Week 3 homework
Lab automation 🦾
Python script for Opentrons artwork
- Generate an artistic design using Ronan’s GUI.
- Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script, which draws your design using the Opentrons. You may use AI assistance for this coding — Google Gemini is integrated into Colab (see the stylized star bottom center); it will do a good job writing functional Python, while you probably need to take charge of the art concept.
- If you use AI to help complete this homework or lab, document how you used AI and which models made contributions.
Consistent with this week’s highly automated and digitized theme, for this assignment, I drew inspiration from an image popularized by the Internet, KC Green’s web comic strip “On Fire”, which, in 2014, became a famous -and my personal favorite- online meme (Figure 3.1). As many other people from all over the world, I deeply relate to this meme, which, I feel, accurately describes my life.

 Figure 3.1 Panel from KC Green’s web comic strip “On Fire”, which generated the popular “This is fine” online meme in 2014. Figure from NPR’s report on the related ‘All things considered’ podcast episode.
Figure 3.1 Panel from KC Green’s web comic strip “On Fire”, which generated the popular “This is fine” online meme in 2014. Figure from NPR’s report on the related ‘All things considered’ podcast episode.
As a first step, I fed the right part of the meme into Ronan’s automation art interface, however, the generated artwork required a lot of additional manual processing to resemble the original image (Figure 3.2).
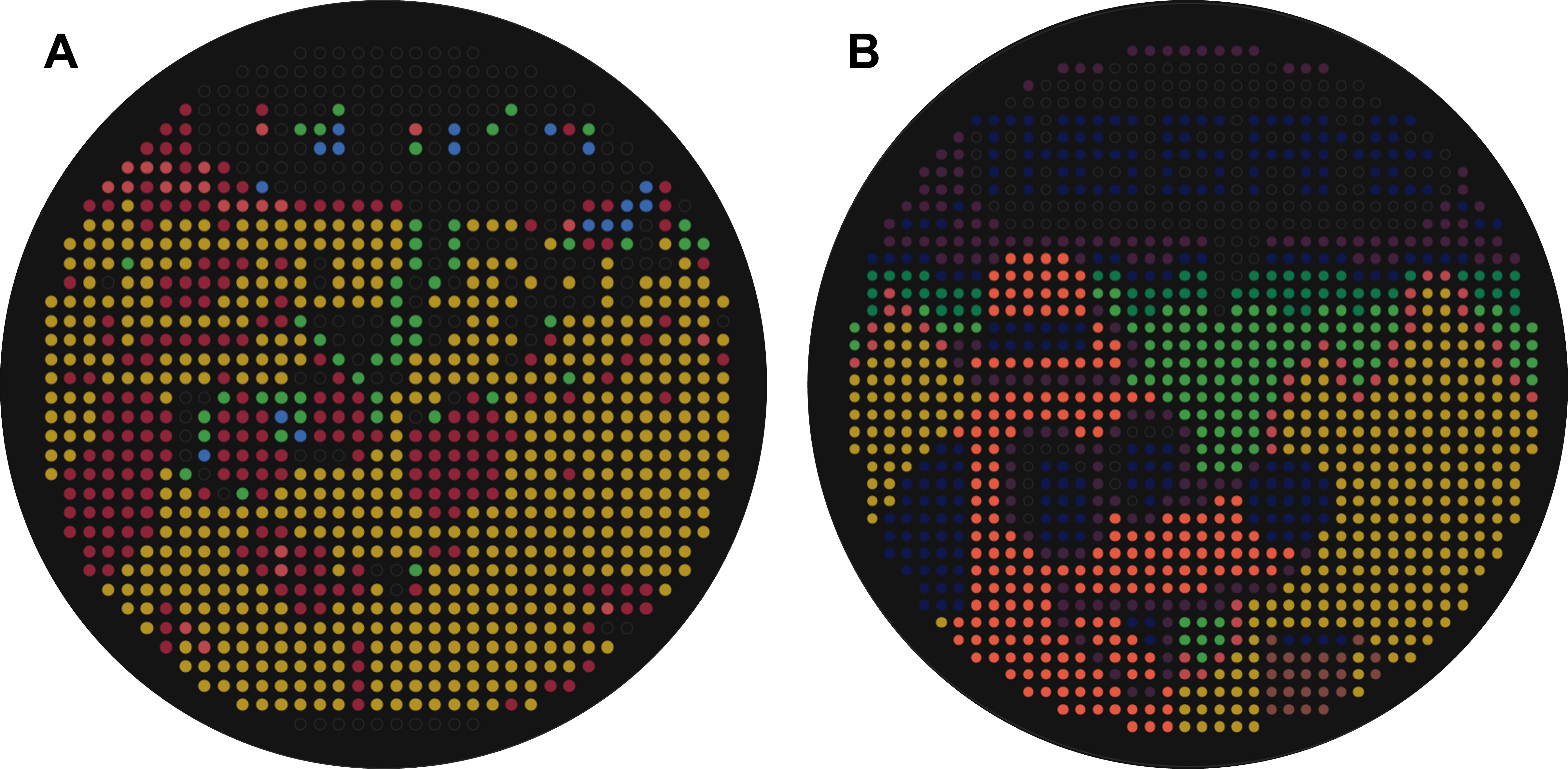
 Figure 3.2 (A) Initial image produced by inserting the right-hand ptych of the “This is fine” meme into Ronan’s automation art interface. (B) Final artwork generated after manual rendering of the initial image.
