HTGAA - Week 11: Bioproduction & Cloud Labs


Cloud laboratories are making science accessible, affordable, and reproducible. Our aim this semester is to showcase how they can enable human creativity at scale, and how they provide a platform for collaboration and community.
How To Grow (Almost) Anything is about synthetic biology, bioengineering, robotics, automation, art, and AI. But it is also about friendship, shared purpose, and the freedom to build beyond what we know and to be inspired by what can be. To that end, the goal with this cloud lab unit and homework assignment is to inspire collaboration and creativity while designing a scientifically rigorous cell-free fluorescent protein optimization experiment together.
Lecture (Tues, Apr 14)
Bioproduction & Cloud Labs
(▶️Recording)
Reshma Shetty
Recitation (Wed, Apr 15)
Cloud laboratories
(▶️Recording | 💻Slides)
Ronan Donovan
My Homework
WEEK 11 - X
This week focuses on.
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
Assignees for the following sections
| MIT/Harvard students | Required |
| Committed Listeners | Required |
1. Contribute at least one pixel to this global artwork experiment before the editing ends on Sunday 4/19 at 11:59 PM EST.
2. Make a note on your HTGAA webpages including:
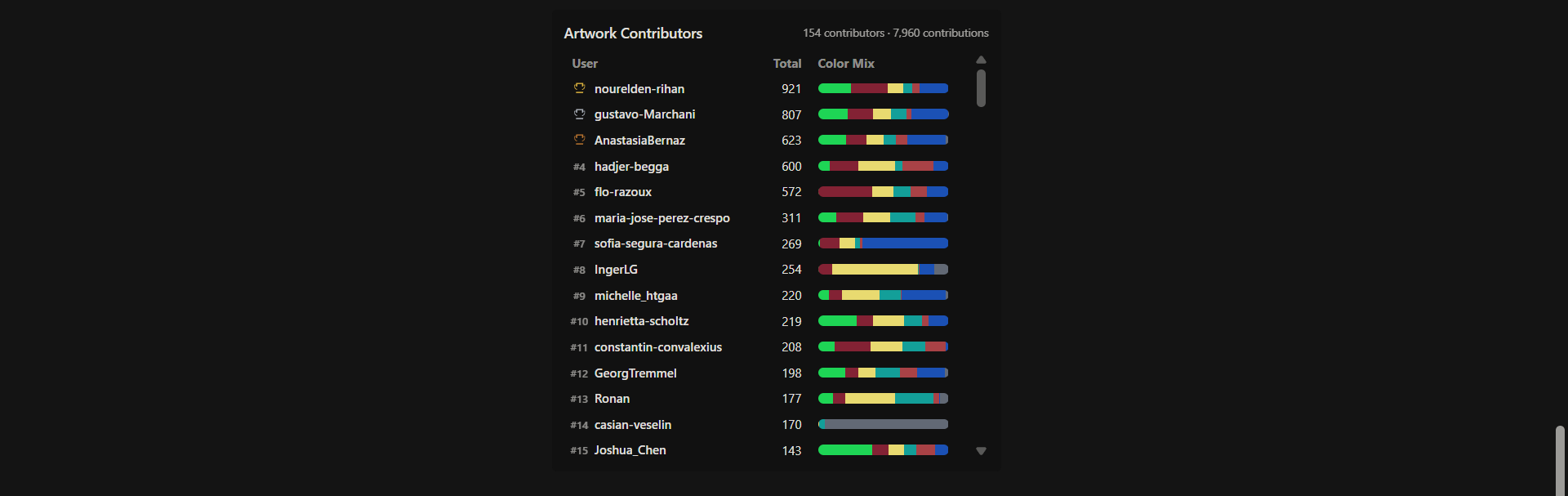
My contribution to the community bioart project
The canvas changed many times during the alocated time period, sometimes you could see some defined shapes, others they just looked like a colorfull mess. In the early satages, I contributed to a wide patch of “blue” pixels made out of Electra2, I was trying to make the figures that were already there a little more defined without erasing other people’s work. At the end, a lot of that blue patch was replaces for other patterns an little remained, but part of it still was preserved and blue pixels can be faund scattered on both, the right and left sides of the canvas with the Electra2 configuration. Also, I helped with some of the yellow mKO2 pixels at the left top corner of the piece with the “2026” design; especifically the yellow ones surrounding and filling the first “2” and the “0” have my name.
I did contributed to other color pixels, but they where minority compared to the Electra2 and mKO2 ones, by now they are mostly gone, so most of my register is plasterd with blue Electra2 ones.

