HTGAA - Week 7: Genetic Circuits Part II: Neuromorphic Circuits


My Homework
WEEK 7 - SMART COMPUTATIONAL SOLUTIONS
This week covers neuromorphic genetic circuits, showing how engineered gene networks can implement neural-network “perceptron”-like computation and learning.
Lecture (Tues, Mar 17)
Genetic Circuits Part II: Neuromorphic Circuits
(▶️Recording)
Ron Weiss
Recitation (Wed, Mar 18)
Neuromorphic circuits & Biomaterials
(▶️Recording |
💻Slides)
Evan Holbrook, Ren Ramlan
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
Assignees for the following sections
| MIT/Harvard students | Required |
| Committed Listeners | Required |
1. What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
About IANNs
Intracellular Artificial Neural Networks (IANNs) are engineered gene networks inside living cells that mimic the behavior of artificial neural networks, particularly simple models like perceptrons.
In a traditional artificial neural network, you have inputs, weights, a summation step, and an activation function. IANNs recreate these same components using biological parts:
- Inputs → concentrations of molecules (transcription factors, small molecules, or signals)
- Weights → regulatory strengths (such as promoter strength, ribosome binding sites, or binding affinities between regulators and DNA)
- Summation → combined effect of multiple regulators acting on a promoter
- Activation function → nonlinear gene expression response (e.g., sigmoidal response of transcription)
So instead of silicon-based computation, the “computation” happens through gene expression and molecular interactions inside the cell.
What makes IANNs especially interesting is that they allow cells to perform analog, multi-input decision-making, rather than simple Boolean logic. For example, a cell could integrate several environmental signals and produce an output only if a weighted combination of those signals crosses a threshold—just like a perceptron classifying data. Additionally, some IANN designs incorporate mechanisms for tuning or learning, where the effective “weights” can be adjusted (for example, by modifying gene expression levels or regulatory interactions), allowing the system to adapt to new conditions.
In short, IANNs are a bridge between:
- Synthetic biology (engineering gene circuits)
- Machine learning concepts (like neural networks and learning)
Enabling living cells to carry out more sophisticated computations such as pattern recognition, classification, and adaptive responses.
Advantages
IANNs can process continuous (analog) inputs rather than being limited to binary ON/OFF states. Traditional Boolean circuits treat signals as discrete, which restricts the complexity of responses. In contrast, IANNs allow graded responses to varying concentrations of molecules, making them more biologically realistic.
IANNs enable integration of multiple inputs in a weighted manner. Instead of simple logical operations like AND or OR, they can assign different “weights” to each input, allowing more nuanced decision-making—similar to how perceptrons work in artificial neural networks.
They provide greater computational complexity and flexibility. Boolean circuits scale poorly when trying to implement complex behaviors, often requiring many layers and components. IANNs can implement sophisticated functions (like classification or pattern recognition) more efficiently within a single network.
IANNs are capable of learning and adaptability. While traditional genetic circuits are typically static once designed, IANNs can, in principle, be engineered to adjust their parameters (like weights) in response to environmental signals, enabling adaptive behavior.
Also, IANNs mimic natural cellular decision-making processes better, which are rarely purely binary. This makes them especially useful for applications in synthetic biology where cells must respond to complex, noisy, and dynamic environments.
Overall, IANNs expand the capabilities of synthetic gene networks from simple logical operations to more powerful, flexible, and biologically relevant computation.
2. Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
A) Conceptual idealistic application: IANN-Based Metabolic Optimization Under Space Stress
Concept
An Intracellular Artificial Neural Network (IANN) can be engineered to function as a closed-loop metabolic controller, allowing cells to dynamically redistribute metabolic resources under extreme conditions (like microgravity, radiation, or nutrient limitations in space) encountered in Space Medicine.
Instead of relying on fixed metabolic pathways, the system continuously evaluates the internal physiological state of the cell and adjusts metabolic fluxes to maintain homeostasis and viability.
