HTGAA - Week 1: Principles and Practices


My Homework
WEEK 1 - SAVE THE WORLD OR DESTROY THE WORLD
This week lays the foundation for ethics, safety, and governance in biotechnology.
Lecture (Tues, Feb 3)
Principles & Practices
(▶️Recording)
David Kong
George Church
Joe Jacobson
Recitation (Wed, Feb 4)
Principles, Ethics, Practices
(▶️Recording |
💻Slides)
Ronan Donovan
Suvin Sundararajan
Subastian Kamau
Greg Galperin
Instructions
First, describe a biological engineering application or tool you want to develop and why. This could be inspired by an idea for your HTGAA class project and/or something for which you are already doing in your research, or something you are just curious about.
Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals. Below is one example framework (developed in the context of synthetic genomics) you can choose to use or adapt, or you can develop your own. The example was developed to consider policy goals of ensuring safety and security, alongside other goals, like promoting constructive uses, but you could propose other goals for example, those relating to equity or autonomy.
Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”). Try to outline a mix of actions (e.g. a new requirement/rule, incentive, or technical strategy) pursued by different “actors” (e.g. academic researchers, companies, federal regulators, law enforcement, etc). Draw upon your existing knowledge and a little additional digging, and feel free to use analogies to other domains (e.g. 3D printing, drones, financial systems, etc.).
Example
- Purpose: What is done now and what changes are you proposing?
- Design: What is needed to make it “work”? (including the actor(s) involved - who must opt-in, fund, approve, or implement, etc)
- Assumptions: What could you have wrong (incorrect assumptions, uncertainties)?
- Risks of Failure & “Success”: How might this fail, including any unintended consequences of the “success” of your proposed actions?
Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals. The following is one framework but feel free to make your own:
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | |||
| • By helping respond | |||
| Foster Lab Safety | |||
| • By preventing incident | |||
| • By helping respond | |||
| Protect the environment | |||
| • By preventing incidents | |||
| • By helping respond | |||
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | |||
| • Feasibility? | |||
| • Not impede research | |||
| • Promote constructive applications |
- Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties. For this, you can choose one or more relevant audiences for your recommendation, which could range from the very local (e.g. to MIT leadership or Cambridge Mayoral Office) to the national (e.g. to President Biden or the head of a Federal Agency) to the international (e.g. to the United Nations Office of the Secretary-General, or the leadership of a multinational firm or industry consortia). These could also be one of the “actor” groups in your matrix.
PART 1. FIXING THE COURSE
- First, describe a biological engineering application or tool you want to develop and why. This could be inspired by an idea for your HTGAA class project and/or something for which you are already doing in your research, or something you are just curious about.
NOTE: This project is just the initial idea, it can be subjected to changes and upgrades in the near future.
Project: Reversible Cell-Free Biosensor for ROS-Mediated Radiation Damage
This project aims to design a reversible, cell-free biosensor capable of reporting radiation-induced oxidative damage through a visible biochemical signal.
The system is based on a DNA-programmed TX–TL circuit embedded within a hydrogel matrix, inspired by biological systems that can transition between active and inactive states under physical stress. Upon exposure to radiation-induced reactive oxygen species (ROS), the biosensor activates a transient fluorescent response, which gradually returns to a basal state once the stimulus is removed, enabling reuse of the material.
By decoupling damage sensing from living cells, this platform provides a controllable and modular approach to studying radiation effects on biological matter.
One-sentence project goal
The goal of this project is to engineer a reversible, reusable, cell-free biosensor that translates radiation-induced oxidative damage into a transient biochemical signal.
Background, application and why does it matter
The primary application of this biosensor is in radiation physics, medical physics and even space science, where it can be used as a reusable biological dosimetry platform to study oxidative damage induced by ionizing radiation.
Rather than measuring radiation directly, the system reports biologically relevant damage, specifically ROS generation, offering insight into how physical radiation translates into molecular stress in biological systems. This makes the material particularly valuable for experimental radiation setups, calibration studies, and comparative stress assays, without the need for living models.