What I liked about the project
I personally enjoy the “being part of something” of this iniciative, it was nice to include us all in a group project! :D
What about this collaborative art experiment could be made better for next year
Definetly the teamwork logistics! I know we had Discourse for this project, but it was still proven insufficient for clear organization since everyone had their own agenda for the design. My Node even mention that the lab logo they had work very hard on was removed SEVERAL times, wich was very sad. Next time, maybe add a rule asking for a initial design idea before starting the actual thing for wich we all can agree to, or let everyone submit a version and then vote for the one that was the most loved, then recreat it together in the final canvas using that as a blue pirnt.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
Assignees for the following sections
| MIT/Harvard students | Required |
| Committed Listeners | Required |
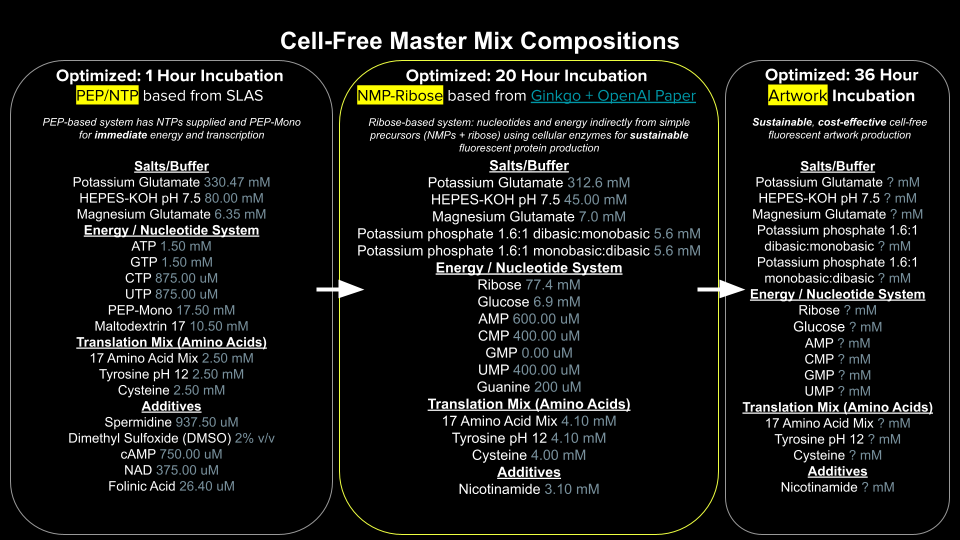

1. Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
| E. coli Lysate |
|---|
| BL21 (DE3) Star Lysate (includes T7 RNA Polymerase): Provides the complete translational machinery (ribosomes, tRNAs, aminoacyl-tRNA synthetases, elongation factors, chaperones) and the T7 RNA polymerase required to transcribe the DNA template into mRNA, enabling coupled transcription-translation in a single reaction. |
| Salts/Buffer |
|---|
| Potassium Glutamate: Acts as the primary counter-ion that stabilizes macromolecular interactions and optimizes ribosome function; glutamate is preferred over chloride as it is less inhibitory to translation. |
| HEPES-KOH pH 7.5: Maintains a stable physiological pH throughout the reaction, preventing acidification that could denature enzymes or inhibit translation. |
| Magnesium Glutamate: Provides Mg²⁺ as an essential cofactor for ribosome assembly, tRNA binding to the ribosome, and polymerase activity; its concentration is critical for translation efficiency. |
| Potassium phosphate monobasic/dibasic: Contributes to buffering capacity and provides phosphate ions that serve as building bloc |
| Energy / Nucleotide System |
|---|
| Ribose: Serves as a carbon source and precursor for de novo nucleotide synthesis through the pentose phosphate pathway, enabling sustained NTP regeneration in long reactions. |
| Glucose: Primary energy substrate that fuels glycolytic and oxidative phosphorylation pathways within the lysate to regenerate ATP and other NTPs. |
| AMP, CMP, GMP, UMP: Nucleoside monophosphates that act as substrates for cellular kinases in the lysate, which phosphorylate them to regenerate the corresponding NTPs (ATP, CTP, GTP, UTP) required for transcription. |
| Guanine: Free base that can be salvaged and converted to GMP via phosphoribosyltransferases and kinases present in the lysate, providing an alternative route for GTP regeneration. |
| Translation Mix (Amino Acids) |
|---|
| 17 Amino Acid Mix: Provides the standard set of proteinogenic amino acids (excluding tyrosine and cysteine) as substrates for protein synthesis by the ribosome. |
| Tyrosine: Supplemented separately due to its poor solubility at neutral pH; it is essential for proteins containing tyrosine residues and for proper folding of certain fluorescent proteins. |
| Cysteine: Added separately because it is chemically unstable and prone to oxidation; it is critical for disulfide bond formation and the structural integrity of many proteins. |
| Additives |
|---|
| Nicotinamide: Serves as a precursor for NAD⁺ biosynthesis, supporting redox balance and energy metabolism within the lysate during extended incubations. |
| Backfill |
|---|
| Nuclease Free Water: Used to bring the reaction to the final volume without introducing RNases or DNases that would degrade the DNA template or synthesized mRNA. |
2. Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
The 1-hour PEP-NTP mix is optimized for immediate high-yield protein production by supplying pre-formed NTPs (ATP, GTP, CTP, UTP) and phosphoenolpyruvate (PEP) as a direct energy source, enabling rapid transcription and translation but exhausting its energy reserves quickly. In contrast, the 20-hour NMP-Ribose-Glucose mix uses a regenerative strategy where simple precursors (nucleoside monophosphates + ribose + glucose) are converted into NTPs over time by the endogenous metabolic enzymes remaining in the E. coli lysate, creating a slower but sustainable energy and nucleotide supply that supports protein synthesis for extended durations.