Figure 3.2 (A) Initial image produced by inserting the right-hand ptych of the “This is fine” meme into Ronan’s automation art interface. (B) Final artwork generated after manual rendering of the initial image.
After composing the final artwork (Figure 3.2B), I imported the nine bacterial dyes I utilized (mClover3, mLychee_TF, mWatermelon, Ultramarine, mKO2, dsRed, mScarlet_I, mCherry, mKate2) along with their respective coordinates into Gemini 2.5 Flash (which was incorporated into my personal copy of HTGAA26 Opentrons Colab notebook) and asked it to write a Python script that would generate the “This is fine”-inspired artwork on a petri dish with the Opentrons system. The Pyhton script obtained from this prompt, slightly augmented with some basic tweaking, including the addition of comments at several steps and functions, as well as renaming the bacterial dyes with their corresponding Hex codes, can be found below:
# Optional, for importing the needed libraries
import subprocess, sys
subprocess.check_call([sys.executable, "-m", "pip", "install", "numpy", "pandas", "opentrons"])
from opentrons import types
metadata = { # see https://docs.opentrons.com/v2/tutorial.html#tutorial-metadata
'Sofia_Oikonomou': '',
'This_is_fine_meme_Opentrons_artwork': '',
'Generate_a_representation_of_the_This_is_fine_meme_with_9_different_bacteria-synthesized_dyes': '',
'source': 'HTGAA 2026 Opentrons Lab',
'apiLevel': '2.20'
}
# Robot deck setup constants - don't change these
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
well_colors = {
'A1' : '#409945',
'A2' : '#40223D',
'A3' : '#E45741',
'A4' : '#B9474B',
'A5' : '#B39223',
'A6' : '#0F7449',
'A7' : '#0F184C',
'A8' : '#45203E',
'A9' : '#7C463E'
}
def run(protocol):
# Load labware, modules and pipettes
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Modules
temperature_module = protocol.load_module('temperature module gen2', COLORS_DECK_SLOT)
# Temperature Module Plate
temperature_plate = temperature_module.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul',
'Cold Plate')
# Choose where to take the colors from
color_plate = temperature_plate
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate') ## TA MUST CALIBRATE EACH PLATE!
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Patterning
# Helper functions for this lab
# pass this e.g. 'Red' and get back a Location which can be passed to aspirate()
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return color_plate[well]
raise ValueError(f"No well found with color {color_string}")
# For this lab, instead of calling pipette.dispense(1, loc) use this: dispense_and_detach(pipette, 1, loc)
def dispense_and_detach(pipette, volume, location):
"""
Move laterally 5mm above the plate (to avoid smearing a drop); then drop down to the plate,
dispense, move back up 5mm to detach drop, and stay high to be ready for next lateral move.
5mm because a 4uL drop is 2mm diameter; and a 2deg tilt in the agar pour is >3mm difference across a plate.
"""
assert(isinstance(volume, (int, float)))
above_location = location.move(types.Point(z=location.point.z + 5)) # 5mm above
pipette.move_to(above_location) # Go to 5mm above the dispensing location