The material functions as a reversible biological stress reporter. Instead of permanently activating or degrading under radiation-induced stress, it temporarily switches state to signal damage and then returns to baseline, enabling repeated use and long-term monitoring.
In medical physics and radiobiology, many existing sensing systems present fundamental limitations:
- They degrade over time
- They saturate under high stimulus
- They are single-use
- They cannot be reset or recovered
Similarly, most biological sensors:
- lose viability
- or remain irreversibly activated after damage
This creates a gap between physical radiation sensing and biologically meaningful damage reporting. The hydrogel is not just a container. While individual stress-responsive genetic elements are well characterized, their integration into a reusable, reversible cell-free biomaterial capable of multiple stress-response cycles remains largely unexplored.
Inspiration
The project is inspired by simple biological systems, such as jellyfish, which exhibit functional resilience and reversible state transitions despite minimal organizational complexity. These organisms demonstrate that biological function does not always require permanent activation or structural complexity, but can instead rely on transient, physics-driven responses to environmental stress.
Translating this principle into a synthetic, cell-free context, the proposed biosensor explores how biological states—such as gene expression and signal emission—can be reversibly triggered by physical damage and allowed to relax back to a stable baseline.
What makes this a synthetic biology project
This project constitutes a synthetic biology approach by designing and programming a DNA-based TX–TL circuit that links oxidative stress sensing to a controlled biochemical output to manifest a visible fluorescent signal. The circuit architecture, combined with material constraints imposed by the hydrogel matrix, enables tunable activation, decay, and reversibility of the signal.
Signal intensity correlates with stress magnitude, while signal reversibility reflects the system’s ability to recover to a baseline state. System reversibility is achieved through the co-design of a stress-responsive genetic circuit and a diffusion-regulated material matrix, enabling transient activation and passive return to a basal state without permanent system alteration. The system does not shut down because it fails; it shuts down because it is designed to relax back to its original state.
This platform is thinked to be modular, allowing future expansion to additional damage types. Rather than engineering a new organism, the project focuses on engineering biological function, emphasizing control, modularity, and reusability.
Conceptual state transition
- The system starts in an OFF (basal) state
- Oxidative stress is applied (e.g. H₂O₂ or radiation-induced ROS)
- The system enters a “damage state”
- A fluorescent signal is activated
- The stress is removed
- The system relaxes back to its basal state
Engineering design decisions
| Biological Circuit Controls | Material (Hydrogel) Controls |
|---|---|
| What is detected (ROS, damage, stress) | How much stimulus enters the system |
| What signal is produced (fluorescence) | How fast the stimulus diffuses |
| Activation threshold and sensitivity | How long the stimulus is retained |
| Timing of signal initiation | Rate of stimulus clearance |
| Duration of protein expression | Smoothness of system shutdown |
| Signal termination mechanisms | Buffering of damage spikes |
| Susceptibility to noise or false positives | Protection of TX–TL components |
Key tunable parameters in the system design include:
- duration of protein expression
- protein degradation rate
- response speed
- energy consumption
- lifetime of the TX–TL system
Primary and secondary reporting strategy
- Primary signal: fluorescence intensity
- Secondary signal: temporal dynamics of activation and decay
Interpretation:
- Fluorescence intensity reflects the magnitude of ROS-induced damage
- Signal duration and decay profile reflect the dynamic response of the system under stress
Simplifying
How much it glows → magnitud of the damage
How fast it starts glowing → intensity of the stress
How the signal declines → dynamics of the system under damage
Reversibility is not interpreted as a property of the damage itself, but as a designed feature of the biosensor, enabling repeated use under multiple damage cycles.
Circuit architecture
[ROS-sensitive promoter]
↓
[Fluorescent protein + degron]
↓
[Terminator]
Why this is non-trivial (and why it’s innovative)
Poor design choices lead to failure modes such as:
- Gel too dense → stimulus never reaches the circuit → no activation
- Gel too loose → excessive activation → no shutdown
- Reporter too stable → permanent signal → no reuse
- Circuit too sensitive → noise and false positives
PART 2. PROJECT CONSIDERATIONS
- Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals. Below is one example framework (developed in the context of synthetic genomics) you can choose to use or adapt, or you can develop your own. The example was developed to consider policy goals of ensuring safety and security, alongside other goals, like promoting constructive uses, but you could propose other goals for example, those relating to equity or autonomy.