3. Bonus question: How can transcription occur if GMP is not included but Guanine is?
Transcription can proceed because the E. coli lysate contains phosphoribosyltransferases (such as hypoxanthine-guanine phosphoribosyltransferase, HGPRT) and nucleoside/nucleotide kinases that can salvage the free guanine base. These enzymes convert guanine into GMP by transferring a phosphoribosyl group from phosphoribosyl pyrophosphate (PRPP), which is generated from ribose-5-phosphate (derived from the supplied ribose via the pentose phosphate pathway). Subsequently, cellular kinases phosphorylate GMP to GDP and then to GTP, making it available for T7 RNA polymerase during transcription. This salvage pathway demonstrates the metabolic versatility of the crude lysate system.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
Assignees for the following sections
| MIT/Harvard students | Required |
| Committed Listeners | Required |
1. Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each).
| 1. sfGFP |
|---|
| Property: Fast folding kinetics and high solubility with rapid maturation. sfGFP was engineered with six mutations (S30R, Y39N, N105T, Y145F, I171V, A206V) that enhance β-barrel folding efficiency and resistance to chemical denaturants. Its chromophore (derived from Ser65-Tyr66-Gly67) matures quickly once the protein folds, making it highly reliable in cell-free systems where rapid signal development is desired. |
| Effect on cell-free expression: The fast folding minimizes aggregation in the lysate environment, but chromophore formation still requires molecular oxygen. In sealed 384-well plates used for long incubations, oxygen depletion can become limiting, potentially slowing maturation despite fast folding. |
| 2. mRFP1 |
|---|
| Property: Slow maturation with a characteristic half-time (t₅₀) of ~22-24 minutes at 37°C and strict dependence on molecular oxygen for chromophore oxidation. The chromophore forms from the tripeptide Gln66-Tyr67-Gly68 through cyclization and oxidation steps, with oxidation being rate-limiting. |