pipette.dispense(volume, location) # Go straight downwards and dispense
pipette.move_to(above_location) # Go straight up to detach drop and stay high
# YOUR CODE HERE to create your design
# Define all points for each dye
all_dye_points = {
#mClover3-Background wall lower
'#409945': [(-11, 11),(-9, 11),(-31, 9),(-9, 9),(-3, 9),(-1, 9),(1, 9),(5, 9),(7, 9),(9, 9),(15, 9),(17, 9),(21, 9),(23, 9),(33, 9),(35, 9),(-39, 7),(-29, 7),(-27, 7),(-25, 7),(-7, 7),(-5, 7),(-3, 7),(-1, 7),(1, 7),(3, 7),(5, 7),(7, 7),(9, 7),(11, 7),(13, 7),(15, 7),(17, 7),(19, 7),(21, 7),(23, 7),(35, 7),(37, 7),(39, 7),(-39, 5),(-5, 5),(-3, 5),(-1, 5),(1, 5),(3, 5),(5, 5),(7, 5),(9, 5),(11, 5),(13, 5),(17, 5),(19, 5),(21, 5),(37, 5),(39, 5),(-5, 3),(-3, 3),(-1, 3),(1, 3),(3, 3),(5, 3),(7, 3),(9, 3),(11, 3),(19, 3),(21, 3),(37, 3),(39, 3),(-7, 1),(-5, 1),(-3, 1),(-1, 1),(1, 1),(3, 1),(5, 1),(7, 1),(9, 1),(19, 1),(39, 1),(-3, -1),(-1, -1),(1, -1),(3, -1),(5, -1),(7, -1),(9, -1),(-1, -3),(1, -3),(3, -3),(5, -3),(7, -3),(1, -5),(3, -5),(5, -5),(7, -5),(1, -7),(3, -7),(5, -7),(7, -7),(1, -9),(3, -9),(5, -9),(-1, -27),(1, -27),(3, -27),(-1, -29),(1, -29),(3, -29),(1, -31)],
#mCherry-Dog outline
'#40223D': [(-29, 25),(-27, 25),(-25, 15),(-23, 15),(-13, 15),(-11, 9),(-9, 7),(-27, 5),(-25, 5),(-7, 5),(-27, 3),(-7, 3),(-25, 1),(-23, 1),(-21, 1),(-19, 1),(-17, 1),(-15, 1),(-13, 1),(-11, 1),(-9, 1),(-7, -3),(-5, -3),(-3, -3),(-19, -5),(-17, -5),(-15, -5),(-9, -5),(-1, -5),(-21, -7),(-13, -7),(-11, -7),(-1, -7),(-21, -9),(-11, -9),(-1, -9),(7, -9),(-21, -11),(-11, -11),(-1, -11),(1, -11),(3, -11),(5, -11),(-21, -13),(-11, -13),(-9, -13),(-3, -13),(-1, -13),(1, -13),(-21, -15),(-11, -15),(-7, -15),(-5, -15),(-19, -17),(-11, -17),(-17, -19),(-15, -19),(-13, -19),(13, -19),(11, -21),(3, -23),(5, -23),(7, 23),(9, -23),(-9, -25),(-7, -25),(-5, -25),(-3, -25),(-1, -25),(1, -25),(3, -25),(-3, -27),(-3, -29),(-11, -31),(-3, -31),(-9, -33),(-3, -33),(-7, -35),(-5, -35)],
#mLychee_TF-Dog and hat
'#E45741': [(-21, 15),(-19, 15),(-17, 15),(-15, 15),(-23, 13),(-21, 13),(-19, 13),(-17, 13),(-15, 13),(-13, 13),(-23, 11),(-21, 11),(-19, 11),(-17, 11),(-15, 11),(-13, 11),(-23, 9),(-21, 9),(-19, 9),(-17, 9),(-15, 9),(-13, 9),(-11, 7),(-11, 5),(-9, 5),(-25, 3),(-23, 3),(-21, 3),(-19, 3),(-17, 3),(-15, 3),(-13, 3),(-11, 3),(-9, 3),(-23, -1),(-21, -1),(-19, -1),(-17, -1),(-15, -1),(-13, -1),(-11, -1),(-9, -1),(-7, -1),(-5, -1),(-25, -3),(-23, -3),(-21, -3),(-19, -3),(-17, -3),(-15, -3),(-13, -3),(-11, -3),(-9, -3),(-27, -5),(-25, -5),(-23, -5),(-21, -5),(-13, -5),(-11, -5),(-25, -7),(-23, -7),(-25, -9),(-23, -9),(-25, -11),(-23, -11),(-25, -13),(-23, -13),(3, -13),(5, -13),(-25, -15),(-23, -15),(-9, -15),(-3, -15),(-1, -15),(1, -15),(3, -15),(5, -15),(-25, -17),(-23, -17),(-21, -17),(-9, -17),(-7, -17),(-5, -17),(-3, -17),(-1, -17),(1, -17),(3, -17), (5, -17),(7, -17),(-25, -19),(-23, -19),(-21, -19),(-19, -19),(-11, -19),(-9, -19),(-7, -19),(-5, -19),(-3, -19),(-1, -19),(1, -19),(3, -19),(5, -19),(7, -19),(9, -19),(11, -19),(-25, -21),(-23, -21),(-21, -21),(-19, -21),(-17, -21),(-15, -21),(-13, -21),(-11, -21),(-9, -21),(-7, -21),(-5, -21),(-3, -21),(-1, -21),(1, -21),(3, -21),(5, -21),(7, -21),(9, -21),(-25, -23),(-23, -23),(-21, -23),(-19, -23),(-17, -23),(-13, -23