Governance and Policy Considerations
flowchart TB
G["Governance & Policy Goals"]
G --> A["Non-malfeasance<br/>(Preventing Harm)"]
A --> A1["Cell-free TX–TL limits dual-use potential"]
A --> A2["Avoids human or clinical deployment"]
A --> A3["Environment friendly"]
G --> B["Safe and Responsible Research"]
B --> B1["Transparency in system limitations"]
B --> B2["Reproducibility and containment"]
B --> B3["Ensuring personal safety and capacitation"]
B --> B4["Financial responsability"]
G --> C["Constructive & Equitable Use"]
C --> C1["Accessibility of the platform"]
C --> C2["Supports education and interdisciplinary research"]
C --> C3["Promotion of heuristic rules/method"]
- Sub-Goal 1A. Cell-free TX–TL limits dual-use potential• The biosensor is designed as a cell-free system, preventing replication, evolution, or environmental persistence, thereby reducing biosafety and biosecurity risks.
- Sub-Goal 2A. Avoids human or clinical deployment• The system is not intended for in vivo, clinical, or diagnostic use; clear communication of this limitation helps prevent inappropriate application and fends emerging ethical concerns about animal and human clinical trials.
- Sub-Goal 3A. Environment friendly• This project prioritizes environmentally responsible design by relying on hydrogel matrices derived from biodegradable, bio-based, or naturally sourced polymers. Such materials are often obtained from renewable resources or industrial by-products, reducing environmental impact compared to synthetic, non-degradable sensing technologies. Additionally, the reusability of the biosensor minimizes material waste and lowers the frequency of disposal, contributing to a more sustainable experimental practice.
- Sub-Goal 1B. Transparency in system limitations• The biosensor reports oxidative damage via ROS signaling rather than direct radiation dose, and this distinction must be clearly stated to avoid misinterpretation.
- Sub-Goal 2B. Reproducibility and containment• The use of in silico circuit design and controlled TX–TL systems improves reproducibility while minimizing unintended biological interactions.
- Sub-Goal 3B. Ensuring personal welfare and capacitation• Because the system is intended for studying radiation-induced damage in controlled environments, its use must be accompanied by appropriate safety protocols and user training. This biosensor is explicitly not designed to replace personal dosimeters or occupational safety monitoring devices. Clear operational guidelines, radiation-handling protocols, and user capacitation are required to ensure that the biosensor is employed strictly as an experimental tool, without increasing risk to personnel.
- Sub-Goal 4B. Financial responsability• The proposed system emphasizes cost-effective design through the use of low-cost materials, minimal infrastructure requirements, and a reusable sensing strategy. By enabling multiple experimental cycles within the same biosensor material, the system reduces recurring expenses associated with single-use sensors or consumables. This extended operational lifetime represents a significant financial advantage for laboratories and institutions, supporting responsible allocation of economic resources.
- Sub-Goal 1C. Accessibility of the platform• Cell-free and hydrogel-based systems lower infrastructure barriers, making the platform more accessible to educational and research laboratories.
- Sub-Goal 2C. Supports education and interdisciplinary research• The project bridges synthetic biology, materials science, and medical physics while maintaining clear ethical boundaries around scope and use.
- Sub-Goal 3C. Promotion of heuristic rules• This project adopts a heuristic-driven design philosophy, leveraging simple, interpretable rules to guide system construction and experimentation. Material properties, circuit dynamics, and experimental steps are intentionally ordered to maximize efficiency—favoring low-cost, low-complexity processes early and reserving more resource-intensive steps for later stages. This approach improves time efficiency, reduces unnecessary expenditures, and promotes accessible, transferable design strategies that can be adapted across laboratories and disciplines.
PART 3. THE WHO AND THE HOW
- Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”). Try to outline a mix of actions (e.g. a new requirement/rule, incentive, or technical strategy) pursued by different “actors” (e.g. academic researchers, companies, federal regulators, law enforcement, etc). Draw upon your existing knowledge and a little additional digging, and feel free to use analogies to other domains (e.g. 3D printing, drones, financial systems, etc.).