| Effect on cell-free expression: The slow maturation creates a significant delay between protein synthesis and fluorescence detection. In cell-free reactions where oxygen is rapidly consumed by metabolic processes, mRFP1 maturation can be further delayed, making it challenging to achieve maximum fluorescence during short incubations. |
| 3. mKO2 |
|---|
| Property: Moderate acid sensitivity (pKa ~5.5-6.0) and strong dependence on oxygen tension for chromophore maturation. The chromophore requires a second oxidation step to form an acylimine linkage followed by spontaneous formation of a thiazole ring from Cys65. mKO2 exhibits slower oxidation kinetics compared to green FPs like mAG, with a pO₂·50 (oxygen tension for 50% fluorescence reduction) of 0.9%. |
| Effect on cell-free expression: The combination of acid sensitivity and high oxygen demand makes mKO2 vulnerable to pH drift and hypoxic conditions in extended cell-free reactions. Acidification of the lysate over time can protonate the chromophore, reducing fluorescence quantum yield. |
| 4. mTurquoise2 |
|---|
| Property: Exceptionally high quantum yield (QY = 0.93) and rapid maturation, but exhibits “complex maturation” kinetics where the rate progressively changes over time rather than following simple first-order exponential decay. The I146F mutation improves chromophore packing through enhanced van der Waals interactions, increasing brightness but potentially making folding more sensitive to ionic conditions. |
| Effect on cell-free expression: While its high QY provides excellent signal-to-noise ratio, the complex maturation kinetics suggest that mTurquoise2 may be sensitive to fluctuations in Mg²⁺ and K⁺ concentrations that affect ribosome-associated chaperones during extended synthesis. |
| 5. mScarlet_I |
|---|
