Week 1 HW: Principles and Practices
- First, describe a biological engineering application or tool you want to develop and why. This could be inspired by an idea for your HTGAA class project and/or something for which you are already doing in your research, or something you are just curious about.
I have worked with the concept of CA before within design and 3d space generative making through creating tools for generating patterns and environments, so it was really fascinating to see it being brought up during class. So, for my idea I’d like to merge my previous digital experience with CA and synthetic biology tooling in a form of a computer aided design tool for spatial synthetic biology
- Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals.
Ensuring biosafety: a) Possibility of the prediction of biological behaviour b) Testable behaviour (not just false confidence in whatever is happening, both for a and b!) c) safety protocols built in within the ux/ui of the software d) training provided
Transparency whilst preventing harm: a) Maintenance of accountability over the created projects (as in not fully automated) b) Responsible use c) (Ideally!) some sort of encouragement of socially beneficial applications
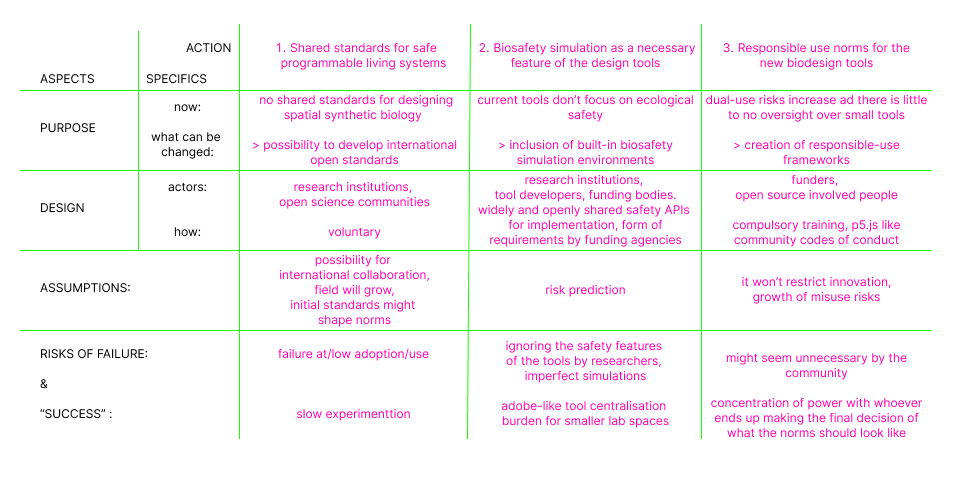
- Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”). Try to outline a mix of actions (e.g. a new requirement/rule, incentive, or technical strategy) pursued by different “actors” (e.g. academic researchers, companies, federal regulators, law enforcement, etc). Draw upon your existing knowledge and a little additional digging, and feel free to use analogies to other domains (e.g. 3D printing, drones, financial systems, etc.).
- Purpose: What is done now and what changes are you proposing?
- Design: What is needed to make it “work”? (including the actor(s) involved - who must opt-in, fund, approve, or implement, etc)
- Assumptions: What could you have wrong (incorrect assumptions, uncertainties)?
- Risks of Failure & “Success”: How might this fail, including any unintended consequences of the “success” of your proposed actions?

- Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals. The following is one framework but feel free to make your own:
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | 2 | 1 | 2 |
| • By helping respond | 3 | 2 | 2 |
| Foster Lab Safety | |||
| • By preventing incident | 1 | 1 | 2 |
| • By helping respond | 2 | 1 | 3 |
| Protect the environment | |||
| • By preventing incidents | 3 | 2 | 1 |
| • By helping respond | 2 | 2 | 1 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 3 | 2 | 1 |
| • Feasibility? | 2 | 2 | 1 |
| • Not impede research | 2 | 3 | 1 |
| • Promote constructive applications | 1 | 1 | 2 |
Homework Questions from Professor Jacobson: [Lecture 2 slides]
1)Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase?
1:10⁶
How does this compare to the length of the human genome?
approximately 3.2 Gbp
How does biology deal with that discrepancy?
Through 5’-3’ error-correcting exonuclease and 3’-5’ proofreading exonucleas + there is also MutS Repair System
2)How many different ways are there to code (DNA nucleotide code) for an average human protein?
2³⁴⁵ or higher (as there is 1,036 bp and 345 amino acids)
In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
- because of the secondary structure formation
- RNA cleavage
- mRNA instability
- different organisms might have preferences for certain codons
Homework Questions from Dr. LeProust: [Lecture 2 slides]
1.What’s the most commonly used method for oligo synthesis currently?
The phosphoramidite method
2.Why is it difficult to make oligos longer than 200nt via direct synthesis?
Because of the 1)error accumulation, 2)”enhanced chemistry” is required
3.Why can’t you make a 2000bp gene via direct oligo synthesis?
Error accumulation and length limitations > assembly approach is required
Homework Question from George Church: [Lecture 2 slides]
[Given slides #2 & 4 (AA:NA and NA:NA codes)] What code would you suggest for AA:AA interactions?
A protein–protein interaction code