Instead of fixed pathways, the system does: “Given my current stress and resources, how should I reroute metabolism?”
| Inputs (continuous intracellular signals) | Processing (IANN computation) |
|---|---|
| The IANN integrates multiple metabolic and stress-related variables, such as: • ATP/ADP ratio (cellular energy status) • NADH/NAD⁺ ratio (redox balance) • Reactive oxygen species (ROS) levels • Availability of key nutrients (e.g., glucose, amino acids) These inputs are inherently continuous and noisy, making them well-suited for neural-like computation rather than Boolean logic. | Each input is assigned a regulatory weight based on its impact on cellular fitness. The IANN performs a weighted integration of these signals and applies a nonlinear activation function to classify the overall metabolic state, for example: • Energy-efficient state • Oxidative stress state • Nutrient-limited state This enables the system to interpret complex intracellular conditions as distinct physiological regimes. |
| Outputs (active metabolic control) | Usefulness |
| Based on this classification, the IANN actively regulates cellular metabolism by: • Upregulating or downregulating key metabolic enzymes • Redirecting flux between pathways (glycolysis vs. oxidative phosphorylation) • Activating antioxidant and stress-response pathways Unlike a biosensor, the system directly modifies cellular behavior, forming a true closed-loop control system. | In space environments, cells experience combined stresses such as radiation, microgravity, and limited nutrients. These conditions disrupt metabolic balance. An IANN-based controller allows cells to: • Adapt in real time to fluctuating conditions • Maintain energy and redox balance • Improve survival and functional stability |
| Limitations and Challenges | Why IANNs are necessary |
| • Complexity of metabolic networks: Metabolic pathways are highly interconnected, making precise control difficult. • Weight tuning and calibration: Determining appropriate regulatory weights is challenging and may vary across conditions. • Cellular burden: Implementing complex circuits may reduce overall cellular efficiency. • Dynamic instability: Feedback regulation could lead to oscillations or unintended metabolic states. | Traditional genetic circuits based on Boolean logic cannot effectively handle: • Continuous metabolic variables • Multi-factorial regulation • Dynamic adaptation IANNs enable graded, multi-input decision-making, making them uniquely suited for real-time metabolic control. |
This application highlights how IANNs can transform engineered cells from passive sensors into adaptive metabolic systems capable of maintaining homeostasis under extreme and unpredictable environments.
B) Tested real ans successful application:
Concept
A machine learning system (Random Forest) applied to continuous acoustic emission data from a laboratory fault. The system learns to predict the time remaining before failure (lab earthquake) by identifying subtle patterns in the signal that are not detectable by humans. Unlike traditional methods based on recurrence intervals, this approach uses instantaneous physical signal features and reveals previously overlooked precursors hidden in what was thought to be noise.
| Parameters | Description |
|---|---|
| Inputs (continuous physical signals) | The model analyzes continuous acoustic emission (AE) signals generated by the fault. From these signals, it extracts statistical features such as:
These inputs are continuous, noisy, and high-dimensional, making them ideal for machine learning rather than classical threshold-based analysis. |
| Processing (ML computation) | The system uses a Random Forest algorithm, which consists of multiple decision trees. Each tree evaluates the statistical features extracted from short time windows of the acoustic signal. The model:
|
| Outputs (prediction / system response) | The system outputs a continuous prediction of time remaining before the next failure event. Key characteristics:
This demonstrates that the system continuously tracks the progression toward failure. |
| Usefulness | This approach overcomes limitations of traditional earthquake prediction methods by:
|
| Key scientific insight | The study reveals that low-amplitude acoustic signals—previously ignored—contain critical information about the system’s state. These signals likely originate from continuous grain motion in the fault gouge, reflecting gradual stress accumulation. The system shows that failure is not sudden, but a progressive, measurable process. |
| Limitations and Challenges |
|
This is a real demonstration of a system that:
- Integrates continuous noisy signals
- Performs nonlinear multi-parameter analysis
- Produces real-time predictions
- This system is computational (external ML model)
- IANNs would implement similar logic inside living cells
3. Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.
