Week 5 HW: Protein Design Part II
Part A: SOD1 Binder Peptide Design (From Pranam)
Part 1: Generate Binders with PepMLM
sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2 MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ (taken from https://rest.uniprot.org/uniprotkb/P00441.fasta ) -> muted form A-> V MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
- (2-5)
# Binder Pseudo Perplexity Notes 0 WHYGAAQAAHWX 7.60026377248636 High confidence 1 WLYGASAAAWKK 7.46473740432208 Highest confidence!!! 2 WLYGAAGVAWKE 10.9325804754158 Moderate confidence 3 WLYYPQAAKLKK 15.5499787120909 Lowest confidence — FLYRWLPSRRGG 20.9180890005569 Known binder (control)
Part 2: Evaluate Binders with AlphaFold3
So what I found out is that in WHYGAAQAAHWX X is an unknown amino acid and that AlphaFold3 is not going to work with it so I’m just skipping it for now.
| Peptide | ipTM | pTM | Binding Location |
|---|---|---|---|
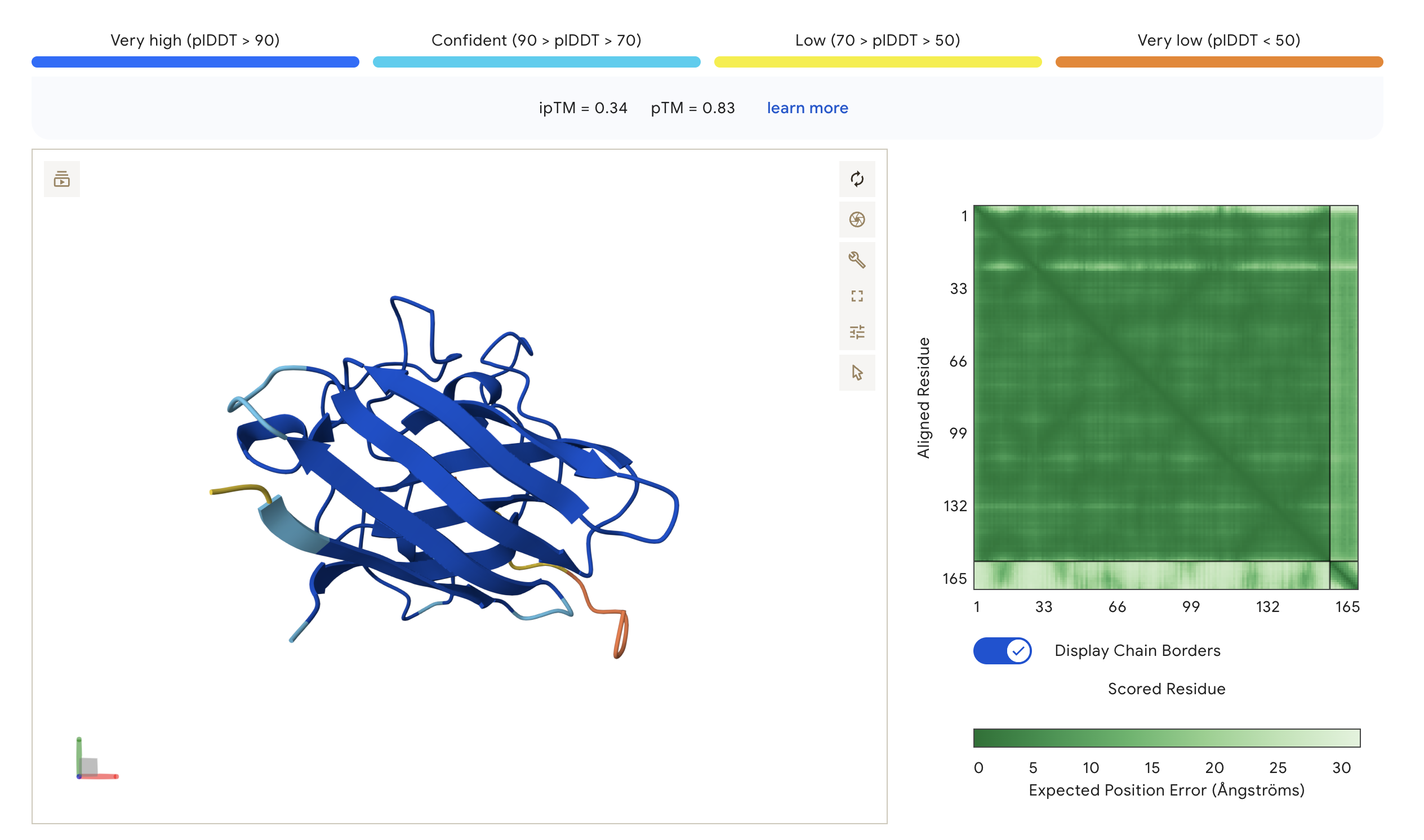
| FLYRWLPSRRGG (known) | 0.34 | 0.83 | Surface-bound, near bottom/loop region |
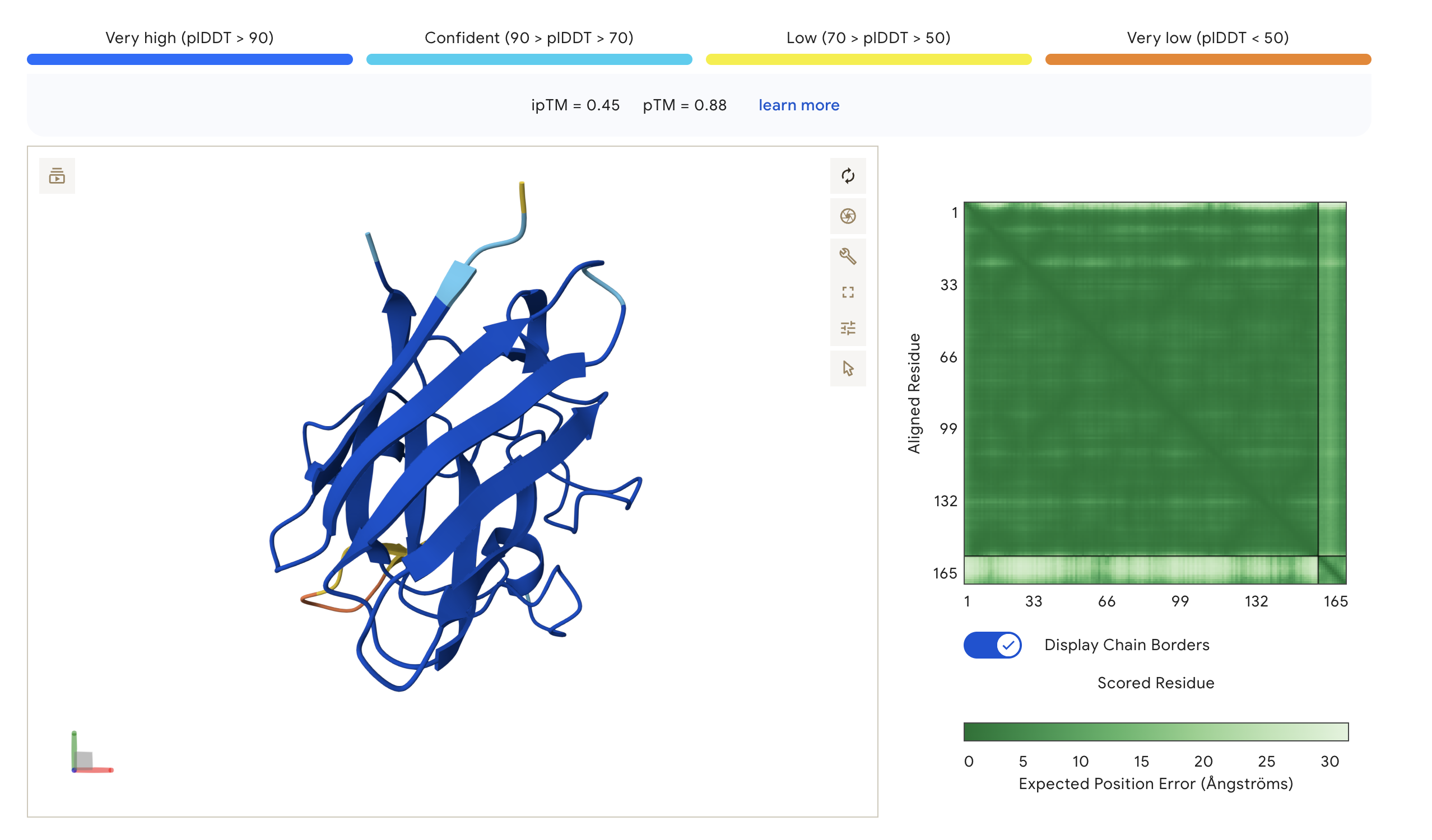
| WLYGAAGVAWKE | 0.45 | 0.88 | Engages β-barrel, partially buried near core |