Example
- Purpose: What is done now and what changes are you proposing?
- Design: What is needed to make it “work”? (including the actor(s) involved - who must opt-in, fund, approve, or implement, etc)
- Assumptions: What could you have wrong (incorrect assumptions, uncertainties)?
- Risks of Failure & “Success”: How might this fail, including any unintended consequences of the “success” of your proposed actions?
Governance Action 1 — Mandatory contextual labeling and use limitation
Actor(s): Academic researchers, research institutions, funding agencies.
Purpose
Currently, biosensors designed for radiation-related applications can be misinterpreted as direct radiation detectors or clinical tools. This project proposes a mandatory contextual labeling requirement stating that the system detects ROS-mediated damage, not radiation dose, and is intended strictly for in vitro experimental use. The change ensures that the tool is not misapplied in clinical, occupational, or regulatory contexts.
Design
To make this work, institutions and funding bodies would require that:
- All documentation, publications, and public-facing descriptions explicitly state the system’s scope and limitations.
- Experimental protocols include a standardized disclaimer clarifying that the biosensor does not replace dosimeters or personal safety devices.
- Course projects and academic demonstrations reinforce correct interpretation through documentation templates and reporting guidelines.
Assumptions
This action assumes that misinterpretation is a primary pathway for harm and that clear documentation meaningfully influences user behavior. It also assumes that researchers and students will comply with labeling norms when they are formally required.
Risks of Failure & “Success”
- Failure risk: Labels may be ignored, especially when the system performs well and appears “sensor-like.”
- Risk of success: If widely adopted, the tool could become a de facto standard for damage reporting, tempting users to extend it beyond its intended domain without appropriate validation.
Governance Action 2 — Safety training and protocol integration as a prerequisite for use
Actor(s): Research institutions, laboratory safety committees, instructors.
Purpose
Radiation-related experimentation already requires specialized training, but novel biosensors can create a false sense of safety. This action proposes that use of the biosensor be explicitly tied to existing radiation safety training and protocols, reinforcing that the tool supplements—but does not replace—established safety infrastructure.
Design
This action would require:
- Integration of the biosensor into institutional radiation safety manuals as an experimental reporting tool.
- Mandatory user training that explains what the biosensor measures, what it does not measure, and how to interpret its output.
- Oversight by institutional safety committees when the system is used in radiation-adjacent experiments.
Assumptions
This approach assumes that institutions already have safety frameworks capable of absorbing new tools, and that users are more likely to behave responsibly when a technology is embedded within formal safety structures.
Risks of Failure & “Success”
- Failure risk: Training could become procedural rather than substantive, reducing its effectiveness.
- Risk of success: If the biosensor becomes normalized within safety workflows, it may be incorrectly perceived as an authoritative indicator of safety rather than an experimental proxy.
Governance Action 3 — Incentivizing reusable, low-waste biosensing systems
Actor(s): Funding agencies, academic programs, sustainability-focused research initiatives.
Purpose
Many sensing technologies are single-use, expensive, or environmentally burdensome. This action proposes incentivizing reusable and low-waste biosensor designs, positioning reusability and material efficiency as desirable research outcomes rather than secondary considerations.
Design
This could be implemented through:
- Establish evaluation criteria that favor reusability, material sustainability, and life cycle efficiency.
- Creation and promotion of open, repositories that document reuse cycles, material performance, and design adaptations for biosensing platforms.
- Recognition or funding bonuses for designs that reduce consumables and experimental waste.
Assumptions
This action assumes that researchers respond to incentive structures and that sustainability metrics can be meaningfully evaluated without stifling innovation or creativity.
Risks of Failure & “Success”
- Failure risk: Incentives may encourage superficial reuse claims without rigorous validation.
- Risk of success: Strong emphasis on reuse could discourage exploration of necessary single-use or high-sensitivity designs in certain contexts.
PART 4. HOW WELL DO YOU DO?
- Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals.