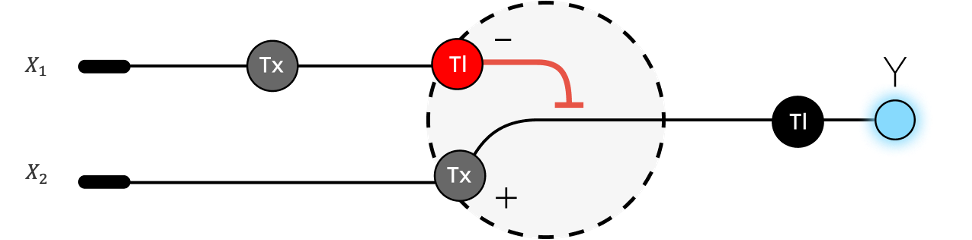

The diagram represents an intracellular single-layer perceptron, where biological components mimic a simple neural network:
- Inputs are encoded as DNA sequences
- Processing occurs through gene expression (transcription + translation)
- Output is a measurable protein signal
System Breakdown
Inputs
- X₁ (DNA → Csy4 endoribonuclease): Encodes an enzyme capable of cleaving RNA.
- X₂ (DNA → fluorescent protein): Encodes the output protein, but its expression is regulated post-transcriptionally.
Processing (Tx + Tl)
- Tx (Transcription): DNA → mRNA
- Tl (Translation): mRNA → Protein
👉 X₁ produces Csy4 enzyme
👉 X₂ produces fluorescent protein mRNA
Regulation Mechanism
- The Csy4 endoribonuclease recognizes and cleaves specific sequences in the mRNA of the fluorescent protein.
- This acts like a negative weight (inhibition)
Output
Final fluorescence depends on:
- Presence of X₂ (can produce protein)
- Presence of X₁ (can suppress it)
Layer 1 flux
- X₂ = main signal
- X₁ = regulator (weight)
- Output = activation level
👉 This is equivalent to a single neuron with weighted inputs. The system does not just detect signals—it computes a weighted response through molecular interactions.
Now, we draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
The original system is limited because:
- Regulation happens in a single step
- No intermediate processing exists
Therefore: A multilayer intracellular perceptron is proposed, where biological regulators act as hidden nodes enabling hierarchical computation.
System flux
Layer 1 — Signal Integration
Goal: Convert multiple inputs into a regulatory signal
Inputs: X₁, X₂ = DNA-encoded signals
Process: Transcription + translation produce an endoribonuclease (E1)
This layer performs:
👉 signal integration
👉 feature extraction
Hidden Node (E1)
- E1 acts as a biological neuron output
- Encodes processed information from Layer 1
Can:
- cleave RNA
- modulate stability
- control translation efficiency
Equivalent to: hidden layer activation in neural networks
Layer 2 — Output Generation
Goal: Transform regulatory signal into measurable output
Input: E1 (from Layer 1)
Process: E1 regulates fluorescent protein mRNA
Output: Fluorescent protein level
The new design allows:
- Nonlinear responses
- Signal filtering
- Amplification or suppression
- Temporal dynamics (if extended)
By introducing an intermediate regulatory layer, intracellular circuits can implement hierarchical computation, where molecular species act as hidden nodes transforming input signals before generating an output response.
Sketch
| Neural Network | Biológico |
|---|---|
| Neuron | Gene circuit |
| Weight | Strength of regulation (cleavage efficiency) |
| Activation | Protein expression level |
| Hidden layer | Regulatory molecule (endoribonuclease) |
The multilayer intracellular perceptron enables hierarchical signal processing, where intermediate biomolecular regulators act as hidden nodes transforming input signals into controlled gene expression outputs.
Assignment Part 2: Fungal Materials
Assignees for the following sections
| MIT/Harvard students | Required |
| Committed Listeners | Required |
1. What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Fungal materials are bio-based composites made primarily from mycelium, the root-like network of fungi. The mycelium acts as a natural binder, growing through agricultural waste (e.g., straw, husks) and forming a solid structure.