),(-11, -23),(-9, -23),(-7, -23),(-5, -23),(-3, -23),(-1, -23),(1, -23),(-25, -25),(-23, -25),(-21, -25),(-19, -25),(-17, -25),(-27, -27),(-25, -27),(-23, -27),(-21, -27),(-19, -27),(-17, -27),(-13, -27),(-11, -27),(-9, -27),(-27, -29),(-25, -29),(-23, -29),(-21, -29),(-19, -29),(-17, -29),(-15, -29),(-11, -29),(-9, -29),(-7, -29),(-25, -31),(-23, -31),(-21, -31),(-19, -31),(-17, -31),(-15, -31),(-13, -31),(-9, -31),(-7, -31),(-5, -31),(-21, -33),(-19, -33),(-17, -33),(-15, -33),(-13, -33),(-11, -33),(-7, -33),(-5, -33),(-19, -35),(-17, -35),(-15, -35),(-13, -35),(-11, -35),(-9, -35),(-3, -35),(-15, -37),(-13, -37),(-11, -37),(-9, -37),(-7, -37),(-5, -37),(-3, -37),(-7, -39),(-5, -39),(-3, -39)],
#mScarlet_I-Flame outline
'#B9474B': [(27, 13),(29, 13),(-35, 11),(25, 11),(31, 11),(-35, 9),(-33, 9),(25, 9),(31, 9),(-37, 7),(-31, 7),(25, 7),(33, 7),(-37, 5),(-29, 5),(15, 5),(23, 5),(35, 5),(-39, 3),(13, 3),(17, 3),(23, 3),(35, 3),(11, 1),(17, 1),(21, 1),(37, 1),(11, -1),(19, -1),(39, -1),(9, -3),(9, -5),(9, -7),(5, -25),(5, -27),(5, -29),(-1, -31),(3, -31),(-1, -33),(1, -33)],
#mKO2-Flame body
'#B39223': [(27, 11),(29, 11),(27, 9),(29, 9),(-35, 7),(-33, 7),(27, 7),(29, 7),(31, 7),(-35, 5),(-33, 5),(-31, 5),(25, 5),(27, 5),(29, 5),(31, 5),(33, 5),(-37, 3),(-35, 3),(-33, 3),(-31, 3),(-29, 3),(15, 3),(25, 3),(27, 3),(29, 3),(31, 3),(33, 3),(-39, 1),(-37, 1),(-35, 1),(-33, 1),(-31, 1),(-29, 1),(-27, 1),(13, 1),(15, 1),(23, 1),(25, 1),(27, 1),(29, 1),(31, 1),(33, 1),(35, 1),(-39, -1),(-37, -1),(-35, -1),(-33, -1),(-31, -1),(-29, -1),(-27, -1),(-25, -1),(13, -1),(15, -1),(17, -1),(21, -1),(23, -1),(25, -1),(27, -1),(29, -1),(31, -1),(33, -1),(35, -1),(37, -1),(-39, -3),(-37, -3),(-35, -3),(-33, -3),(-31, -3),(-29, -3),(-27, -3),(11, -3),(13, -3),(15, -3),(17, -3),(19, -3),(21, -3),(23, -3),(25, -3),(27, -3),(29, -3),(31, -3),(33, -3),(35, -3),(37, -3),(39, -3),(-39, -5),(-37, -5),(-35, -5),(-33, -5),(-31, -5),(-29, -5),(11, -5),(13, -5),(15, -5),(17, -5),(19, -5),(21, -5),(23, -5),(25, -5),(27, -5),(29, -5),(31, -5),(33, -5),(35, -5),(37, -5),(39, -5),(-39, -7),(-37, -7),(-35, -7),(-33, -7),(-31, -7),(11, -7),(13, -7),(15, -7),(17, -7),(19, -7),(21, -7),(23, -7),(25, -7),(27, -7),(29, -7),(31, -7),(33, -7),(35, -7),(37, -7),(39, -7),(-37, -9),(-35, -9),(-33, -9),(15, -9),(17, -9),(19, -9),(21, -9),(23, -9),(25, -9),(27, -9),(29, -9),(31, -9),(33, -9),(35, -9),(37, -9),(-37, -11),(-35, -11),(17, -11),(19, -11),(21, -11),(23, -11),(25, -11),(27, -11),(29, -11),(31, -11),(33, -11),(35, -11),(37, -11),(-37, -13),(-35, -13),(17, -13),(19, -13),(21, -13),(23, -13),(25, -13),(27, -13),(29, -13),(31, -13),(33, -13),(35, -13),(37, -13),(-37, -15),(17, -15),(19, -15),(21, -15),(23, -15),(25, -15),(27, -15),(29, -15),(31, -15),(33, -15),(35, -15),(37, -15),(15, -17),(17, -17),(19, -17),(21, -17),(23, -17),(25, -17),(27, -17),(29, -17),(31, -17),(33, -17),(35, -17),(15, -19),(17, -19),(19, -19),(21, -19),(23, -19),(25, -19),(27, -19),(29, -19),(31, -19),(33, -19),(35, -19),(13, -21),(15, -21),(17, -21),(19, -21),(21, -21),(23, -21),(25, -21),(27, -21),(29, -21),(31, -21),(33, -21),(11, -23),(13, -23),(15, -23),(17, -23),(19, -23),(21, -23),(23, -23),(25, -23),(27, -23),(29, -23),(31, -23