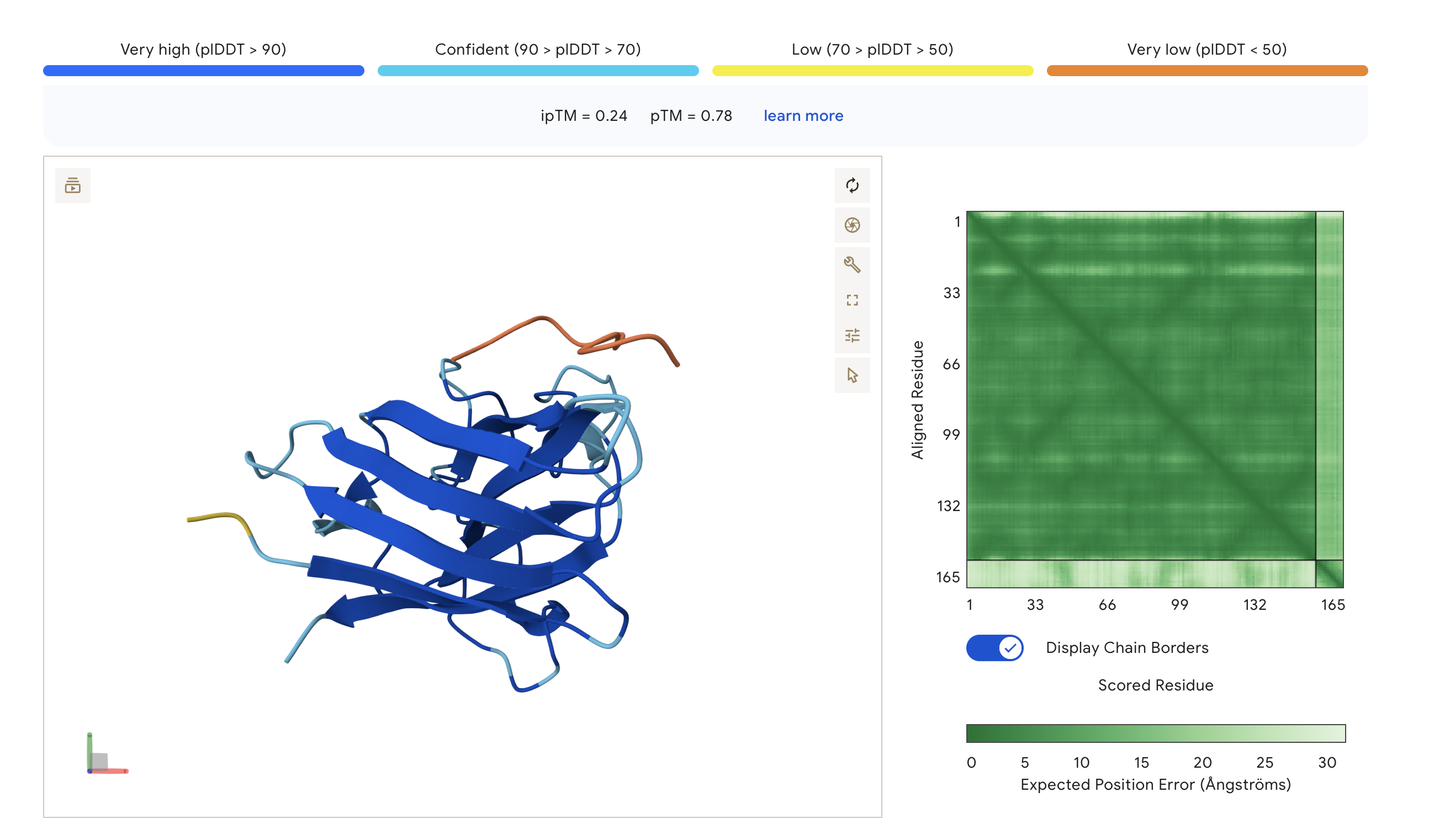
| WLYGASAAAWKK | 0.24 | 0.78 | Surface-bound, near N-terminus/top loops |
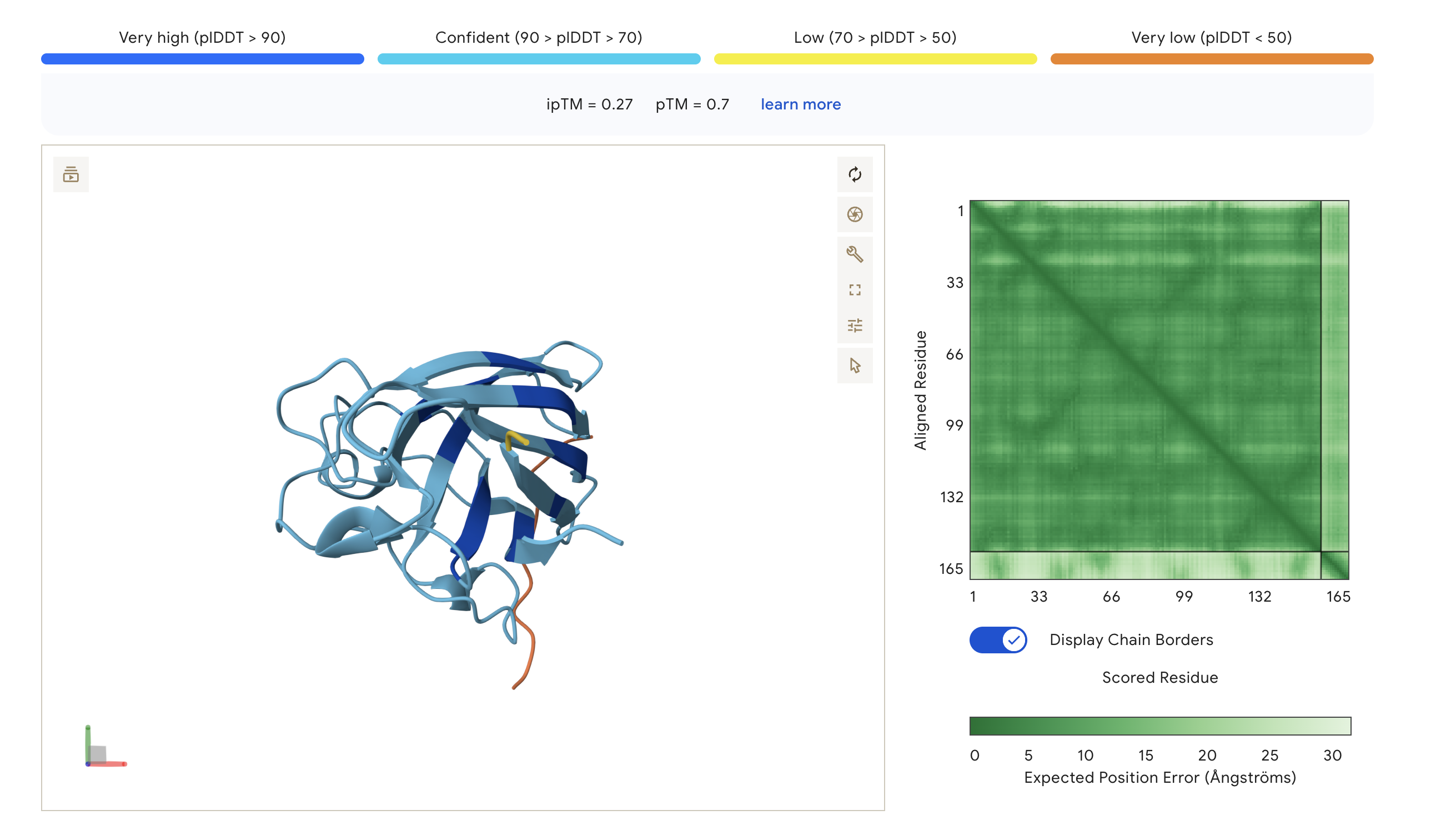
| WLYYPQAAKLKK | 0.27 | 0.70 | Surface-bound, loose association near loops |
- ipTM scores across all peptides ranged from 0.24 to 0.45 -> suggests weak-to-moderate predicted interface confidence. The PepMLM-generated peptide WLYGAAGVAWKE (ipTM = 0.45) outperformed the known binder FLYRWLPSRRGG (ipTM = 0.34), appearing more engaged with the β-barrel core of SOD1. The remaining