| Property: Rapidly maturing variant of mScarlet with a T74I mutation that accelerates cellular maturation significantly (t₅₀ ~26 min at 37°C) at the cost of reduced quantum yield (0.54) and fluorescence lifetime (3.1 ns). It maintains high brightness and monomeric structure with moderate acid sensitivity (pKa ~5.3). |
| Effect on cell-free expression: The fast maturation makes mScarlet-I ideal for time-course experiments, but like all red FPs, it requires oxygen for chromophore formation. The T74I mutation may alter the protein’s interaction with endogenous chaperones in the lysate, potentially affecting folding efficiency in crude extracts. |
| 6. Electra2 |
|---|
| Property: Blue fluorescent protein derived from mRuby3 with a TagBFP-like chromophore that forms via a non-fluorescent precursor in a rate-limiting step. Electra2 exhibits photoactivation behavior where UV illumination can accelerate chromophore formation from the non-fluorescent intermediate, suggesting slow spontaneous maturation kinetics. It is 2.1× brighter than mTagBFP2 but shows aggregation tendencies in some cellular contexts. |
| Effect on cell-free expression: The rate-limiting chromophore formation from the non-fluorescent precursor means that Electra2 may accumulate as non-fluorescent protein during the initial hours of cell-free synthesis. The aggregation tendency suggests sensitivity to the reducing conditions and macromolecular crowding in the lysate. |
The amino acid sequences are shown in the HTGAA Cell-Free Benchling folder.
2. Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
| Hypothesis for 36-Hour Incubation Optimization |
|---|
| Protein: mKO2 |
| Identified Property: Strong oxygen dependence for chromophore maturation (pO₂·50 = 0.9%) and moderate acid sensitivity (pKa ~5.5-6.0). |
| Reagent Adjustment: Increase Potassium phosphate buffer concentration and supplement with additional Magnesium Glutamate. |