Examples of existing fungal materials
| Material | Example | Uses | Advantages | Disadvantages |
|---|---|---|---|---|
| Mycelium-Based Packaging | Ecovative Design | Replacement for polystyrene; protective packaging for electronics and furniture | Biodegradable (compostable in weeks); renewable raw materials; low energy production | Lower durability than plastics; sensitive to moisture; limited shelf life |
| Mycelium Composites (Furniture & Panels) | MycoWorks; mycelium boards in architecture | Furniture (chairs, tables); interior panels; lightweight structural components | Lightweight; good compressive strength; moldable into custom shapes | Lower tensile strength than wood/metal; not suitable for load-bearing structures; variability in properties |
| Mycelium Leather (Biofabricated Leather) | Bolt Threads (Mylo™); MycoWorks (Reishi™) | Fashion (bags, shoes, clothing); upholstery | Animal-free; lower environmental impact than leather; customizable texture and thickness | Scaling challenges; durability not yet equal to high-grade leather; can be expensive |
| Mycelium Bricks & Construction Materials | Experimental architectural applications | Insulation panels; temporary structures; sustainable construction | Fire-resistant; good thermal and acoustic insulation; very low carbon footprint | Low structural strength; not standardized for building codes; degradation over time |
| Mycelium Foams | — | Insulation; shock absorption; acoustic dampening | Replaces petroleum-based foams; biodegradable; non-toxic | Less uniform than synthetic foams; performance variability |
Comparison with traditional materials
| Property | Fungal Materials | Traditional Materials |
|---|---|---|
| Source | Renewable (biological) | Fossil-based / mined |
| Biodegradability | High | Low (plastics persist) |
| Energy use | Low (grown, not manufactured) | High (industrial processing) |
| Mechanical strength | Moderate–low | High |
| Durability | Limited | High |
| Environmental impact | Low | High (CO₂, pollution) |
Key avantages
Sustainability
- Grown from waste
