Week 2 HW: DNA Read, Write, & Edit
Part 1: Benchling & In-silico Gel Art
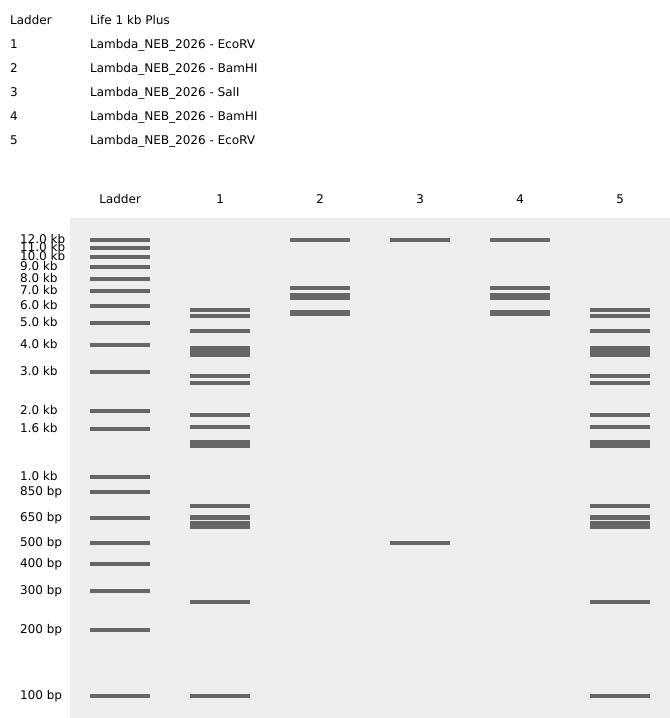
Here’s my version of made in Benchling pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks :)
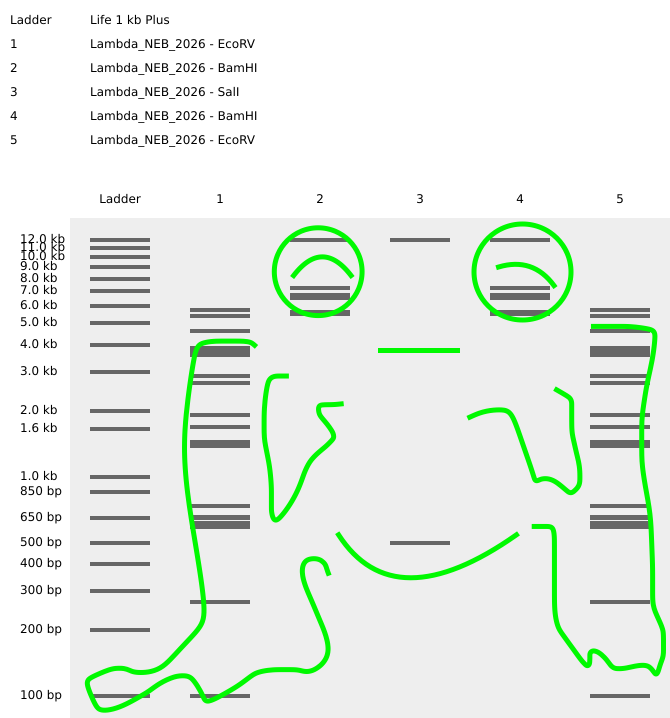

https://benchling.com/s/seq-E1w2i5oXtxnqS8BWnwcZ?m=slm-mz29OlZAWUwDeJTn4nir
I also played around separately to that drawing with Ronan’s tool, and this is what I got ( I wanted to go for something more abstract this time)
| Enzyme | Number of Cuts | Number of Fragments | Fragment Sizes (bp) |
|---|---|---|---|
| EcoRI | 5 | 6 | 21,226 / 7,421 / 5,804 / 5,643 / 4,878 / 3,530 |
| HindIII | 6 | 7 | 9,416 / 6,682 / 4,361 / 3,130 / 2,322 / 2,027 / 564 |
| BamHI | 5 | 6 | 16,841 / 7,233 / 6,770 / 6,527 / 5,626 / 5,505 |
| KpnI | 2 | 3 | 29,942 / 17,057 / 1,503 |
| EcoRV | 21 | 22 | 5,765 / 5,376 / 4,613 / 3,873 / 3,744 / 3,595 / 2,884 / 2,674 / 1,921 / 1,679 / 1,434 / 1,403 / 1,377 / 1,313 / 738 / 655 / 618 / 597 / 588 / 268 / 52 / 35 |
| SacI | 2 | 3 | 24,776 / 22,621 / 1,105 |
| SalI | 2 | 3 | 32,745 / 15,258 / 499 |


Restriction Enzymes Used
- EcoRI
- EcoRV
- HindIII
- KpnI
- BamHI
- SacI
- SalI
Restriction Digest Setup
| Lane | Water | CutSmart Buffer | λ DNA | Enzyme(s) |
|---|---|---|---|---|
| M (Ladder) | 14 μL | 2 μL | 3 μL | - |
| 1 | 13 μL | 2 μL | 3 μL | 1 μL EcoRI |
| 2 | 13 μL | 2 μL | 3 μL | 1 μL KpnI + 1 μL BamHI |
| 3 | 14 μL | 2 μL | 3 μL | 1 μL EcoRI + 1 μL HindIII |
| 4 | 14 μL | 2 μL | 3 μL | 1 μL EcoRV |
| 5 | 13 μL | 2 μL | 3 μL | 1 μL EcoRI + 1 μL KpnI |
| 6 | 13 μL | 2 μL | 3 μL | 1 μL SacI + 1 μL HindIII |
| 7 | 13 μL | 2 μL | 3 μL | 1 μL SacI + 1 μL SacI |
| 8 | 13 μL | 2 μL | 3 μL | 1 μL SalI + 1 μL KpnI |
| 9 | 14 μL | 2 μL | 3 μL | 1 μL SacI + 1 μL SacI |
| 10 | 13 μL | 2 μL | 3 μL | 1 μL HindIII + 1 μL SacI |
Total volume per tube: 20 μL
Restriction Digest Parameters
- Incubation: 37°C for 60 minutes
- Heat Inactivation (optional): 80°C for 20 minutes
DNA Gel Electrophoresis
Goal: 100-150 ng of DNA per lane
Hand-Cast Gel Protocol
Digest Sample:
- 14.7 μL Water
- 3.3 μL Loading Dye
- 2 μL Digest
- Total: 20 μL
Ladder:
- 6.6 μL Water
- 3.3 μL Loading Dye
- 10 μL Ladder (15 ng/μL stock)
- Total: 20 μL
E-Gel Protocol
Digest Sample:
- 18 μL Water
- 2 μL Digest
- Total: 20 μL
Ladder:
- 10 μL Water
- 10 μL Ladder (15 ng/μL stock)
- Total: 20 μL
Part 3: DNA Design Challenge
3.1 For this task, I’d like to use green fluorescent protein also known as GFP as it is, firstly, safe and pretty well characterised (as described here! https://pubmed.ncbi.nlm.nih.gov/8303295/), essential for CA mechanics to be working (https://www.science.org/doi/10.1126/science.abb8205) and just generally directly connects to tool making (https://www.nature.com/articles/s41467-024-53078-8)
sp|P42212|GFP_AEQVI Green fluorescent protein OS=Aequorea victoria OX=6100 GN=GFP PE=1 SV=1 MSKGEELFTGVVPILVELDGDVNGHKFSVSGEGEGDATYGKLTLKFICTTGKLPVPWPT LVTTFSYGVQCFSRYPDHMKQHDFFKSAMPEGYVQERTIFFKDDGNYKTRAEVKFEGDT LVNRIELKGIDFKEDGNILGHKLEYNYNSHNVYIMADKQKNGIKVNFKIRHNIEDGSVQ LADHYQQNTPIGDGPVLLPDNHYLSTQSALSKDPNEKRDHMVLLEFVTAAGITHGMDELYK
3.2 Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
based on this tool https://www.bioinformatics.org/sms2/rev_trans.html
reverse translation of sp|P42212|GFP_AEQVI Green fluorescent protein OS=Aequorea victoria OX=6100 GN=GFP PE=1 SV=1 to a 714 base sequence of consensus codons. atgwsnaarggngargarytnttyacnggngtngtnccnathytngtngarytngayggn gaygtnaayggncayaarttywsngtnwsnggngarggngarggngaygcnacntayggn aarytnacnytnaarttyathtgyacnacnggnaarytnccngtnccntggccnacnytn gtnacnacnttywsntayggngtncartgyttywsnmgntayccngaycayatgaarcar caygayttyttyaarwsngcnatgccngarggntaygtncargarmgnacnathttytty aargaygayggnaaytayaaracnmgngcngargtnaarttygarggngayacnytngtn aaymgnathgarytnaarggnathgayttyaargargayggnaayathytnggncayaar ytngartayaaytayaaywsncayaaygtntayathatggcngayaarcaraaraayggn athaargtnaayttyaarathmgncayaayathgargayggnwsngtncarytngcngay caytaycarcaraayacnccnathggngayggnccngtnytnytnccngayaaycaytay ytnwsnacncarwsngcnytnwsnaargayccnaaygaraarmgngaycayatggtnytn ytngarttygtnacngcngcnggnathacncayggnatggaygarytntayaar