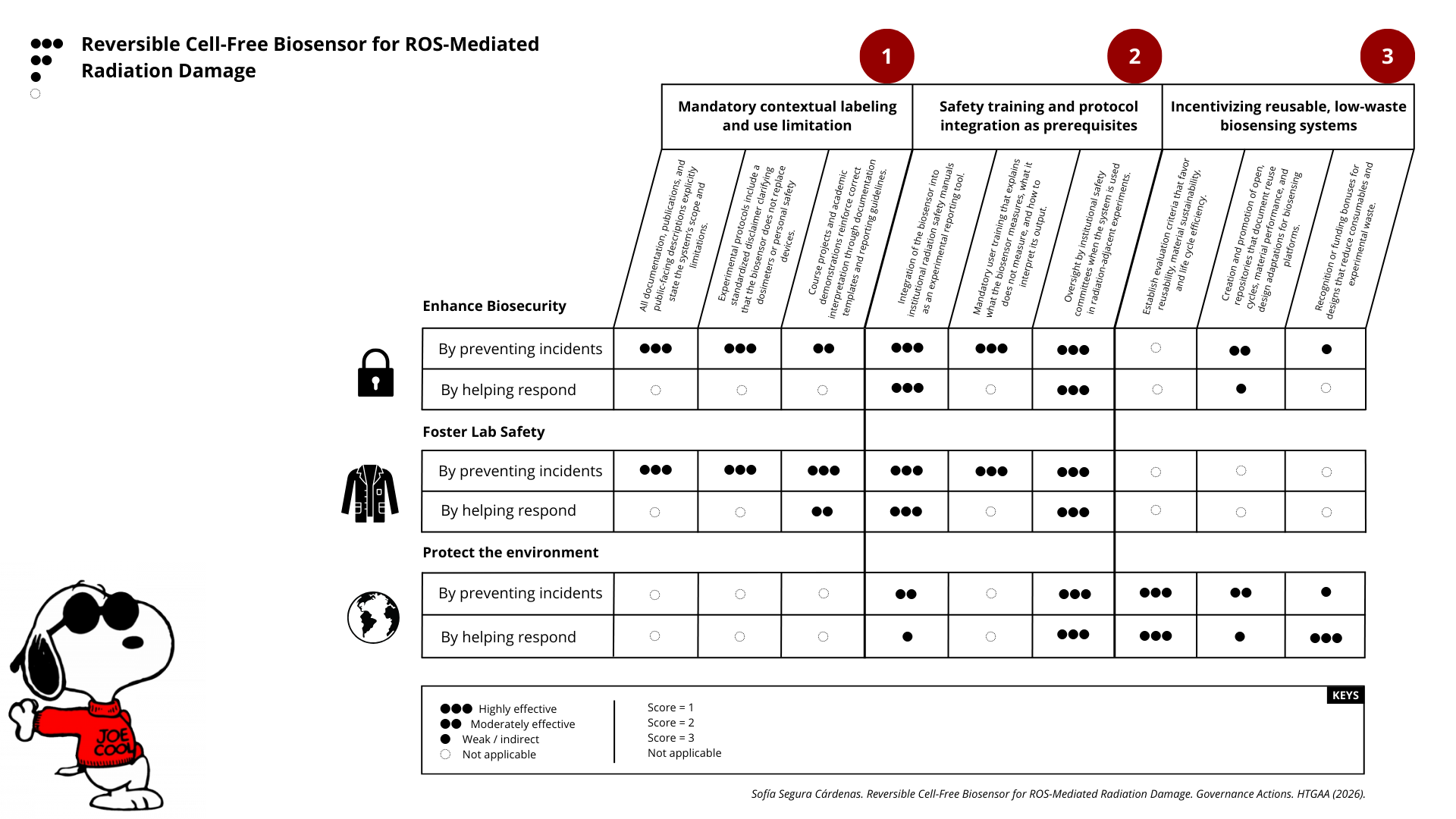

PART 5. PRIORITIES
- Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties. For this, you can choose one or more relevant audiences for your recommendation, which could range from the very local (e.g. to MIT leadership or Cambridge Mayoral Office) to the national (e.g. to President Biden or the head of a Federal Agency) to the international (e.g. to the United Nations Office of the Secretary-General, or the leadership of a multinational firm or industry consortia). These could also be one of the “actor” groups in your matrix.