| Expected Effect: The additional phosphate buffer (increasing from 5.6 mM to ~8-10 mM total phosphate) will enhance pH buffering capacity over 36 hours, preventing acidification that would protonate the mKO2 chromophore and reduce fluorescence. The increased Mg²⁺ (from 7.0 mM to ~8-9 mM) will support sustained activity of metabolic enzymes in the lysate that regenerate ATP and maintain oxygen-consuming pathways, ensuring adequate oxygen availability for the second oxidation step required for mKO2 chromophore maturation. This should increase total accumulated fluorescence by maintaining optimal pH and energy metabolism throughout the extended incubation. |
3. The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here.
Important
In order to be eligible for this, make sure that your final project slide is in the “2026 Committed Listener ONE FINAL PROJECT IDEA” slide deck.
Optimized 36-Hour Artwork Master Mix Composition
Based on the progression from the 1-hour PEP/NTP mix (immediate energy) to the 20-hour NMP-Ribose mix (regenerative energy), the 36-hour Artwork mix must provide sustainable, cost-effective production with enhanced buffering and metabolic stability. The following composition optimizes for long-duration fluorescence maintenance:
| Component | 36-Hour Artwork Concentration | Rationale |
|---|---|---|
| Potassium Glutamate | 300 mM | Slightly reduced from 20-hour mix to balance ionic strength while maintaining ribosome stability |
| HEPES-KOH pH 7.5 | 50 mM | Increased from 45 mM to provide additional pH buffering over 36 hours, preventing acidification that affects mKO2 and mRFP1 |
| Magnesium Glutamate | 8.0 mM | Increased from 7.0 mM to support sustained kinase activity for NTP regeneration and chaperone function |
| Potassium phosphate (1.6:1 dibasic:monobasic) | 8.0 mM | Increased from 5.6 mM to enhance buffering capacity and provide phosphate for energy metabolism |
| Potassium phosphate (1.6:1 monobasic:dibasic) | 8.0 mM | Maintains the dual phosphate system for robust pH control |
| Ribose | 80 mM - 12.0 g/L | Slightly increased from 77.4 mM to ensure sustained PRPP generation for nucleotide salvage and NTP regeneration |