3.3. Codon optimization.
based on this tool - https://en.vectorbuilder.com/tool/codon-optimization/51ebfb2e-a00a-4190-9fe9-c1578d3ecfbe.html
Pasted Sequence: GC=48.60%, CAI=1.00
ATGAGCAAAGGCGAAGAACTGTTTACCGGCGTGGTGCCGATTCTGGTGGAACTGGATGGCGATGTGAACGGCCATAAATTTAGCGTGAGCGGCGAAGGCGAAGGCGATGCGACCTATGGCAAACTGACCCTGAAATTTATTTGCACCACCGGCAAACTGCCGGTGCCGTGGCCGACCCTGGTGACCACCTTTAGCTATGGCGTGCAGTGCTTTAGCCGCTATCCGGATCATATGAAACAGCATGATTTTTTTAAAAGCGCGATGCCGGAAGGCTATGTGCAGGAACGCACCATTTTTTTTAAAGATGATGGCAACTATAAAACCCGCGCGGAAGTGAAATTTGAAGGCGATACCCTGGTGAACCGCATTGAACTGAAAGGCATTGATTTTAAAGAAGATGGCAACATTCTGGGCCATAAACTGGAATATAACTATAACAGCCATAACGTGTATATTATGGCGGATAAACAGAAAAACGGCATTAAAGTGAACTTTAAAATTCGCCATAACATTGAAGATGGCAGCGTGCAGCTGGCGGATCATTATCAGCAGAACACCCCGATTGGCGATGGCCCGGTGCTGCTGCCGGATAACCATTATCTGAGCACCCAGAGCGCGCTGAGCAAAGATCCGAACGAAAAACGCGATCATATGGTGCTGCTGGAATTTGTGACCGCGGCGGGCATTACCCATGGCATGGATGAACTGTATAAA
Improved DNA[1]: GC=48.88%, CAI=0.97
ATGAGCAAAGGCGAAGAACTGTTTACCGGCGTGGTGCCGATTCTGGTGGAACTGGATGGCGATGTGAATGGCCATAAATTTAGCGTGAGCGGCGAAGGTGAAGGCGATGCGACCTATGGCAAACTGACCCTGAAATTTATCTGCACCACCGGTAAACTGCCGGTGCCGTGGCCGACCCTGGTGACCACCTTCAGCTACGGCGTGCAGTGTTTTAGCCGCTACCCGGATCATATGAAACAGCATGATTTTTTTAAAAGCGCGATGCCGGAAGGCTATGTGCAGGAACGCACCATTTTTTTCAAAGATGATGGCAATTACAAAACCCGTGCCGAAGTGAAATTCGAAGGCGATACCCTGGTGAATCGCATTGAACTGAAAGGCATTGATTTTAAAGAAGATGGTAACATTCTGGGCCACAAACTGGAATACAACTATAACAGCCATAACGTGTACATTATGGCGGATAAACAGAAAAATGGCATTAAAGTGAACTTTAAAATTCGCCATAACATTGAAGATGGCTCAGTGCAGCTGGCGGATCACTATCAGCAGAACACCCCGATTGGCGATGGCCCGGTTCTGCTGCCGGATAACCACTATCTGAGCACCCAGAGCGCGCTGTCGAAAGATCCGAACGAAAAACGCGATCACATGGTGCTGCTGGAATTTGTGACCGCCGCGGGCATCACCCATGGTATGGATGAACTGTATAAA Avoid cleavage sites of restriction enzymes: BbsI BsaI
It’s done to get a more reliable and consistent protein expression!
