Week 2 HW: DNA read, write and edit
Homework
Part 1: Benchling and in-silico gel art
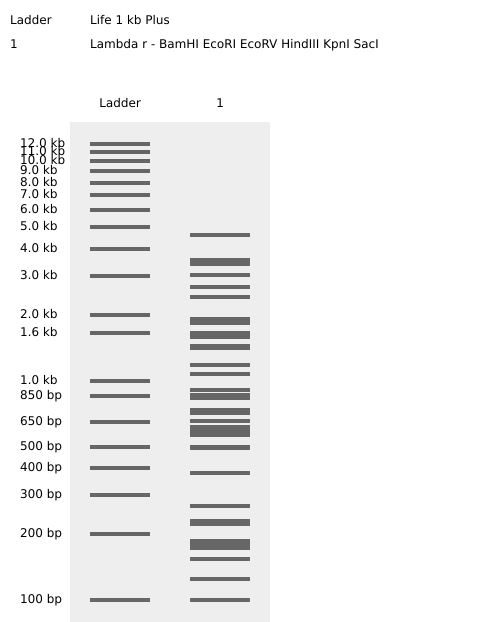

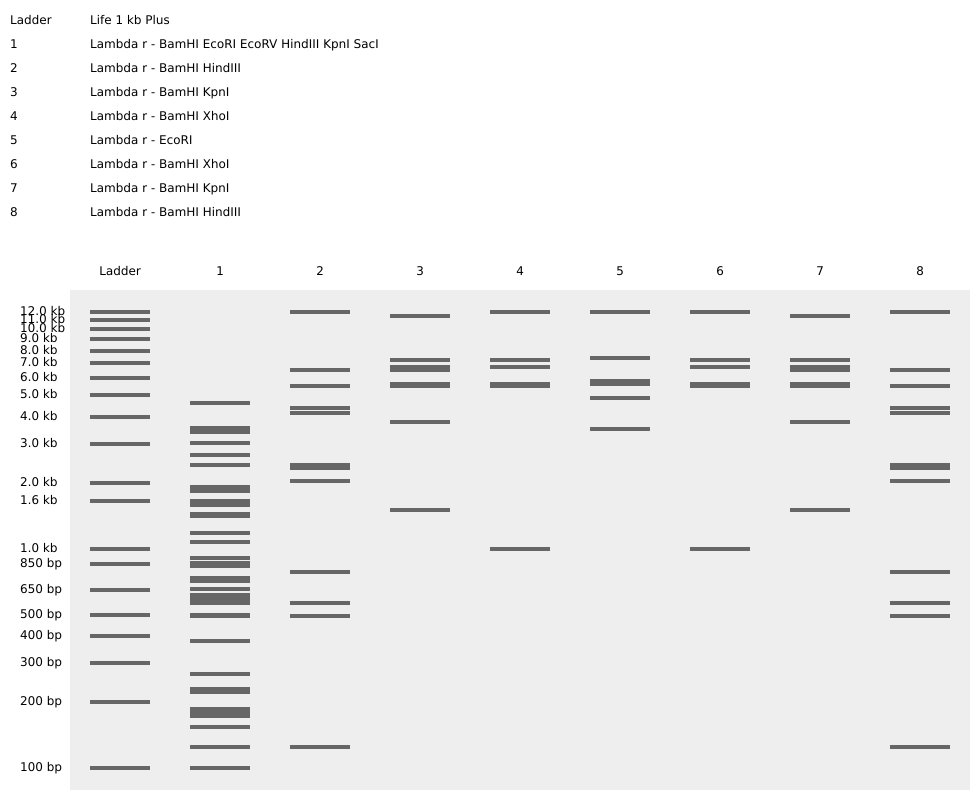
This is the virtual digestion of the Lambda genome with this restriction enzymes: EcoRI, HindIII, BamHI, KpnI, EcoRV, SacI and SalI.
And here is some artwork:

Part 3: DNA design challenge
3.1- I have choose for the DNA nucleotidyllexotransferase, because it is used in the enzymatic DNA synthesis process, and been able to engineer it could conduct to synthesize DNA in vivo, thing that would make advance the field tremendously.