),(7, -25),(9, -25),(11, -25),(13, -25),(15, -25),(17, -25),(19, -25),(21, -25),(23, -25),(25, -25),(27, -25),(29, -25),(31, -25),(-29, -27),(7, -27),(9, -27),(11, -27),(13, -27),(15, -27),(17, -27),(19, -27),(21, -27),(23, -27),(25, -27),(27, -27),(29, -27),(7, -29),(21, -29),(23, -29),(25, -29),(27, -29),(5, -31),(7, -31),(19, -31),(23, -31),(25, -31),(3, -33),(5, -33),(7, -33),(19, -33),(-1, -35),(1, -35),(3, -35),(5, -35),(7, -35),(19, -35),(-1, -37),(1, -37),(3, -37),(5, -37),(7, -37),(-1, -39),(1, -39),(3, -39),(5, -39),(7, -39)],
#mWatermelon-Background wall upper
'#0F7449': [(-37, 13),(-35, 13),(-33, 13),(-31, 13),(-11, 13),(-9, 13),(-1, 13),(1, 13),(7, 13),(9, 13),(11, 13),(13, 13),(15, 13),(17, 13),(25, 13),(31, 13),(33, 13),(35, 13),(37, 13),(-37, 11),(-33, 11),(-31, 11),(-29, 11),(-27, 11),(-25, 11),(-7, 11),(-5, 11),(-3, 11),(-1, 11),(1, 11),(5, 11),(7, 11),(9, 11),(11, 11),(13, 11),(15, 11),(17, 11),(19, 11),(21, 11),(23, 11),(33, 11),(35, 11),(37, 11),(-37, 9),(-29, 9),(-27, 9),(-25, 9),(-7, 9),(-5, 9),(11, 9),(13, 9),(19, 9),(37, 9)],
#Ultramarine-Dark blue elements
'#0F184C': [(-25, 31),(-23, 31),(-21, 31),(-19, 31),(-15, 31),(-13, 31),(-11, 31),(-9, 31),(-7, 31),(-3, 31),(-1, 31),(1, 31),(3, 31),(7, 31),(9, 31),(11, 31),(13, 31),(15, 31),(21, 31),(23, 31),(25, 31),(-23, 29),(-19, 29),(-15, 29),(-13, 29),(-11, 29),(-3, 29),(-1, 29),(7, 29),(13, 29),(15, 29),(17, 29),(21, 29),(23, 29),(-23, 27),(-19, 27),(-17, 27),(-15, 27),(-13, 27),(-11, 27),(-9, 27),(-7, 27),(-3, 27),(-1, 27),(1, 27),(3, 27),(7, 27),(9, 27),(11, 27),(13, 27),(15, 27),(19, 27),(21, 27),(23, 27),(25, 27),(27, 27),(-23, 25),(-19, 25),(-15, 25),(-13, 25),(-7, 25),(-3, 25),(3, 25),(7, 25),(13, 25),(15, 25),(21, 25),(23, 25),(-23, 23),(-19, 23),(-15, 23),(-13, 23),(-11, 23),(-9, 23),(-7, 23),(-3, 23),(-1, 23),(1, 23),(3, 23),(7, 23),(13, 23),(15, 23),(21, 23),(23, 23),(25, 23),(27, 23),(-27, 21),(31, 21),(-33, 19),(-31, 19),(-29, 19),(33, 19),(35, 19),(-37, 15),(-35, 15),(-33, 15),(-31, 15),(-11, 15),(-9, 15),(-1, 15),(1, 15),(9, 15),(11, 15),(13, 15),(15, 15),(17, 15),(25, 15),(27, 15),(29, 15),(31, 15),(-29, 13),(-27, 13),(-25, 13),(-7, 13),(-5, 13),(-3, 13),(19, 13),(21, 13),(23, 13),(-23, 7),(-21, 7),(-19, 7),(-17, 7),(-15, 7),(-13, 7),(-23, 5),(-21, 5),(-19, 5),(-17, 5),(-15, 5),(-13, 5),(-29, -7),(-27, -7),(-7, -7),(-5, -7),(-3, -7),(-31, -9),(-29, -9),(-27, -9),(-17, -9),(-15, -9),(-7, -9),(-5, -9),(-3, -9),(9, -9),(11, -9),(13, -9),(-33, -11),(-31, -11),(-29, -11),(-27, -11),(-17, -11),(-15, -11),(-13, -11),(-7, -11),(-5, -11),(-3, -11),(7, -11),(9, -11),(11, -11),(13, -11),(15, -11),(-33, -13),(-31, -13),(-29, -13),(-27, -13),(-17, -13),(-15, -13),(-13, -13),(-5, -13),(7, -13),(9, -13),(11, -13),(13, -13),(15, -13),(-35, -15),(-33, -15),(-31, -15),(-29, -15),(-27, -15),(-17, -15),(-15, -15),(-13, -15),(7, -15),(9, -15),(11, -15),(13, -15),(15, -15),(-35, -17),(-33, -17),(-31, -17),(-29, -17),(-27, -17),(-17, -17),(-15, -17),(-13, -17),(9, -17),(11, -17),(13, -17),(-35, -19),(-33, -19),(-31, -19),(-29, -19),(-27, -19),(-33, -21),(-31, -21),(-29, -21),(-27, -21),(-31, -23),(-29, -23),(-27, -23),(-31, -25),(-29, -25),(-27, -25),(-7, -27),(-5, -27),(-5, -29),(11, -29),(13, -29),(15, -29),(17, -29)],