I chose Escherichia coli K-12 substr. MG1655 because it’s standard and saf to work with + GFP expression is well established within it
3.4 If it’s a cell-dependent method the DNA sequence can be transcribed and translated into my protein through these steps: 1. Transformation 2.Transcription 3.Translation 4.Protein Folding 5.Fluorescence
Part 4: Prepare a Twist DNA Synthesis Order
https://benchling.com/s/seq-TYyZiuVgAJUcWN179kJj?m=slm-SR2kt8YFMq4ngkmbdM2Q
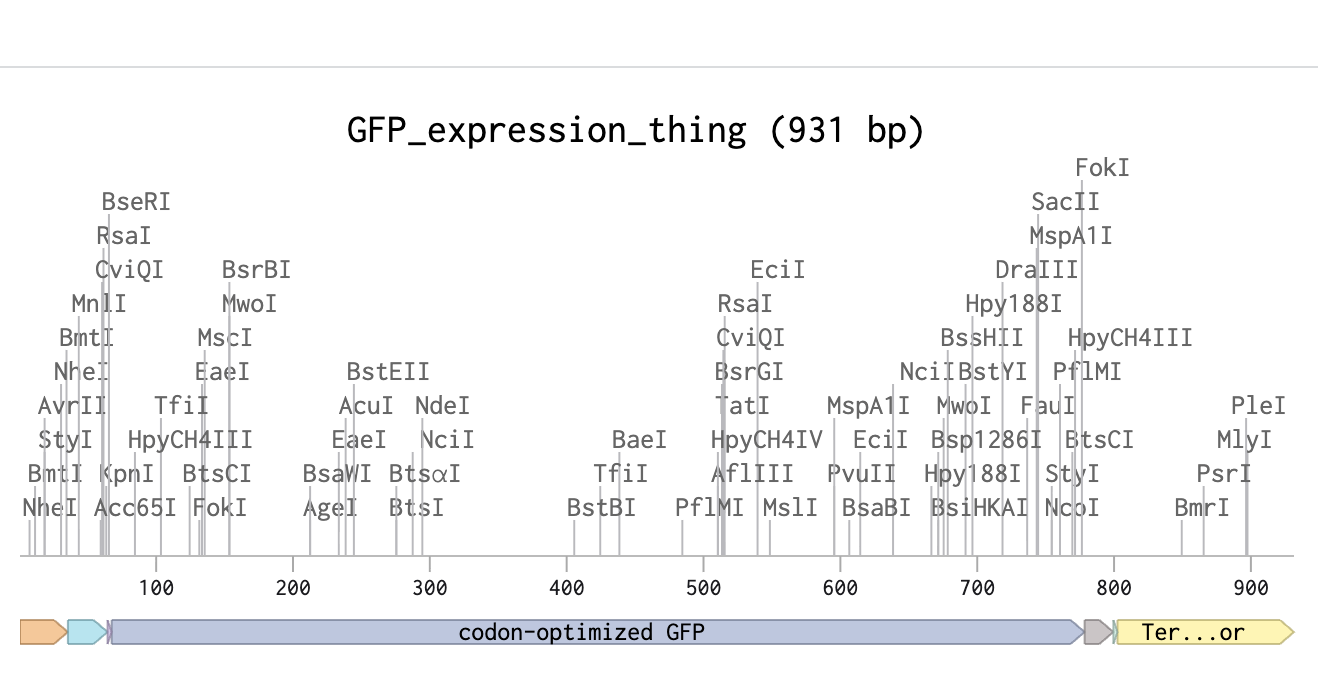

 Afterwards in Twist I got a bunch of errors when importing as when I was making a sequence a copy pasted an article “a"into it as it was a part of it. It is also visible here as 931 is not divisible by 3
fixed version!>