sp|P04053|TDT_HUMAN DNA nucleotidylexotransferase OS=Homo sapiens OX=9606 GN=DNTT PE=1 SV=3 MDPPRASHLSPRKKRPRQTGALMASSPQDIKFQDLVVFILEKKMGTTRRAFLMELARRKG FRVENELSDSVTHIVAENNSGSDVLEWLQAQKVQVSSQPELLDVSWLIECIRAGKPVEMT GKHQLVVRRDYSDSTNPGPPKTPPIAVQKISQYACQRRTTLNNCNQIFTDAFDILAENCE FRENEDSCVTFMRAASVLKSLPFTIISMKDTEGIPCLGSKVKGIIEEIIEDGESSEVKAV LNDERYQSFKLFTSVFGVGLKTSEKWFRMGFRTLSKVRSDKSLKFTRMQKAGFLYYEDLV SCVTRAEAEAVSVLVKEAVWAFLPDAFVTMTGGFRRGKKMGHDVDFLITSPGSTEDEEQL LQKVMNLWEKKGLLLYYDLVESTFEKLRLPSRKVDALDHFQKCFLIFKLPRQRVDSDQSS WQEGKTWKAIRVDLVLCPYERRAFALLGWTGSRQFERDLRRYATHERKMILDNHALYDKT KRIFLKAESEEEIFAHLGLDYIEPWERNA
3.2-
reverse translation of sp|P04053|TDT_HUMAN DNA nucleotidylexotransferase OS=Homo sapiens OX=9606 GN=DNTT PE=1 SV=3 to a 1527 base sequence of most likely codons.
atggatccgccgcgcgcgagccatctgagcccgcgcaaaaaacgcccgcgccagaccggc gcgctgatggcgagcagcccgcaggatattaaatttcaggatctggtggtgtttattctg gaaaaaaaaatgggcaccacccgccgcgcgtttctgatggaactggcgcgccgcaaaggc tttcgcgtggaaaacgaactgagcgatagcgtgacccatattgtggcggaaaacaacagc ggcagcgatgtgctggaatggctgcaggcgcagaaagtgcaggtgagcagccagccggaa ctgctggatgtgagctggctgattgaatgcattcgcgcgggcaaaccggtggaaatgacc ggcaaacatcagctggtggtgcgccgcgattatagcgatagcaccaacccgggcccgccg aaaaccccgccgattgcggtgcagaaaattagccagtatgcgtgccagcgccgcaccacc ctgaacaactgcaaccagatttttaccgatgcgtttgatattctggcggaaaactgcgaa tttcgcgaaaacgaagatagctgcgtgacctttatgcgcgcggcgagcgtgctgaaaagc ctgccgtttaccattattagcatgaaagataccgaaggcattccgtgcctgggcagcaaa gtgaaaggcattattgaagaaattattgaagatggcgaaagcagcgaagtgaaagcggtg ctgaacgatgaacgctatcagagctttaaactgtttaccagcgtgtttggcgtgggcctg aaaaccagcgaaaaatggtttcgcatgggctttcgcaccctgagcaaagtgcgcagcgat aaaagcctgaaatttacccgcatgcagaaagcgggctttctgtattatgaagatctggtg agctgcgtgacccgcgcggaagcggaagcggtgagcgtgctggtgaaagaagcggtgtgg gcgtttctgccggatgcgtttgtgaccatgaccggcggctttcgccgcggcaaaaaaatg ggccatgatgtggattttctgattaccagcccgggcagcaccgaagatgaagaacagctg ctgcagaaagtgatgaacctgtgggaaaaaaaaggcctgctgctgtattatgatctggtg gaaagcacctttgaaaaactgcgcctgccgagccgcaaagtggatgcgctggatcatttt cagaaatgctttctgatttttaaactgccgcgccagcgcgtggatagcgatcagagcagc tggcaggaaggcaaaacctggaaagcgattcgcgtggatctggtgctgtgcccgtatgaa cgccgcgcgtttgcgctgctgggctggaccggcagccgccagtttgaacgcgatctgcgc cgctatgcgacccatgaacgcaaaatgattctggataaccatgcgctgtatgataaaacc aaacgcatttttctgaaagcggaaagcgaagaagaaatttttgcgcatctgggcctggat tatattgaaccgtgggaacgcaacgcg
3.3- Codon optimazed for Saccharomyces cerevisiae: ATGGATCCACCAAGAGCATCACATTTGTCCCCAAGAAAAAAAAGACCTAGGCAAACTGGTGCATTAATGGCTTCCTCTCCACAGGACATTAAATTCCAAGACTTAGTCGTTTTTATCCTCGAAAAAAAGATGGGTACTACAAGAAGAGCTTTTCTCATGGAATTGGCTAGACGTAAAGGTTTCAGAGTTGAAAATGAACTCAGTGATTCTGTTACACATATTGTTGCTGAAAATAATTCTGGTTCAGATGTTTTGGAATGGTTGCAAGCTCAAAAAGTTCAAGTCTCTTCTCAACCAGAATTGTTAGATGTCAGTTGGTTAATAGAATGCATTAGAGCTGGTAAGCCAGTTGAAATGACTGGCAAACATCAATTAGTAGTAAGAAGAGATTACTCAGATTCAACAAATCCAGGTCCACCAAAAACTCCACCAATAGCTGTTCAAAAAATTTCTCAATACGCTTGTCAACGTAGGACTACTTTAAATAATTGTAATCAAATTTTTACAGATGCTTTTGACATCTTGGCTGAAAACTGTGAATTTAGAGAAAATGAAGATAGCTGCGTTACGTTTATGAGAGCTGCCTCAGTTTTAAAAAGCTTACCTTTTACTATAATTTCTATGAAAGATACCGAAGGAATTCCATGTTTGGGTTCAAAAGTTAAGGGTATTATAGAAGAAATAATTGAAGATGGTGAATCATCAGAAGTTAAAGCAGTGTTGAATGATGAAAGATACCAATCTTTTAAATTATTTACTTCAGTCTTTGGTGTTGGTTTAAAAACCTCCGAAAAATGGTTTAGAATGGGCTTCAGAACCTTGAGTAAGGTGAGATCTGATAAATCATTGAAATTTACGAGAATGCAAAAAGCTGGTTTTTTGTACTACGAGGATTTGGTCAGCTGTGTTACTAGAGCTGAAGCTGAAGCTGTTTCTGTTCTGGTTAAAGAAGCAGTTTGGGCATTTTTGCCTGATGCTTTCGTTACAATGACAGGTGGTTTTAGAAGGGGTAAAAAAATGGGTCATGATGTAGATTTTTTGATTACTTCCCCTGGTTCTACTGAAGATGAAGAACAATTGCTACAAAAAGTGATGAACTTATGGGAGAAAAAAGGTTTGTTGTTGTACTATGATTTAGTTGAGTCTACTTTTGAAAAGTTGAGATTACCATCTCGCAAAGTTGATGCTTTAGATCATTTCCAAAAATGTTTTTTGATCTTTAAGTTGCCAAGACAAAGAGTAGATTCCGATCAGAGTTCCTGGCAAGAAGGTAAGACTTGGAAAGCCATTAGAGTCGACTTGGTTCTGTGTCCATACGAAAGAAGAGCTTTTGCATTATTAGGTTGGACAGGTTCCAGACAATTTGAAAGAGATTTGAGAAGATATGCCACACATGAGAGAAAGATGATTTTAGATAACCATGCATTATATGATAAAACTAAAAGAATCTTCTTGAAAGCCGAATCAGAGGAAGAAATTTTTGCCCATCTAGGTTTGGATTACATTGAACCATGGGAGAGAAATGCT
Avoid cleavage sites of restriction enzymes: BbsI BsaI
3.4- I can produce it in S. cerevisiae via putting the fragment into a vector with the corresponding parts (RBS, promoter, terminator, etc). I can add tags in the 3´ sequence if i wanted to purify it with a desire method. Also cell-free methods could be use to produce it.
Part 4: prepare a Twist DNA Synthesis Order
Part 5: DNA Read/Write/Edit
Part 5.1: DNA Read
5.1- What DNA would you want to sequence (e.g., read) and why?
For the develop of my personal project, i would need to sequence a lot of the plasmids and designs that i will design in it. Also, genomic DNA sequencing to see the integration of the sequences in the corresponding site might be needed.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
TGS. Nanopore, for example. To plasmid and genomic site-especific integration view i would amplify the concentration of the sequences by PCR they, and i would fragmentate (e.g via sonification) the sequences. Then, i would design primers to bind to specific regions (e.g adeine tails) and add adaptors to the sequencing machine and barcodes to aling the fragments, and PCR the sequences. What is next is to prepare the material for the sequencing machine (dilute, pipete). And for transcriptome reading the use of a reverse transcriptase in the firts PCR is needed. For nanopore sequencing, after the preparation of the library, the sample is loaded onto the flow cell, where a flow of ions create a current that passes throught pores. A protein motor guide that pass of the dna throught the pore, where each base disrupt the voltage and the machine is able to interpret it. The raw electrical signals are stored as FAST5 and by algorithms is converted into FASTQ.
5.2 DNA write
(i) What DNA would you want to synthesize (e.g., write) and why?
I would to synthesize the components of the system (e.g transcription factors, kinases, phosphatases) and their respective vectors and cassettes components (promoters, RBSs, terminators). Maybe, as the design process uses tools of protein design, some components may need to be prepared in libraries where certain AA sequences are changign for a more desired output. And, for a more broad characterization, libraries where the components of the cassettes are different.
For now, the components are TF1,TF2, peptidase 1, kinase 1 , cas 13 and their gRNA, condensate andamio, condesate andamio 2, TF3, TF 4 and TF5.
All of this components are general because some would be engineered (modified domains, design of motives, etc).
5.3 DNA edit
(i) What DNA would you want to edit and why?
For my system to work i would need to integrate the components in the genome with, maybe, some recombinase-use engineering.