| Glucose | 8.0 mM - 1.5 g/L | Increased from 6.9 mM to fuel extended glycolytic ATP regeneration without causing excessive acidification |
| AMP | 600 μM - 0.6 mM | Maintained from 20-hour mix; sufficient for adenylate kinase-mediated ATP regeneration |
| CMP | 400 μM - 0.4 mM | Maintained from 20-hour mix |
| GMP | 0 μM | Maintained; guanine salvage provides alternative GTP source |
| UMP | 400 μM - 0.4 mM | Maintained from 20-hour mix |
| Guanine | 200 μM - 0.2 mM | Maintained; salvaged to GMP via HGPRT in lysate |
| 17 Amino Acid Mix | 4.0 mM | Slightly reduced from 4.1 mM to minimize osmotic stress while maintaining synthesis capacity |
| Tyrosine pH 12 | 4.0 mM | Maintained; critical for sfGFP and mTurquoise2 maturation |
| Cysteine | 4.0 mM | Maintained; essential for mKO2 thiazole ring formation and disulfide stability |
| Nicotinamide | 4.0 mM | Increased from 3.1 mM to enhance NAD⁺ regeneration for redox balance and extended metabolic activity |
80 mM (millimolar) ribose to g/L, useing the molar mass of ribose and unit conversion:
- The molar mass of ribose es (C₅H₁₀O₅) is 150.13 g/mol
- Convert millimolar (mM) to (g/L):
- 1 mM = 1 mmol/L = 0.001 mol/L
- 80 mM = 80×10^−3 mol/L = 0.08 mol/L
- g/L = 0.08 mol/L × 150.13 g/mol = 12.0104 g/L
8 mM (millimolar) glucose to g/L, useing the molar mass of glucose and unit conversion:
- The molar mass of glucose (C₆H₁₂O₆) is 180.156 g/mol
- Convert millimolar (mM) to molar (M):
- 8 mM = 8×10^−3 mol/L
- Multiply by molar mass to get grams per liter:
- (8×10^−3 mol/L) × 180.156 g/mol = 1.44125 g/L
600 µM of AMP to mM
- 1 mM = 1000 µM
- 600 μM = 600/1000 mM = 0.6 mM
Reaction composition per well
- 6 μL BL21 (DE3) Star Lysate
- 10 μL 2X Optimized 36-Hour Artwork Master Mix (concentrations above are 1X; 2X stocks double these values)
- 2 μL Assigned fluorescent protein DNA template
- 2 μL Custom reagent supplements (additional phosphate or Mg²⁺ for mKO2/mRFP1 wells)