#mKate2-Puff of smoke
'#45203E': [(-7, 39),(-5, 39),(-3, 39),(-1, 39),(1, 39),(3, 39),(5, 39),(7, 39),(-15, 37),(-13, 37),(-11, 37),(11, 37),(13, 37),(15, 37),(-19, 35),(-27, 29),(-29, 27),(-27, 27),(-31, 25),(31, 25),(-31, 23),(-29, 23),(-27, 23),(31, 23),(-33, 21),(-31, 21),(-29, 21),(-25, 21),(29, 21),(33, 21),(-35, 19),(-27, 19),(-25, 19),(-23, 19),(27, 19),(29, 19),(31, 19),(-35, 17),(-33, 17),(-31, 17),(-29, 17),(-27, 17),(-25, 17),(-23, 17),(-21, 17),(-19, 17),(-17, 17),(-15, 17),(-13, 17),(-11, 17),(-9, 17),(-7, 17),(-5, 17),(-3, 17),(-1, 17),(1, 17),(9, 17),(11, 17),(13, 17),(15, 17),(17, 17),(19, 17),(21, 17),(23, 17),(25, 17),(27, 17),(29, 17),(31, 17),(33, 17),(35, 17),(-29, 15),(-27, 15),(-7, 15),(-5, 15),(-3, 15),(19, 15),(21, 15),(23, 15),(33, 15),(35, 15),(37, 15),(-15, -23),(-15, -25),(-13, -25),(-11, -25),(-15, -27),(-13, -29)],
#dsRed-Coffee mug
'#7C463E': [(9, -29),(19, -29),(9, -31),(11, -31),(13, -31),(15, -31),(17, -31),(21, -31),(9, -33),(11, -33),(13, -33),(15, -33),(17, -33),(21, -33),(9, -35),(11, -35),(13, -35),(15, -35),(17, -35),(9, -37),(11, -37),(13, -37),(15, -37)]
}
for dye_name, points_list in all_dye_points.items():
if not points_list: # Skip if no points for this dye
continue
color_source_well = location_of_color(dye_name)
pipette_20ul.pick_up_tip()
# Aspirate in batches to ensure enough liquid
for i, (x_coord, y_coord) in enumerate(points_list):
if i % 20 == 0 or pipette_20ul.current_volume < 1: # Re-aspirate if volume is low or every 20 points
# Aspirate enough for remaining points or 20uL, whichever is less
volume_to_aspirate = min(20, len(points_list) - i)
if volume_to_aspirate > 0:
pipette_20ul.aspirate(volume_to_aspirate, color_source_well)
adjusted_location = center_location.move(types.Point(x=x_coord, y=y_coord))
dispense_and_detach(pipette_20ul, 1, adjusted_location)
pipette_20ul.drop_tip()
# Don't forget to end with a drop_tip()
# Execute Simulation / Visualization -- don't change this code block
protocol = OpentronsMock(well_colors)
run(protocol)
protocol.visualize()
Lastly, when I simulated the artwork, the Python script presented above did produce the designed image (Figure 3.3).
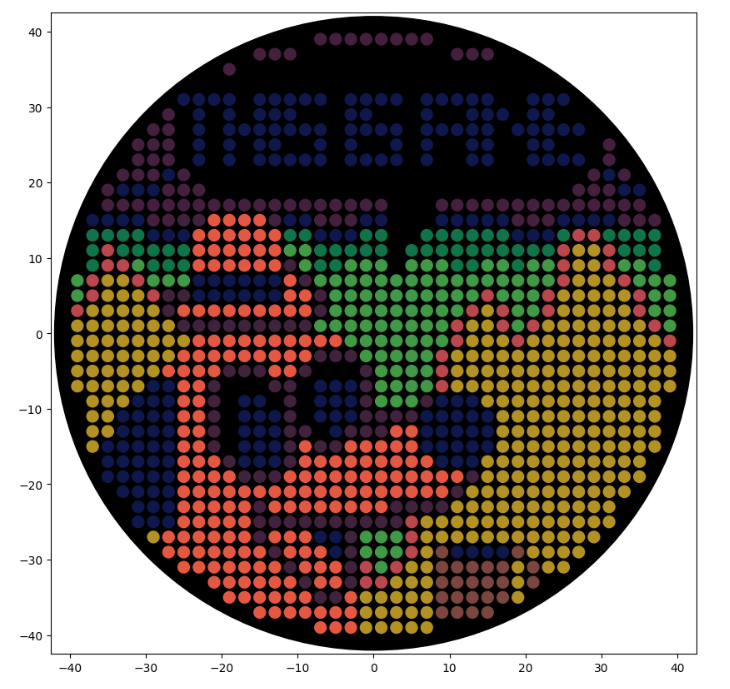
 Figure 3.3 Simulation of the “This is fine” artwork generated by running the Python script displayed above.
Figure 3.3 Simulation of the “This is fine” artwork generated by running the Python script displayed above.
Post-lab questions related to using the Opentrons system
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely. For this week:
- Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
In their paper from March 2026 1, Kostanjšek et al. present the development of Rhodo-Box, a toolkit of standardized genetic parts for the emerging Synthetic Biology chassis Rhodobacter sphaeroides. R. sphaeroides is a purple non-sulfur alphaproteobacterium with a highly versatile metabolism, including photosynthetic pathways, which render it a promising platform for the sustainable biosynthesis of numerous compounds. To realize the microorganism’s full potential, the researchers built and characterized a collection of modular genetic elements specifically tailored for use in R. sphaeroides.