Prioritized governance strategy and rationale
Drawing upon the governance scoring matrix, the most effective strategy for guiding the responsible development and use of the proposed reversible cell-free biosensor is a combined prioritization of Governance Options 1 and 2, with Governance Option 3 acting as a reinforcing, longer-term incentive mechanism.
Primary priority: Governance Options 1 and 2 (combined)
- Option 1 — Mandatory contextual labeling and use limitation +
- Option 2 — Safety training and protocol integration as prerequisites
These two options consistently score highest across biosafety, lab safety, and environmental protection, particularly in their ability to prevent incidents rather than merely respond to them. Together, they address the most immediate risks associated with misuse, misinterpretation, or inappropriate deployment of the biosensor.
Option 1 ensures that the system is clearly framed as:
- A cell-free, non-replicative biosensing platform
- Not a personal radiation dosimeter Not intended for clinical or in vivo use
This directly reduces the risk of over-interpretation of fluorescence signals and prevents the technology from being deployed outside its validated scope.
Option 2 complements this by embedding the biosensor within existing institutional safety cultures, requiring that users receive appropriate training in:
- Radiation handling protocols
- Interpretation of indirect ROS-based signals
- Limitations of TX–TL systems
Importantly, this option does not introduce new regulatory burdens but instead leverages existing laboratory training and approval workflows, making it both feasible and scalable.
Trade-off considered: These measures may slow early adoption or increase onboarding time for new users. However, this is outweighed by the reduction in misuse risk and the preservation of trust in the technology.
Secondary priority: Governance Option 3 (Incentive-based einforcement)
Option 3 — Incentivizing reusable, low-waste biosensing systems
While Option 3 scores lower in immediate incident prevention, it plays a crucial role in shaping long-term research behavior and system design choices. Incentives that reward reusability, lifecycle efficiency, and reduced consumables encourage adoption of the very properties that distinguish this biosensor from traditional single-use sensors.
Rather than acting as a primary safeguard, this option functions best as:
- A structural reinforcement mechanism +
- A signal to researchers and institutions that sustainability and reuse are valued outcomes
Trade-off considered:
Incentive-based mechanisms depend on institutional uptake and may have uneven effects across well-funded versus resource-limited laboratories. Their impact is therefore slower and less uniform than mandatory requirements.
Assumptions and Uncertainties
This prioritization assumes that:
- Institutions and laboratories already possess baseline safety infrastructure
- Users are willing to engage with training and labeling requirements
- Regulatory bodies are receptive to non-single-use technologies
Uncertainties remain regarding:
- How fluorescence-based damage reporting might be interpreted by non-experts
- Variability in institutional enforcement of training standards
- How incentive structures translate into real design decisions over time
Recommended audience
This governance strategy is primarily directed toward:
- Institutional biosafety committees and laboratory leadership
- Funding agencies and regulatory bodies overseeing research infrastructure
- Organizations setting best-practice standards for cell-free and biosensing technologies
By acting at this institutional and regulatory level, the proposed governance combination balances safety, feasibility, innovation, and sustainability, aligning closely with the technical and ethical goals of the project.
WEEK 2 - LECTURE PREP
In preparation for Week 2’s lecture on “DNA Read, Write, and Edit," please review the follow materials
Lecture 2 slides as posted below.
The associated papers that are referenced in those slides.
In addition, answer these questions in each faculty member’s section:
Homework Questions from Professor Jacobson: [Lecture 2 slides]
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
DNA Polymerase error rate and genome fidelity
DNA polymerase, the enzyme responsible for copying DNA during replication, has an intrinsic error rate of approximately 1 mistake per 10⁵ nucleotides incorporated.
The human genome contains about 3 × 10⁹ base pairs. At this raw error rate, tens of thousands of mutations would occur every time a human cell divides, which would be incompatible with life.
