Week 3 HW: Lab automation
Homework
I create the following code by drawing at the opentrons art platform, coping the coordenates and asking chatgpt to generate the code below the code provided by the original opentrons HTGAA google colabs.
mclover3_points = [(-27, 24),(-24, 24),(-21, 24),(-18, 24),(-30, 21),(-27, 21),(-18, 21),(-15, 21),(-3, 21),(0, 21),(-33, 18),(-30, 18),(-15, 18),(-12, 18),(-3, 18),(0, 18),(-33, 15),(-30, 15),(-24, 15),(-21, 15),(-18, 15),(-12, 15),(-3, 15),(0, 15),(-33, 12),(-27, 12),(-24, 12),(-21, 12),(-18, 12),(-15, 12),(-3, 12),(0, 12),(-33, 9),(-27, 9),(-24, 9),(-12, 9),(-3, 9),(0, 9),(-30, 6),(-9, 6),(-6, 6),(-3, 6),(0, 6),(-33, 3),(-3, 3),(0, 3),(-6, 0),(-3, 0),(0, 0),(15, 0),(18, 0),(21, 0),(24, 0),(27, 0),(-3, -3),(0, -3),(12, -3),(15, -3),(18, -3),(24, -3),(27, -3),(30, -3),(-3, -6),(0, -6),(9, -6),(12, -6),(21, -6),(33, -6),(-3, -9),(0, -9),(9, -9),(15, -9),(18, -9),(21, -9),(24, -9),(27, -9),(30, -9),(-3, -12),(0, -12),(12, -12),(15, -12),(18, -12),(21, -12),(24, -12),(27, -12),(30, -12),(-3, -15),(0, -15),(3, -15),(6, -15),(12, -15),(15, -15),(18, -15),(21, -15),(24, -15),(27, -15),(-3, -18),(0, -18),(-3, -21),(0, -21),(-3, -24),(0, -24),(-3, -27),(0, -27),(-3, -30),(0, -30),(-3, -33),(0, -33),(-3, -36),(0, -36),(-3, -39),(0, -39)]
sfgfp_points = [(-24, 21),(-21, 21),(-27, 18),(-24, 18),(-21, 18),(-18, 18),(-27, 15),(-15, 15),(-30, 12),(-12, 12),(-30, 9),(-9, 9),(-33, 6),(-6, 3), (21, -3),(15, -6),(18, -6),(24, -6),(27, -6),(30, -6),(12, -9),(33, -9),(6, -12),(9, -12),(3, -18)]
electra2_points = [(-3, 24),(0, 24)]
def draw_points(color, points):
DISPENSE_VOL = 0.75
pipette_20ul.pick_up_tip()
for x, y in points:
pipette_20ul.aspirate(DISPENSE_VOL, location_of_color(color))
adjusted_location = center_location.move(types.Point(x=x, y=y))
dispense_and_detach(pipette_20ul, DISPENSE_VOL, adjusted_location)
pipette_20ul.drop_tip()
draw_points('Green', mclover3_points)
draw_points('Red', sfgfp_points)
draw_points('Orange', electra2_points)
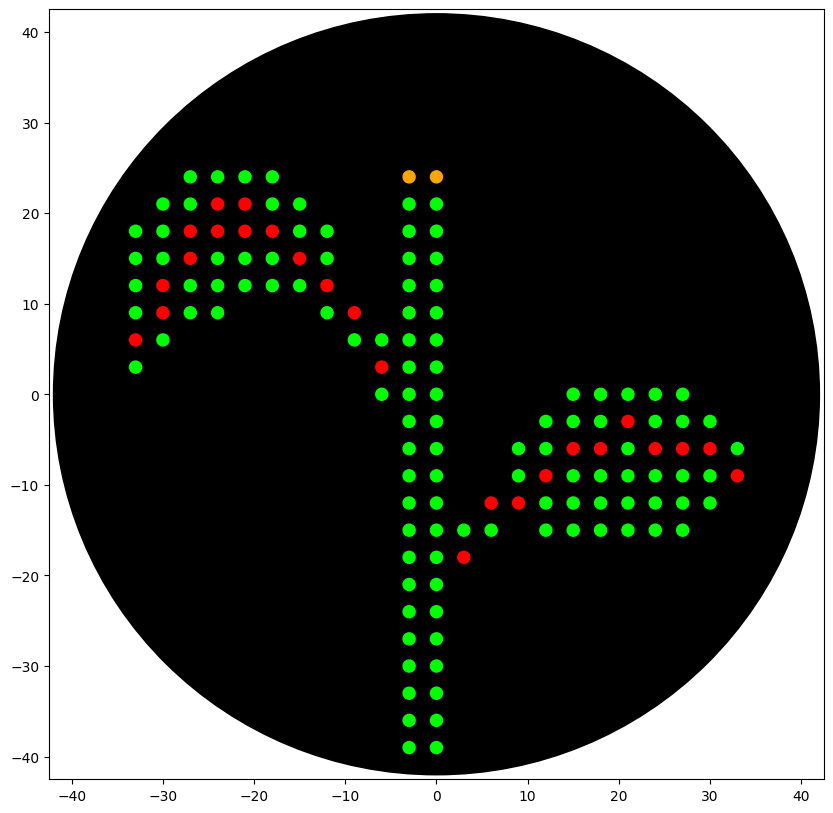
The simulation provide the following artwork.

Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
A massively parallel reporter assay library to screen short synthetic promoters in mammalian cells The paper uses massive parallel reporter assays (MPRAs) as a high-throughput method to cuantify the barcoded mRNA produced by a series of synthetic promoters, and how the regulatory dynamics change over different compounds. They tested 6144 promoters in conventional and specific mammalian cell lines The results were obtein by doing NGS and counting the reads of the mRNAs presents in the cells