Afterwards in Twist I got a bunch of errors when importing as when I was making a sequence a copy pasted an article “a"into it as it was a part of it. It is also visible here as 931 is not divisible by 3
fixed version!>
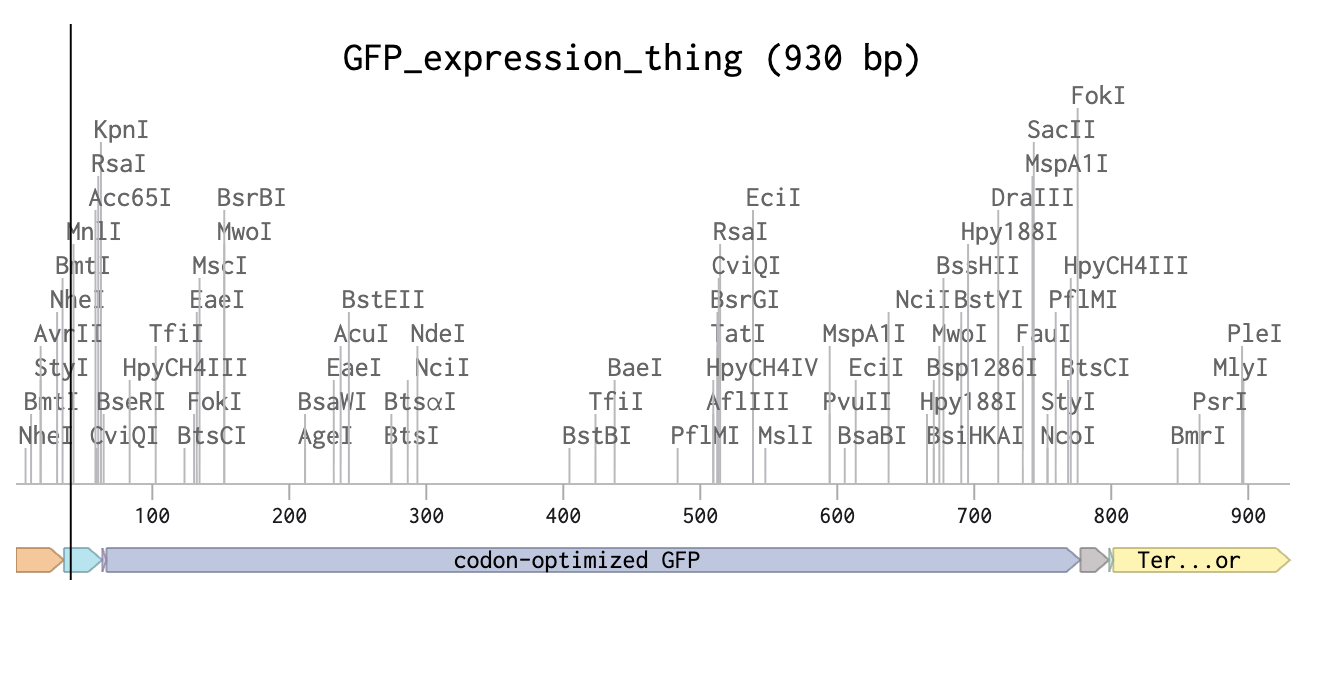

 how it works>
how it works>
final link https://benchling.com/s/seq-jNpnBifWoDGPy7RLJoHu?m=slm-nUfWbFSbXctEauGsTTaq
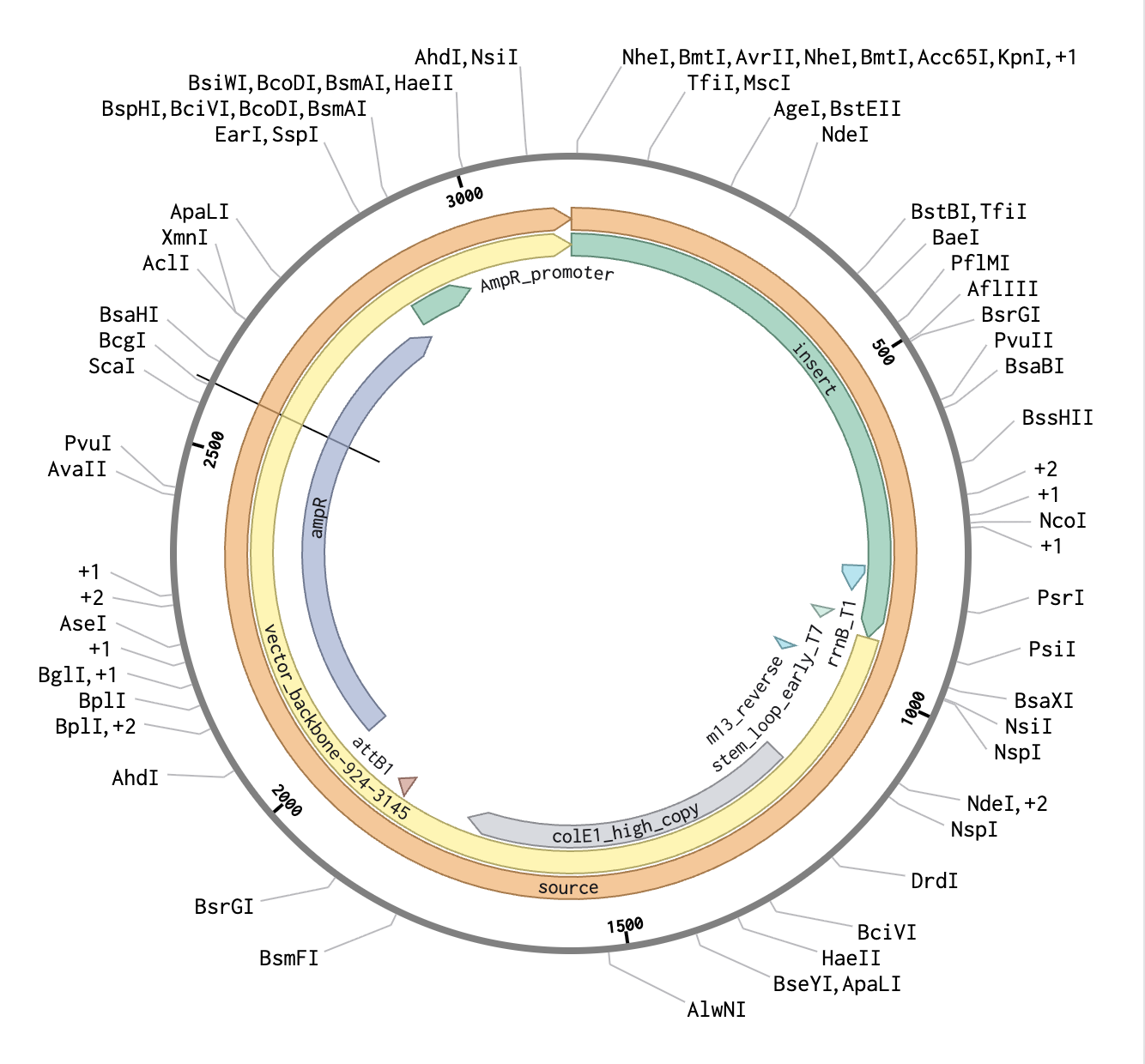

Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
So far some sort of plasmids encoding CA-based pattern formation circuits in bacterial populations to be able to design patterns with them for the final project tool
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Probably something like Illumina as it is pretty commonly used which means it will be easier to find
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
GFP-based circuits so I could prototype programmable living pattern systems
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
PCR or Gibson assembly to do precise construction of my custom genetic circuit
And then in terms of computer based tech Benchling and Twist
5.3 DNA Edit
(i) What DNA would you want to edit and why?
So far based on my research for this project E. coli K-12 MG1655as it is safe, well-documented, accessible and also works amazing with my project
(ii) What technology or technologies would you use to perform these DNA edits and why?
I’m not fully sure what’s possible to use for the projects yet but if I can CRISPR-Cas9 as it can precisely edit genetic circuits