How biology addresses this discrepancy
Biological systems reduce replication errors through multiple layers of error correction:
Proofreading by DNA polymerase
Many DNA polymerases possess 3′→5′ exonuclease activity, which allows them to remove incorrectly incorporated nucleotides immediately. This improves fidelity to roughly 1 error per 10⁷ nucleotides.Post-replication mismatch repair (MMR)
Additional cellular repair systems detect and correct mismatches that escape proofreading, further reducing the error rate to approximately 1 error per 10⁹–10¹⁰ nucleotides.
As a result, the final error rate is low enough that most cell divisions occur without introducing harmful mutations.
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
Coding Capacity of DNA for an Average Human Protein
An average human protein is approximately 300 amino acids long. Each amino acid is encoded by a codon, a sequence of three nucleotides.
Because there are 4 possible nucleotides (A, T, C, G), there are:
- ( 4^3 = 64 ) possible codons
- Only 20 amino acids (plus stop signals)
This means the genetic code is degenerate, and most amino acids are encoded by multiple codons.
Number of Possible DNA Sequences for One Protein
If an average amino acid is encoded by ~3 synonymous codons, then the total number of possible DNA sequences that could encode a 300–amino acid protein is approximately:
[ 3^{300} ]
This is an astronomically large number, meaning there are many distinct DNA sequences that can, in theory, encode the same protein.
Why Most Possible Codes Do Not Work in Practice
Despite this theoretical flexibility, not all synonymous DNA sequences function equally well due to several biological constraints:
Codon usage bias
Organisms preferentially use certain codons over others. Rare codons can slow translation or cause ribosome stalling.mRNA secondary structure
Certain nucleotide sequences form stable secondary structures that hinder ribosome binding or elongation.Translational accuracy and efficiency
Codon choice can affect misincorporation rates and protein folding during translation.Regulatory elements embedded in coding sequences
Coding regions may overlap with regulatory signals affecting splicing, mRNA stability, or localization.GC content and genome stability
Extreme nucleotide compositions can impact DNA replication and transcription efficiency.
Because of these factors, only a small subset of all theoretically possible DNA sequences are biologically viable for producing a functional protein at appropriate levels.
Homework Questions from Dr. LeProust: [Lecture 2 slides]
What’s the most commonly used method for oligo synthesis currently?
The most commonly used method for oligonucleotide synthesis is solid-phase phosphoramidite chemistry.
In this method, DNA is synthesized stepwise from the 3′ to the 5′ end on a solid support. Each cycle consists of four main steps: deprotection, coupling of a phosphoramidite nucleotide, capping of unreacted chains, and oxidation. This approach is highly automated, fast, and reliable, making it the standard technique used by commercial DNA synthesis providers.
Why is it difficult to make oligos longer than 200nt via direct synthesis?
It is difficult to synthesize oligos longer than ~200 nucleotides because errors accumulate with each synthesis cycle.
Each nucleotide addition has a small but nonzero failure rate (incomplete coupling, side reactions, or deletions). As the oligo length increases, these errors compound exponentially, leading to a low fraction of full-length, correct sequences. Additionally, longer oligos are harder to purify effectively, since truncated products differ only slightly in length from the desired product.
Why can’t you make a 2000bp gene via direct oligo synthesis?
A 2000 bp gene cannot be made via direct oligo synthesis because the cumulative error rate would be extremely high, resulting in an almost negligible yield of error-free full-length DNA.
Beyond error accumulation, chemical synthesis efficiency, purification limitations, and cost make direct synthesis impractical at this scale. Instead, long genes are constructed by assembling shorter, overlapping oligos using enzymatic methods such as PCR-based assembly or Gibson assembly, followed by cloning and sequence verification.
Homework Question from George Church: [Lecture 2 slides]
Choose ONE of the following three questions to answer; and please cite AI prompts or paper citations used, if any.
[Using Google & Prof. Church’s slide #4] What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
The 10 essential amino acids in animals are those that cannot be synthesized de novo and therefore must be obtained from the diet:
- Histidine
- Isoleucine
- Leucine
- Lysine
- Methionine
- Phenylalanine
- Threonine
- Tryptophan
- Valine
- Arginine (essential in all animals during growth; in many adult animals it is conditionally essential)
How does this affect the view of the “Lysine Contingency”?