two peptides WLYGASAAAWKK (ipTM = 0.24) and WLYYPQAAKLKK (ipTM = 0.27) scored below FLYRWLPSRRGG and appeared loosely surface-bound. None exceeded an ipTM of 0.5, which should be expected for short peptides against a structured target
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
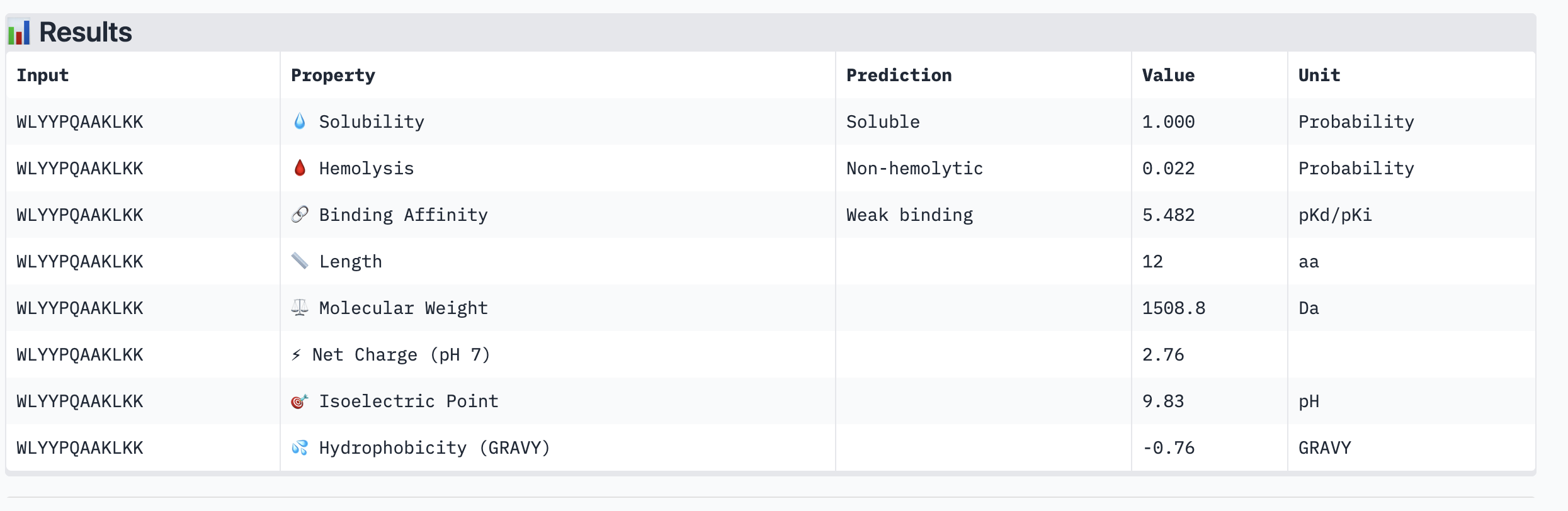
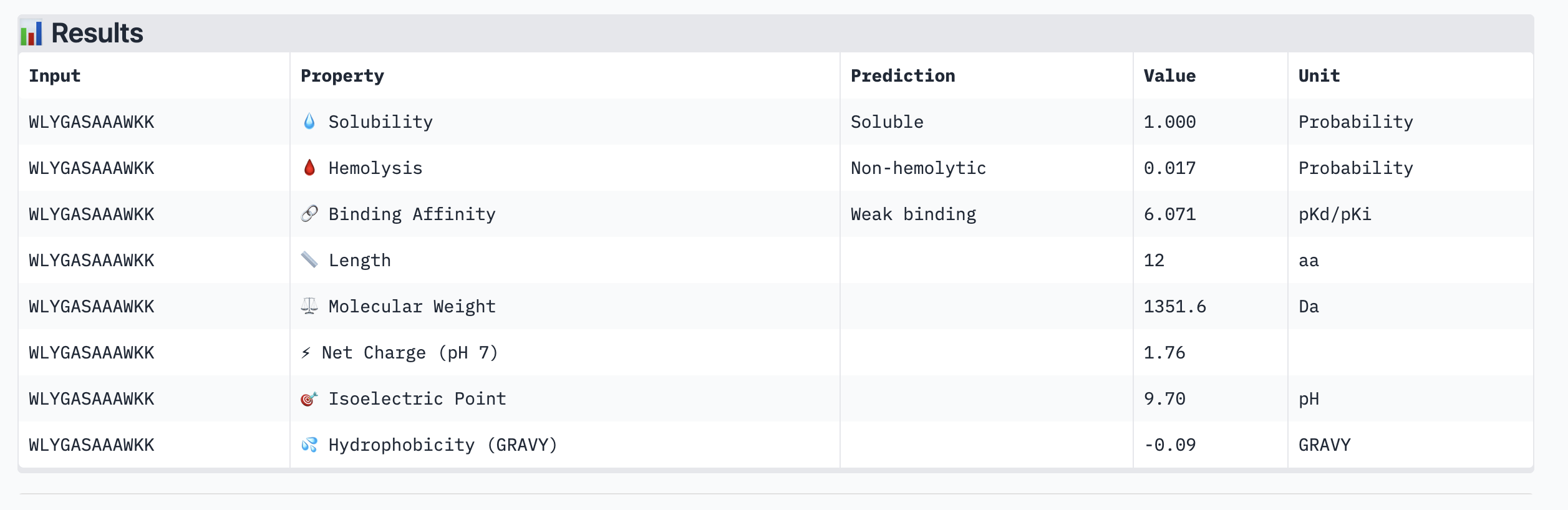
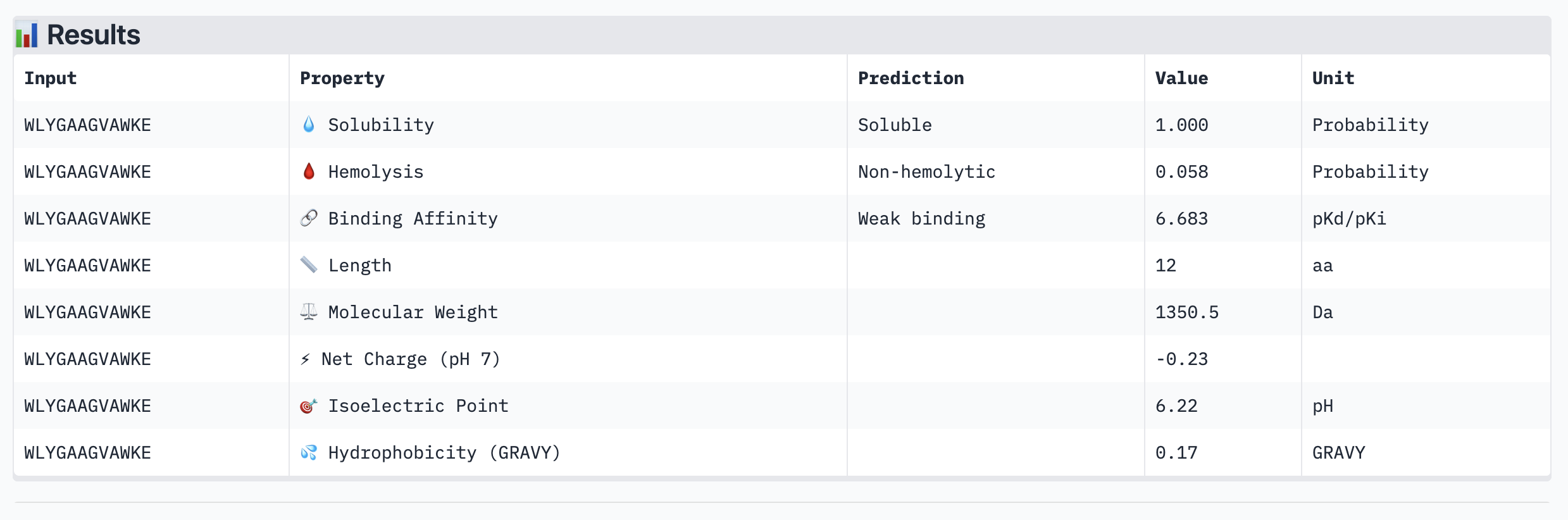
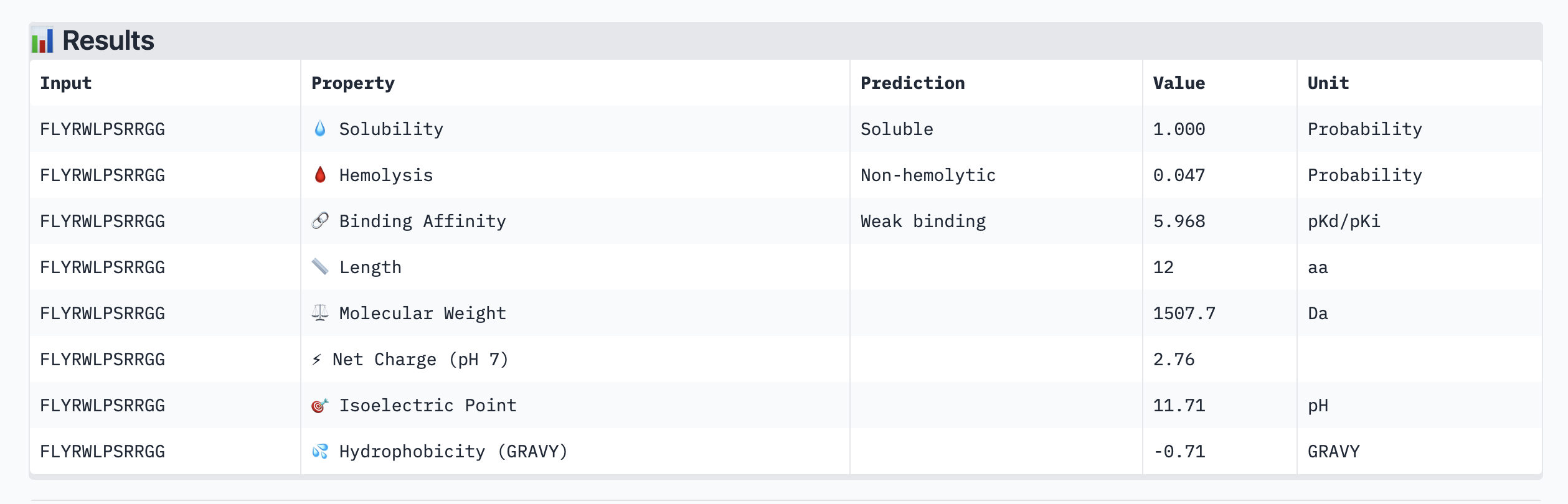
| Peptide | ipTM | Binding Affinity (pKd) | Solubility | Hemolysis | Net Charge |
|---|---|---|---|---|---|