Total: 20 μL reaction
4. The final phase of this lab will be analyzing the fluorescence data we collect to determine whether we can draw any conclusions about favorable reagent compositions for our fluorescent proteins. This will be due a week after the data is returned (date TBD!).
The reaction composition for each well will be as follows:
- 6 μL of Lysate
- 10 μL of 2X Optimized Master Mix from above
- 2 μL of assigned fluorescent protein DNA template
- 2 μL of your custom reagent supplements
Total: 20 μL reaction
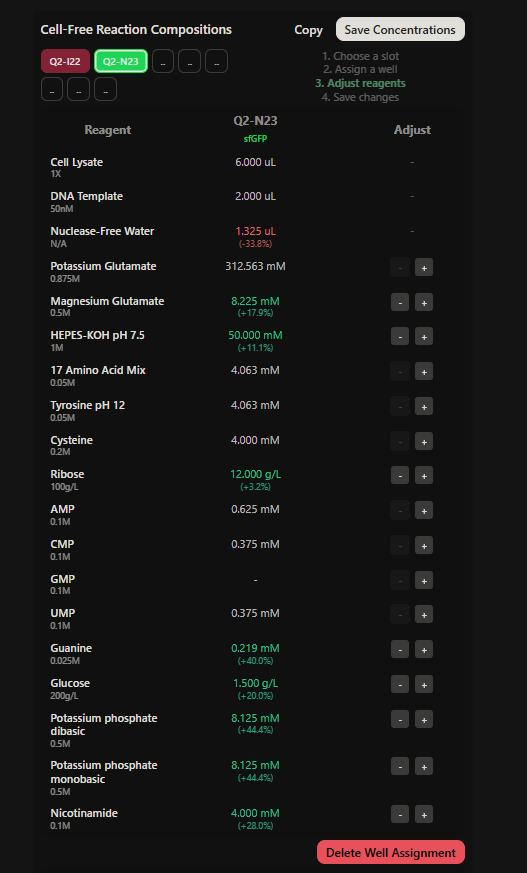
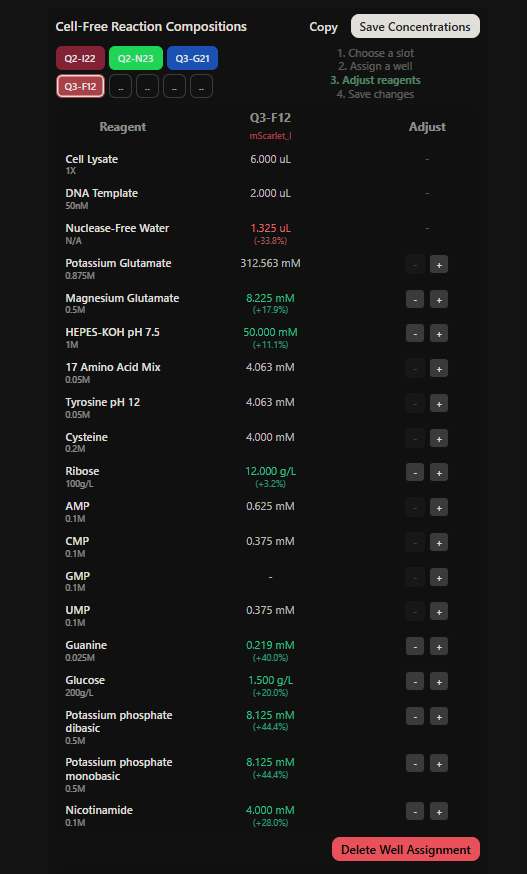
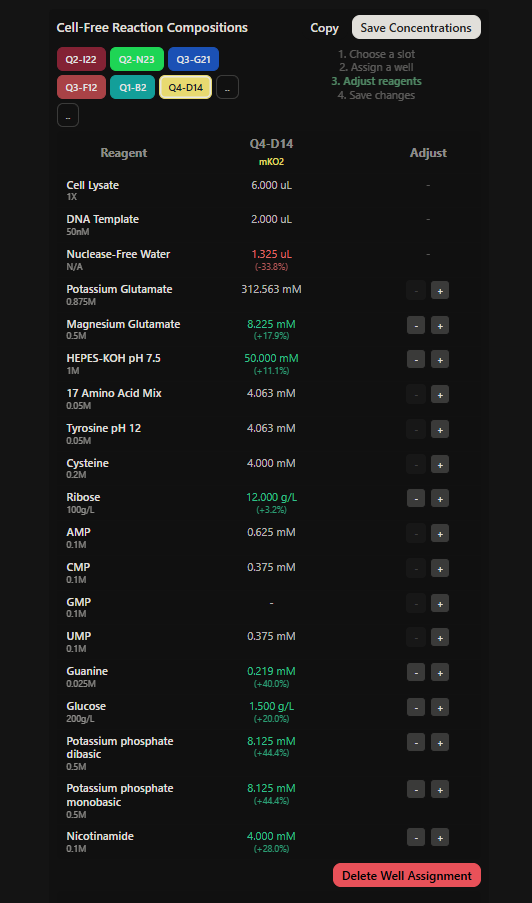
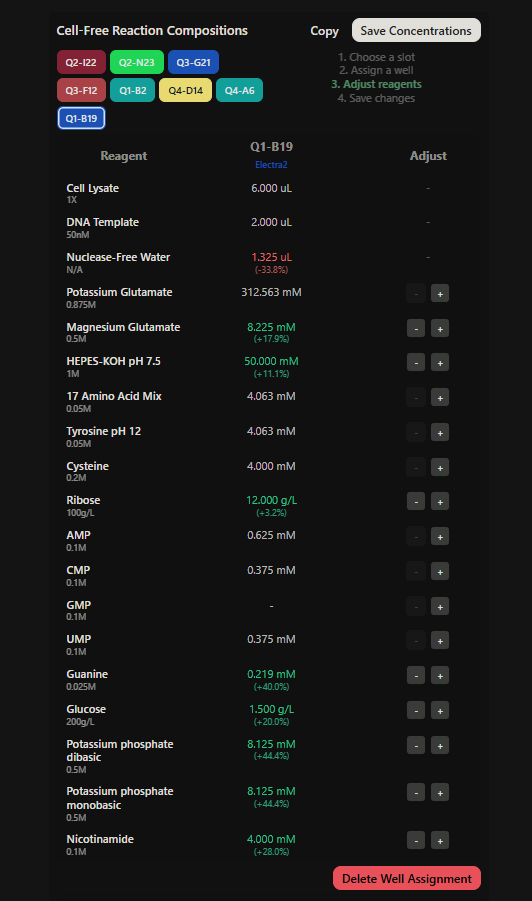
Final optimized 8 wells: Master Mix combos
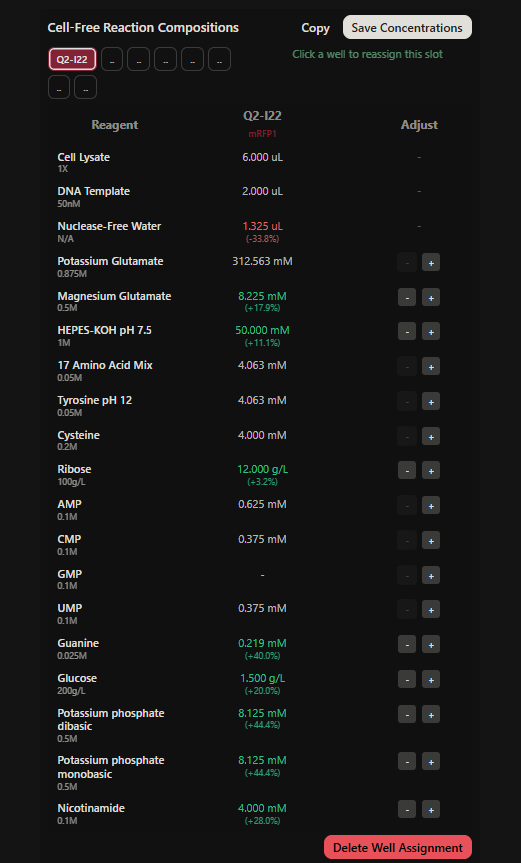
 | 
 |
|---|
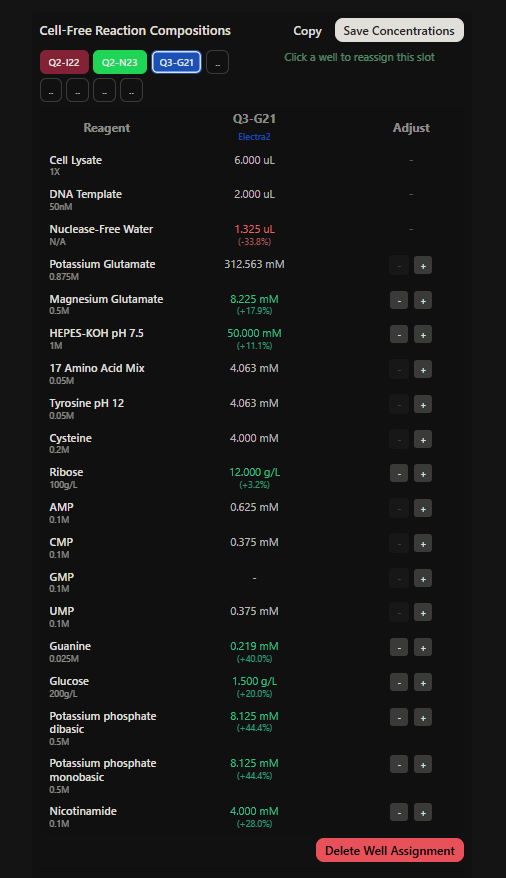
 | 
 |
|---|
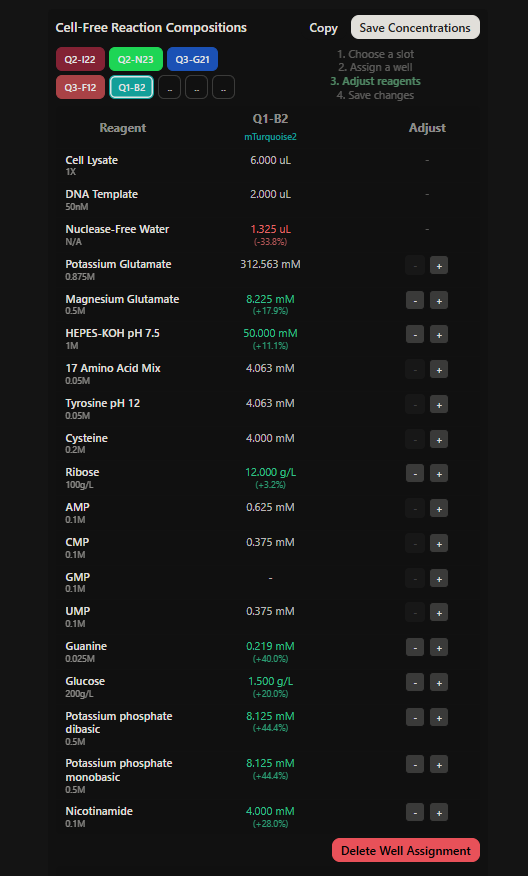
 | 
 |
|---|
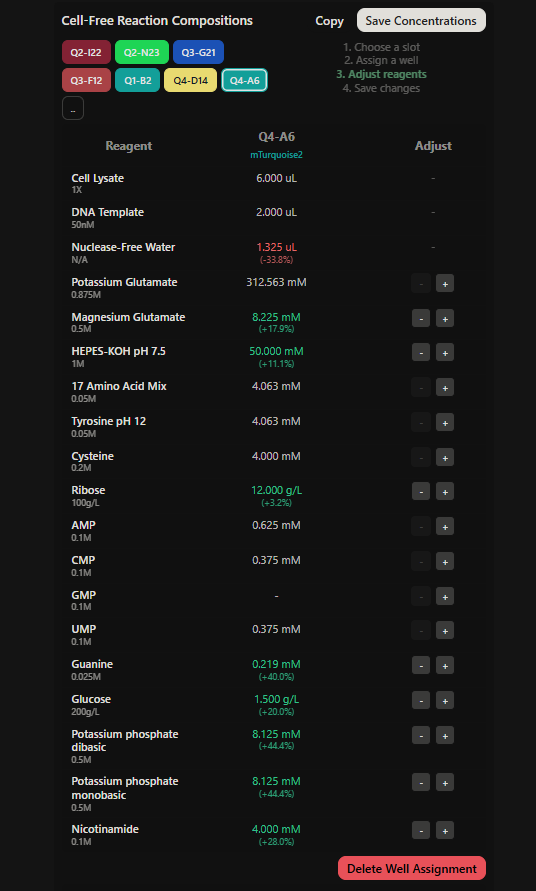
 | 
 |
|---|
Reagent Supplement JSON
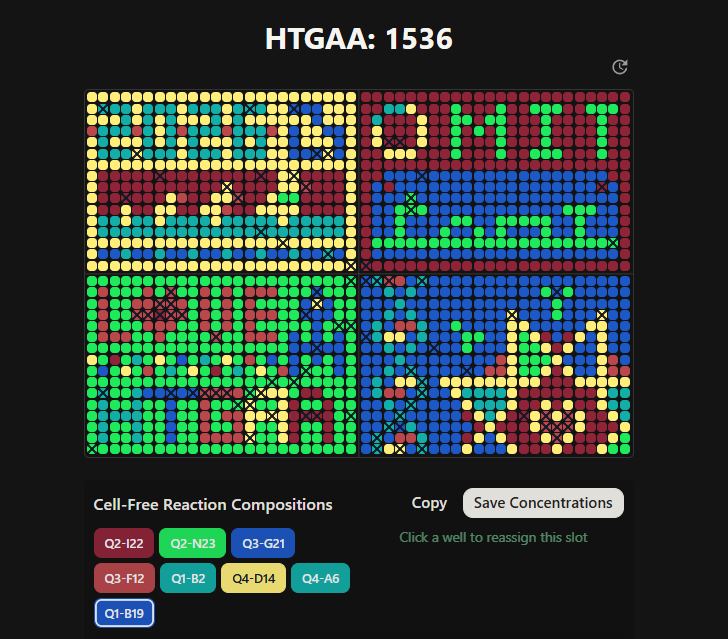
Master Mix contributions: Canvas final look

 | 
 |
|---|
Part D: Build-A-Cloud-Lab | (optional) Bonus Assignment
Assignees for the following sections
| MIT/Harvard students | Optional |
| Committed Listeners | Optional |

Ginkgo Nebula Cloud Laboratory Rendering, 2025
- Use this simulation tool to create an interesting looking cloud lab out of the Ginkgo Reconfigurable Automation Carts. This is just a minimal implementation so far, but I would love to see some fun designs!
Tip
Note from Ronan: If you are interested in helping me build out future HTGAA cloud lab software, please fill out this form!
Resources
Reading:
- Recitation slides from week 3
- Nebula RACs TA Onboarding slides from HTGAA Summer Research
- Using a GPT-5-driven autonomous lab to optimize the cost and titer of cell-free protein synthesis
- Design-driven optimization of low-cost reagent formulations for reproducible and high-yielding cell-free gene expression
Common Nebula protocols & their parameters