- Compostable
- Circular economy compatible
Energy Efficiency
- No high-temperature processing
- Self-assembling material
Design Flexibility
- Can grow into molds
- Tunable properties via growth conditions
Key limitations
- Mechanical Constraints: Not as strong as steel, concrete, or advanced polymers
- Environmental Sensitivity: Moisture and biological degradation
- Standardization Issues
- Variability between batches
- Hard to scale consistently
2. What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Environmental Context – Mexico Oil Spill (YeT aGaIN! 🙃🫠)
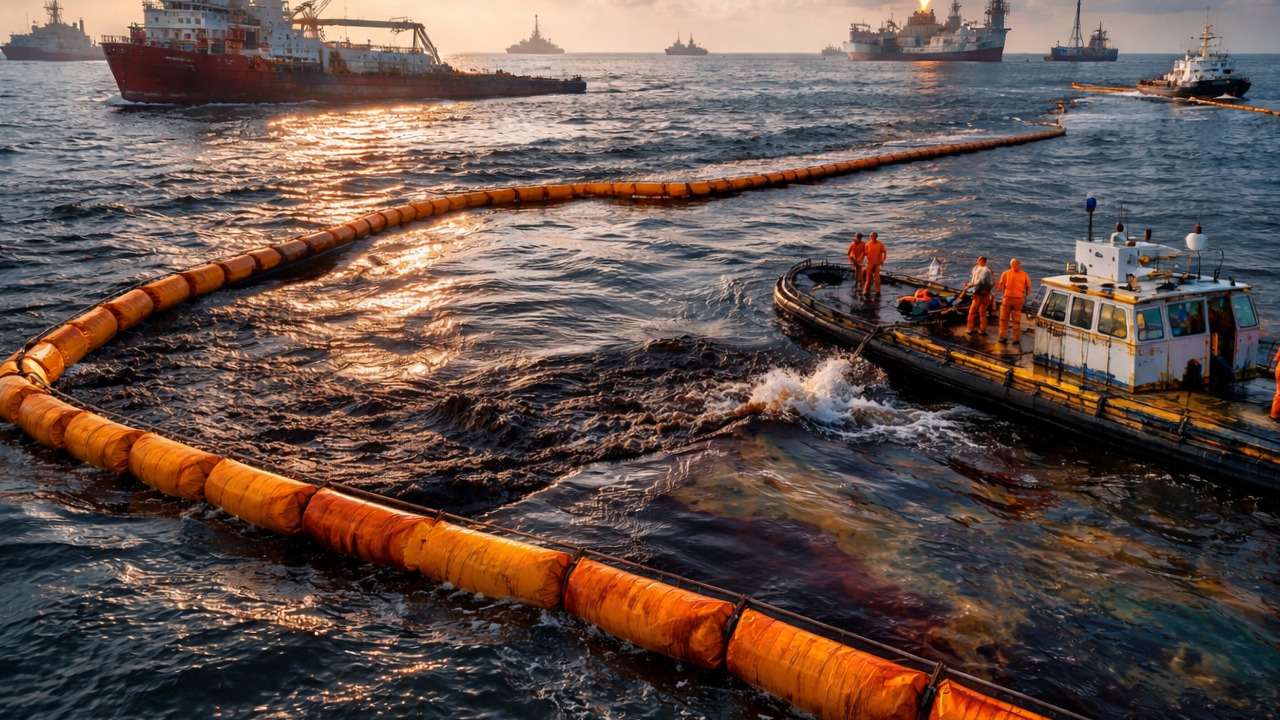
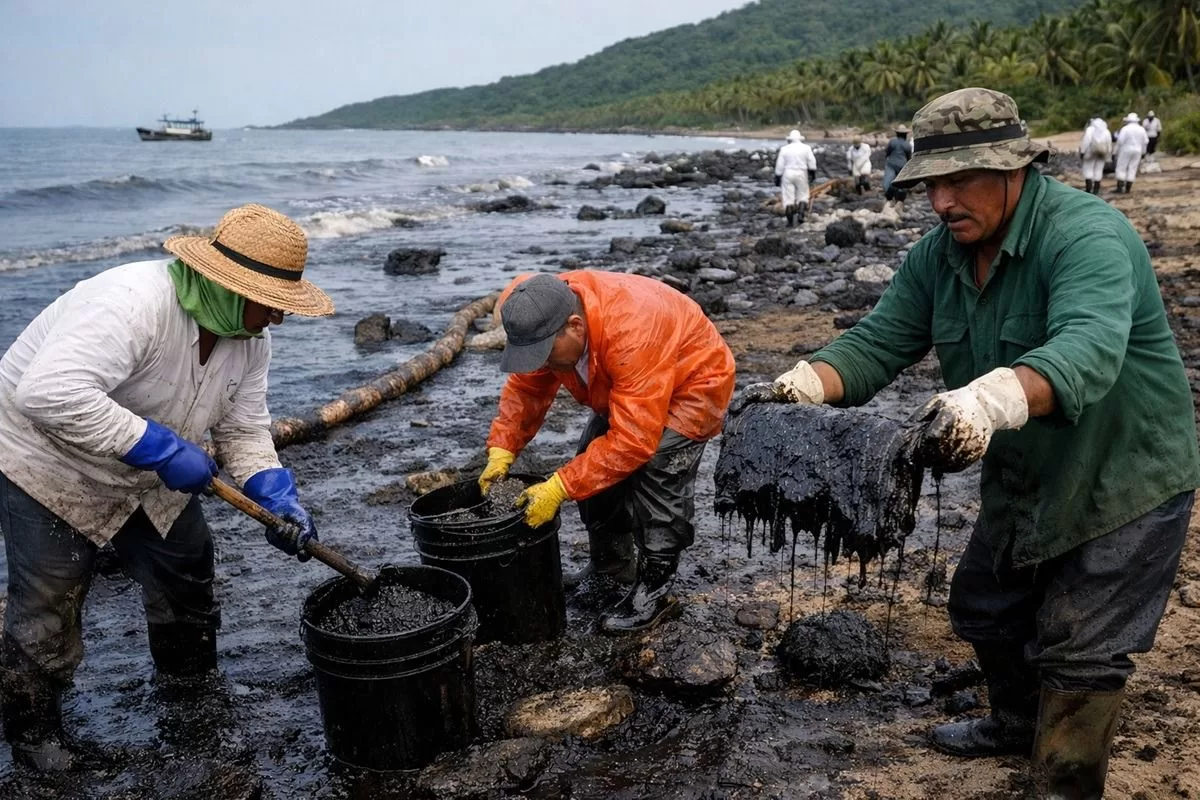
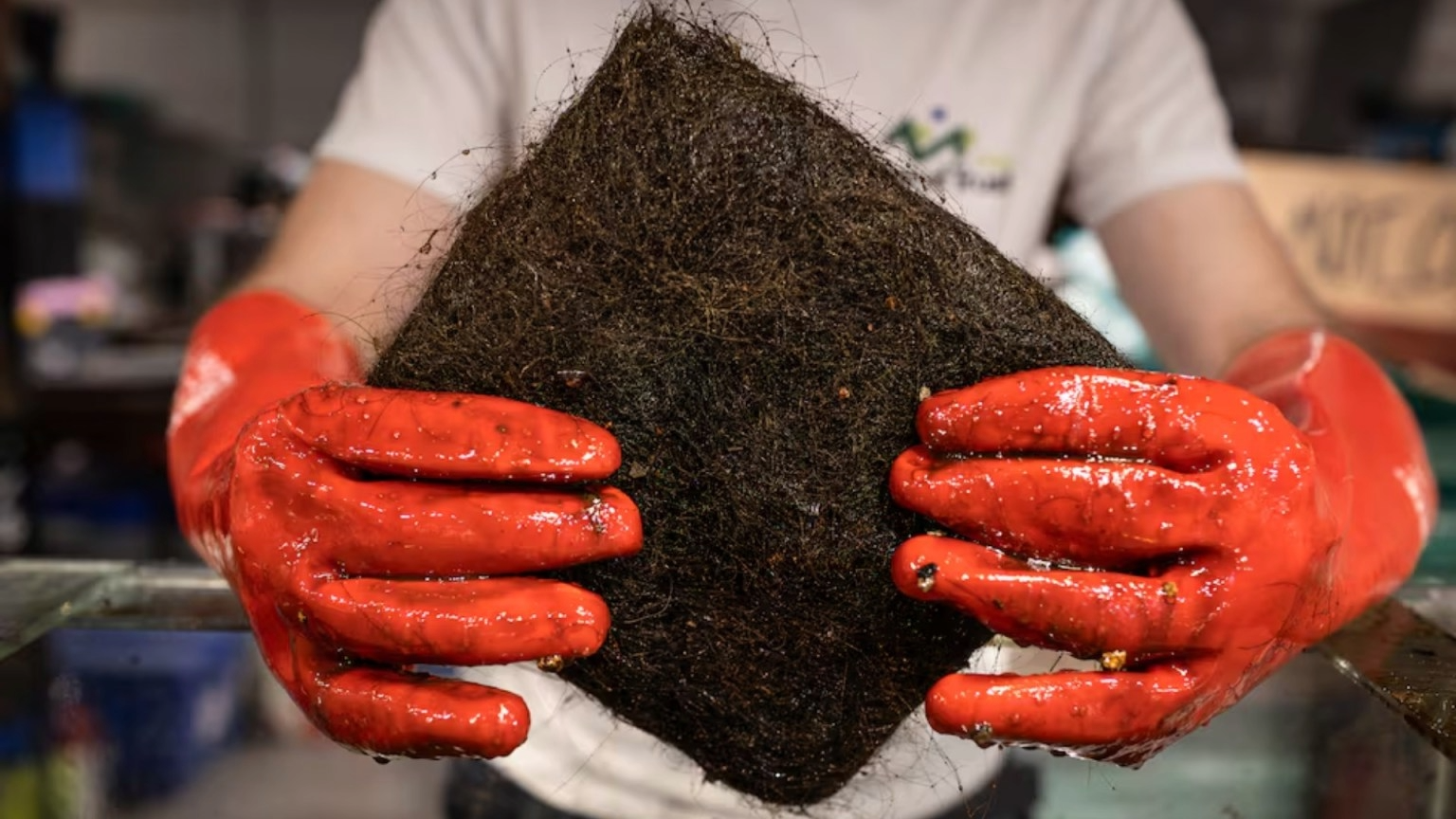
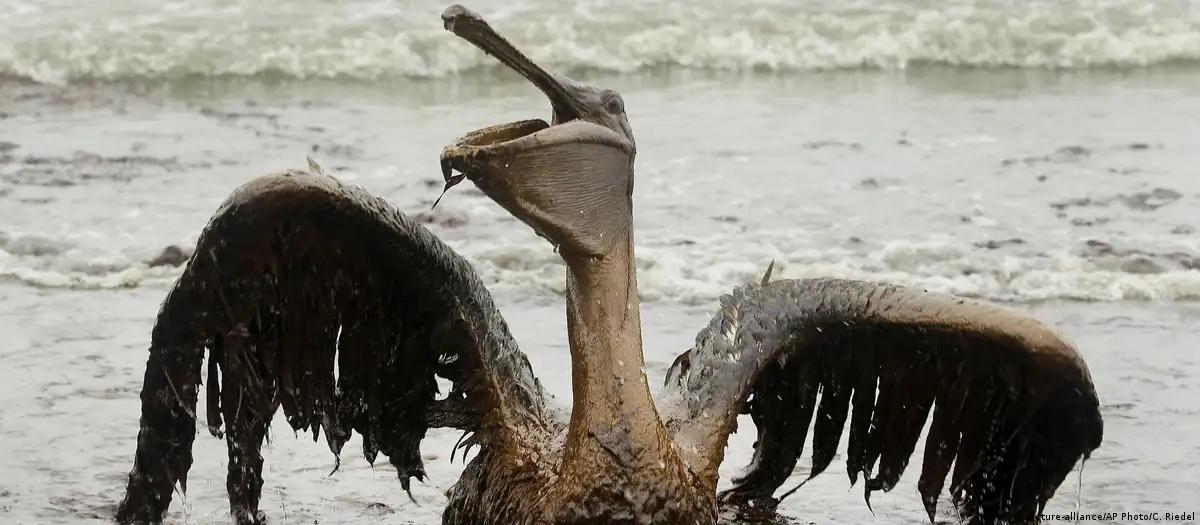
Recent petroleum spills in my cute little Mexico have once again highlighted the vulnerability of marine and coastal ecosystems to hydrocarbon contamination. These events have led to severe environmental consequences, including water pollution, damage to marine habitats, and visible distress and mortality in wildlife. In response, local communities and volunteers around the country have mobilized grassroots efforts, such as collecting human and animal hair to create absorbent barriers for oil cleanup. While these initiatives demonstrate remarkable social engagement, they also underscore the limitations of current response strategies, which are often reactive, labor-intensive, and insufficient for large-scale remediation. This situation emphasizes the urgent need for innovative, scalable, and biologically driven solutions. Engineering fungi capable of degrading petroleum compounds offers a promising approach, as such systems could actively break down pollutants in situ, complementing physical cleanup efforts and contributing to faster and more sustainable ecosystem recovery.
| Environmental damage |
|---|

 |

 |

 |

 |
Engineering Fungi for Oil Spill Remediation
The proposal is to genetically engineer fungi to detect, absorb, and degrade petroleum hydrocarbons in contaminated marine and coastal environments.