Write a description about what you intend to do with automation tools for your final project
For my final project i can use cell-free protein synthesis to screen the post-translational modifications with fluorescence based arrays, in a cloud laboratory. Im going to express the cdks-cyclins complexes, a peptidase, and the correspond TF that would have the post-translational modifications. If im going to use a peptidase that depends on those cdks/cyclins complexes i would be easy, but it firts needs to exist, and not have other activations, otherwise the system would be active at a different time, and i find this very difficult. Another option could be design it If i design the peptidase, firts i should test it in the same type of array. I can build libraries of different changes in the domains and test the perfomance in a well plate.
A possible automated workflow would be:
Echo transfer DNA templates encoding CDK-cyclin complexes, transcription factor variants, protease/peptidase candidates, and fluorescent reporter constructs into specified wells of a 96-well or 384-well plate. Echo transfer any required cofactors, kinase reagents, ATP supplementation conditions, or small molecules into selected wells depending on the test condition. Bravo dispense or stamp cell-free protein synthesis master mix into all wells. Multiflo add lysate or additional common reagents to initiate expression. PlateLoc seal the plate to prevent evaporation. Inheco incubate the plate under defined temperature conditions to allow protein synthesis and post-translational regulation to occur. XPeel remove the seal after incubation. PHERAstar measure fluorescence or time-course reporter output to compare activation, background signal, and dynamic range across designs.
Example pseudocode for automated screening workflow
constructs = load_construct_library([
"CDK_cyclin_variants",
"TF_variants",
"protease_variants",
"reporter_constructs"
])
conditions = define_conditions([
"with_CDK_cyclin",
"without_CDK_cyclin",
"with_protease",
"without_protease",
"different_cofactors",
"different_temperatures"
])
plate = create_plate(format="384-well")
for well in plate:
assign_construct_combination(well, constructs, conditions)
echo_transfer_DNA(well)
echo_transfer_cofactors(well)
bravo_add_mastermix(plate, reagent="CFPS_master_mix")
multiflo_add_lysate(plate)
seal_plate(plate)
incubate_plate(plate, temperature=37, time="defined incubation period")
unseal_plate(plate)
results = measure_fluorescence(plate)
analyze_dynamic_range(results)
rank_best_designs(results)