The modular parts include three broad-host origins of replication (ORIs), namely RSF1010, pBBR1, and RK2 functioning as high-, medium-, and medium-copy-number ORIs respectively. Also found in the toolbox are 13 constitutive promoters of native (like PJ95025), heterologous (such as P crtE), and artificial origin (like PJ23100), spanning a 270-fold dynamic range, as well as 11 ribosome binding sites (RBSs), originally designed for E. coli, for instance, B0034, or for R. sphaeroides, such as J95028, for translational regulation, spanning a 49-fold dynamic range. Another significant feature of Rhodo-Box are the four inducible expression systems assessed in the context of the study, namely NahR-P salTTC, LacI-P lacT7A1_O3O4, VanR-P vanCC, and XylS-Pm. Among them, NahR-P salTTC and VanR-P vanCC, induced by salicylic and vanillic acid each, appeared to be the most promising ones as they displayed high tunability and low basal expression, with the former comprising an appealing option for industrial-scale biomanufacturing due to the affordability of salicylic acid. To enable further flexibility and orthogonality of the modular cloning strategy, the authors proceeded to construct plasmid backbones for Rhodo-Box by combining the ORIs mentioned above with common antibiotic resistance markers, while ensuring that the interaction of the different promoter and RBS sequences tested would not generate genetic context-dependent effects influencing the level of expression.
To this end, meaning to cross-screen for context-dependent interactions among Rhodo-Box’s components, Kostanjšek et al. implemented a semi-automated cloning workflow to build a R. sphaeroides strain library in a time-efficient manner using an Opentrons OT2 platform. In particular, they programmed the Opentrons liquid-handling robot to assemble five promoters with six RBSs and four transcriptional terminators to yield a total of 42 constructs (Figure 3.4). Through this process, they obtained 38 correctly assembled strains, corresponding to an overall 90% success rate in genetic circuit construction, and, concurrently, reduced the time required to build this library by 50%.
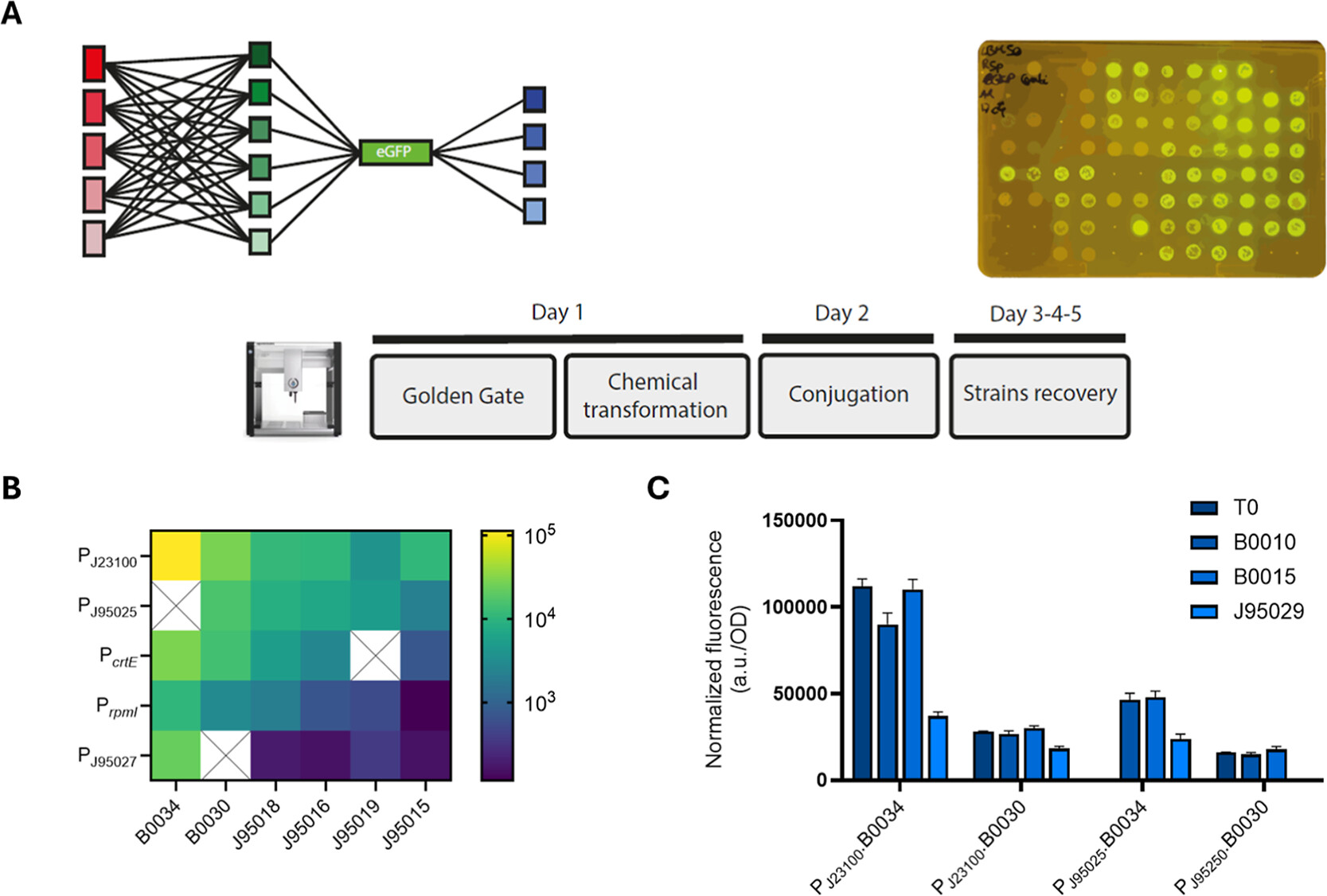 Figure 3.4 Combinatorial characterization of parts from the Rhodo-Box toolkit using a semiautomated cloning workflow on the Opentrons OT2 platform. (A) General layout of the semiautomated workflow, combining five promoters (red) with six RBSs (green), and characterizing four terminators (blue) with eGFP. (B) Heat-map representing the combinatorial promoters and RBSs' strengths in *R. sphaeroides*. (C) Normalized fluorescence of tested terminators T0, B0010, B0015, J95029 expressed in combinations of P~J23100~ and P~J95025~ with B0034 and B0030. Graphs (B, C) represent the average fluorescence values normalized with OD~600~, with the standard deviation of three technical replicates depicted in (C). Figure from Kostanjšek et al., 2026 [^1].