The “Lysine Contingency” refers to the idea that life—particularly animals—became evolutionarily dependent on lysine availability from external sources, because animals lost the ability to synthesize lysine. Since lysine is universally essential in animals and often limiting in plant-based diets (especially cereal grains), this creates a strong nutritional and evolutionary constraint.
This reinforces the view that the lysine contingency is real and biologically significant:
- Animals are metabolically constrained by the loss of lysine biosynthesis pathways.
- Ecosystems and food webs are shaped by lysine availability and by organisms (plants, fungi, bacteria) that can synthesize it.
- It helps explain why lysine supplementation or biofortification (e.g., high-lysine crops) has a major impact on nutrition and health.
Overall, the universality of lysine as an essential amino acid in animals supports the idea that lysine availability is a key evolutionary and nutritional bottleneck rather than a trivial dietary detail.
[Given slides #2 & 4 (AA:NA and NA:NA codes)] What code would you suggest for AA:AA interactions?
[(Advanced students)] Given the one paragraph abstracts for these real 2026 grant programs sketch a response to one of them or devise one of your own:
https://arpa-h.gov/explore-funding/programs/boss
https://www.darpa.mil/research/programs/smart-rbc
https://www.darpa.mil/research/programs/go
Resources
- Nguyen, P. Q., Soenksen, L. R., Donghia, N. M., Angenent-Mari, N. M., de Puig, H., Huang, A., Lee, R. A., Slomovic, S., Galbersanini, T., Lansberry, G., Sallum, H. M., Zhao, E. M., Niemi, J. B. & Collins, J. J. (2021). Wearable materials with embedded synthetic biology sensors for biomolecule detection. Nature Biotechnology, 39(11). https://doi.org/10.1038/s41587-021-00950-3
- Karim, M. M. and Lasker, T. (2025). Electrochemical Biosensors for Cancer Biomarker Detection: Basic Concept, Design Strategy and Cutting‐Edge Development. Electrochemical Science Advances. https://doi.org/10.1002/elsa.70007
- Liang, Q., Lu, Y. & Zhang, Q. (2022). Hydrogels‐Based Electronic Devices for Biosensing Applications. In Smart Stimuli-Responsive Polymers, Films, and Gels. https://doi.org/10.1002/9783527832385.ch10
- Zhang, M., Xu, T., Liu, K., Zhu, L., Miao, C., Chen, T., Gao, M., Wang, J. & Si, C. (2024). Modulation and Mechanisms of Cellulose‐Based Hydrogels for Flexible Sensors. SusMat, 5. https://doi.org/10.1002/sus2.255
- Ahmed, S. N. (2015). Physics and Engineering of Radiation Detection. Choice Reviews Online. https://doi.org/10.1016/C2013-0-15270-1
- Ng, K., Ung, N. & Hill, R. (2022). Problems and Solutions in Medical Physics: Radiotherapy Physics. CRC Press. https://doi.org/10.1201/9780429159466
- Abaza, A. M. H. (2017). New Trend in Radiation Dosimeters. American Journal of Modern Physics, 7(1), 21-30. https://doi.org/10.11648/j.ajmp.20180701.13
- Bartoloni, A. & Strigari, L. (2025). Space Radiobiology: Synergies between Astroparticle and Medical Physics. World Scientific.
- Zhang, J., Liu, J., Qiao, L., Zhang, Q., Hu, J. & Zhang, C. (2024). Recent Advance in Single-Molecule Fluorescent Biosensors for Tumor Biomarker Detection. Biosensors, 14(11). https://doi.org/10.3390/bios14110540
- Cao, X., Lv, D., Zhang, L & Xing, Z. (2020). Adaptive Governance, Loose Coupling, Forward-Looking Strategies and Responsible Innovation (September 2020). https://doi.org/10.1109/ACCESS.2020.3046095
- OpenAI. (2026). ChatGPT (GPT-5.2) [Large language model]. https://chat.openai.com/


