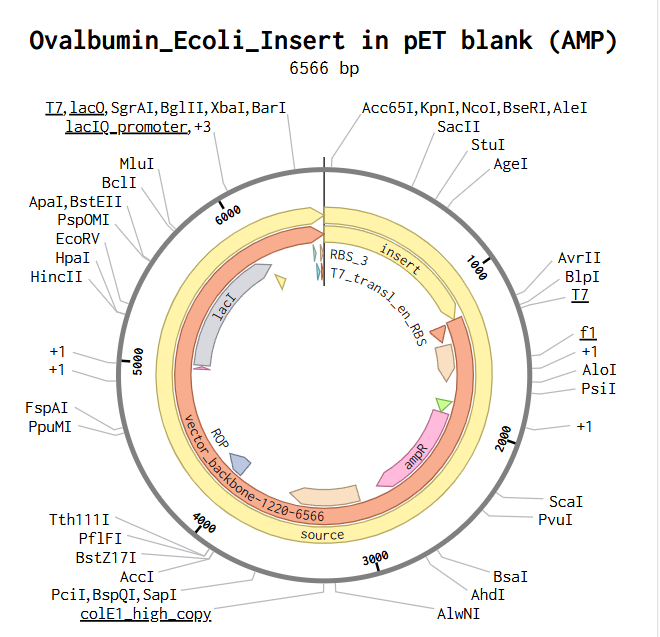
For plasmid assembly, I selected the pTwist Amp High Copy vector, which is designed for high-copy replication in E. coli and contains an ampicillin resistance marker for selection. This backbone supports efficient bacterial transformation and protein expression. The codon-optimized ovalbumin expression cassette was inserted into this circular backbone, resulting in a 6504 bp plasmid construct.

Part 5: DNA Read
(i) What DNA would you like to sequence and why?
I’d like to sequence synthetics DNA used for digital data storage. I remember we briefly went over this topic in the last lecture, but I found that it was very interesting that data could be stored in such a way. DNA-based storage encodes binary information into nucleotide sequences. This sequencing can allow the retrieval of the stored data by reading the encoded bases and reconstructing the digital information. I’d like to do this specifically because feel that it bridges biology and computation, and it demonstrates how DNA can be a high-density storage medium beyond its biological role.
(ii) What sequencing technology would you use?
Illumina is a second generation sequencing technology that offers veru high accuracy and massive parallel sequencing, which means it is well suited for decoding short synthetic DNA fragments used for digital storage systems.
The output consists of millions of short DNA reads in FASTQ format, containing both sequence information and quality scores. These reads can be computationally assembled to reconstruct the original stored data.
5.2 DNA Write
(i) What DNA would you want to synthesize and why?
Long-term archival data storage DNA strands. Digital files to binary code to nucleotide sequencers through mapping schemes. This offers very high data density, and long-term stability compared to traditional hard drives.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
DNA Phosoramidite Synthesis. Nucleotides are added sequentially to a growing DNA strand attached to a solid support.
5.3 DNA Edit
(i) What DNA would you want to edit and why?
TERT gene in human somatic cells plays a role in cellular aging. Telomeres shorten during cell division. Understanding and modulating TERT expression can provide insights into aging, longevity and age-related diseases.
(ii) What technology would you use?
CRISPR-based base editing to modify the TERT gene (telomerase reverse transcriptase) in human somatic cells. Telomeres shorten with age, contributing to cellular senescence, and carefully regulating telomerase activity may help improve regenerative capacity. Base editing allows precise single-nucleotide changes without creating double-strand breaks, reducing the risk of large mutations. A Cas9 nickase fused to a deaminase enzyme, guided by RNA, converts specific bases within a targeted region. However, limitations include off-target edits and the risk of increased cancer if telomerase activity becomes uncontrolled.