Fungi would be designed to:
- Secrete hydrocarbon-degrading enzymes, such as laccases and peroxidases, to break down complex petroleum compounds into less toxic molecules.
- Enhance their mycelial network structure to physically trap and retain oil, similar to how materials like human hair are currently used in cleanup efforts.
- Sense the presence of hydrocarbons and upregulate degradation pathways only when pollutants are detected, improving efficiency and reducing unnecessary metabolic burden.
The motivation for this approach comes from the urgent need for more effective and scalable responses to oil spills. Current methods are often limited to physical removal or absorption, which do not fully eliminate contaminants. In contrast, engineered fungi could provide a self-sustaining, in situ bioremediation system that not only contains but actively neutralizes pollutants, accelerating ecosystem recovery.
Conceptual system overview
Why use fungi instead of bacteria?
Advantages of Fungi
1. Physical Structure (Mycelium Networks)
Fungi form extensive filamentous networks that can:
- Penetrate contaminated sediments and shorelines
- Physically trap oil particles
- Cover large surface areas
This makes them both a material and a metabolic system, unlike bacteria.
2. Ability to Degrade Complex Compounds
Fungi naturally degrade highly complex organic materials such as lignin, which is structurally similar to many petroleum compounds. This gives them a strong advantage in breaking down recalcitrant hydrocarbons.
3. Environmental Robustness
Fungi can survive in:
- Low oxygen environments
- Nutrient-poor conditions
- Harsh and variable ecosystems
Ideal for real-world spill conditions.
4. In Situ Growth and Self-Propagation
Once deployed, fungal mycelium can:
- Expand across contaminated areas
- Continuously produce degrading enzymes
- Self-repair and persist over time
5. Potential for Integrated Sensing and Response
Fungi can be engineered with regulatory circuits to:
- Detect pollutants
- Dynamically adjust enzyme production
Enabling smart, adaptive bioremediation systems
Limitations compared to bacteria
- Slower growth rates
- More complex genetic engineering
- Fewer standardized tools and models
- Greater difficulty in precise control
Assignment Part 3: First DNA Twist Order
Assignees for the following sections
| MIT/Harvard students | Required |
| Committed Listeners | Required |
- Review the Individual Final Project documentation guidelines.
- Submit this Google Form with your draft Aim 1, final project summary, HTGAA industry council selections, and shared folder for DNA designs. DUE MARCH 20 FOR MIT/HARVARD/WELLESLEY STUDENTS
- Review Part 3: DNA Design Challenge of the week 2 homework. Design at least 1 insert sequence and place it into the Benchling/Kernel/Other folder you shared in the Google Form above. Document the backbone vector it will be synthesized in on your website.
Resources
- The perceptron, the basis of artificial neural networks: https://www.geeksforgeeks.org/deep-learning/what-is-perceptron-the-simplest-artificial-neural-network/
- Many examples of artificial neural networks made using biomolecules: https://doi.org/10.1016/j.biosystems.2024.105164
- Montesinos López O. A., Montesinos López A., & Crossa, J. Chapter 10: Fundamentals of Artificial Neural Networks and Deep Learning. Multivariate Statistical Machine Learning Methods for Genomic Prediction. (2022). Springer. https://www.ncbi.nlm.nih.gov/books/NBK583971/
- Nilsson, A., Peters, J. M., Meimetis, N., et al. (2022). Artificial neural networks enable genome-scale simulations of intracellular signaling. Massachusetts Institute of Technology. Nature Communications, 13 (1). https://hdl.handle.net/1721.1/147780
- Rouet-Leduc, B., Hulbert, C.,Lubbers, N., et al. (2017). Machine learningpredicts laboratory earth-quakes. Geophysical Research Letters, 44, 9276–9282. https://doi.org/10.1002/2017GL074677
- OpenAI. (2026). ChatGPT (GPT-5.2) [Large language model]. https://chat.openai.com/