Figure 3.4 Combinatorial characterization of parts from the Rhodo-Box toolkit using a semiautomated cloning workflow on the Opentrons OT2 platform. (A) General layout of the semiautomated workflow, combining five promoters (red) with six RBSs (green), and characterizing four terminators (blue) with eGFP. (B) Heat-map representing the combinatorial promoters and RBSs' strengths in *R. sphaeroides*. (C) Normalized fluorescence of tested terminators T0, B0010, B0015, J95029 expressed in combinations of P~J23100~ and P~J95025~ with B0034 and B0030. Graphs (B, C) represent the average fluorescence values normalized with OD~600~, with the standard deviation of three technical replicates depicted in (C). Figure from Kostanjšek et al., 2026 [^1].By coupling the validation of the modular design principles underlying the Rhodo-Box toolkit with the automated approach facilitated by the Opentrons OT2 platform, the researchers hope to further expand the repertoire of genetic circuits built for expression in R. sphaeroides, so that this novel chassis can be employed for more advanced Metabolic Engineering and Synthetic Biology applications.
- Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details. While your description/project idea doesn’t need to be set in stone, we would like to see core details of what you would automate.
A significantly large portion of my experience of Synthetic Biology stems from working in the wet lab, so I hope you can understand my reluctance to adopt a more automated approach. However, I believe that automation can substantially help with highly repetitive tasks in order to accelerate several processes, rendering them more high-throughput.
In the context of one of my final ideas in particular, I would like to combine a genetically modified fungus with a cyanobacterium or a microalga to generate a synthetic lichen. One of the major challenges of building a lichen from the bottom up lies in establishing the appropriate organisms partaking in the symbiotic relationship. Although there are hundreds of naturally occurring lichen species 2 that could function as a basic foundation for this investigation, it would be interesting to experiment with different combinations of fungi and photobionts (possibly more than one from each category per combination). Exploring novel microorganism combinations for the generation of a synthetic lichen could lead to optimizing existing species but also to creating an artificial lichen that constitutes a particularly suitable platform for the proposed application, which is a color-shifting building coating. Opentrons could be deployed both to prepare the initial separate axenic cultures of the different fungi and photobionts and to generate the mixed co-culture combinations. Another difficulty arises from growth incompatibility between the different symbiotic partners, so Opentrons could also contribute to visually inspecting the several co-cultures through a spectrophotometer function to pinpoint the combinations where the growth rate of the fungus (or fungi) matches the respective growth rate of the photobiont(s) in a stable and consistent manner. Additionally, this kind of visual inspection performed by the Opentrons system could ensure that the cultures stay uncontaminated (which can be revealed through alterations in light absorption too), as well as identify the co-cultures which display the potential for a lichen’s structural complexity. Rather that merely co-existing in the same culture, the symbionts should also demonstrate the capacity to recreate the highly organized, layered structure of natural lichens (called a thallus), which constitutes an additional criterion for distinguishing the most promising candidates. To this end, apart from utilizing the Opentrons platform for co-culture maintenance and surveillance, I could design and 3D-print a 24-well plate embedded with a micro-scaffold that would “encourage” the formation of a rudimentary thallus by the symbiotic microorganisms. Lastly, as genetic parts for putative lichen partners are often not as well-established as for model organisms, such as E. coli, several versions of the color-changing genetic construct should be assembled, each combining different genetic parts. In that case, Opentrons could be extremely helpful in screening the performance of all different circuit combinations in a high-throughput manner, as well as in characterizing a number of novel parts, engineered or optimized for the purposed of the project. Since all three of my proposals involve the assembly and integration of a genetic construct into a biological system, the same screening of genetic circuits could be conducted by Opentrons to a similar end for my other final project ideas too.
Kostanjšek M, Raynal A, Dimopoulos G, et al. Rhodo-Box: A Synthetic Biology Toolbox to Facilitate Metabolic Engineering of Rhodobacter sphaeroides. ACS Synth Biol. March 15, 2026. doi:10.1021/acssynbio.5c00808 ↩︎
Lutzoni F, Miadlikowska J. Lichens. Curr Biol. 2009;19(13):R502-3. doi:10.1016/j.cub.2009.04.034 ↩︎