| FLYRWLPSRRGG (known) | 0.34 | 5.968 | Soluble | Non-hemolytic | +2.76 |
| WLYGAAGVAWKE | 0.45 | 6.683 | Soluble | Non-hemolytic | -0.23 |
| WLYGASAAAWKK | 0.24 | 6.071 | Soluble | Non-hemolytic | +1.76 |
| WLYYPQAAKLKK | 0.27 | 5.482 | Soluble | Non-hemolytic | +2.76 |
WLYGAAGVAWKE had both the highest ipTM (0.45) and the strongest predicted binding affinity (6.683 pKd), suggesting that structural confidence and binding prediction align for this peptide. All peptides were soluble and non-hemolytic, meaning none raised safety red flags. Notably, higher ipTM did loosely correlate with stronger affinity — WLYGAAGVAWKE topped both metrics while WLYYPQAAKLKK scored lowest on affinity and had a weak ipTM.
Peptide to Advance: WLYGAAGVAWKE!!!
WLYGAAGVAWKE has the best structural binding confidence (ipTM = 0.45), the strongest predicted affinity (6.683 pKd), is fully soluble, and non-hemolytic. It outperforms the known binder FLYRWLPSRRGG on every key metric! Therefore, we should use this one
Part 4: Generate Optimized Peptides with moPPIt
I tried to run for Motif positions 1, 2, 3, 4, 5, 6, 7 but it’s just taking so long, so, I’m nit going to finosh running it but here’re some results that I got so far:
moPPIt Generated Peptides
| # | Peptide | Target Residues |
|---|---|---|
| 1 | GKTEKTYTDCCD | 1, 2, 3, 4, 5, 6, 7 |
| 2 | EEQNTCIQTTKA | 1, 2, 3, 4, 5, 6, 7 |
Comparison: moPPIt vs PepMLM Peptides:
moPPIt peptides differ notably from PepMLM-generated ones in both composition and design intent. PepMLM peptides (e.g., WLYGAAGVAWKE, WLYGASAAAWKK) were dominated by W at position 1 and showed a hydrophobic character, as the model simply sampled sequences likely to bind SOD1 without any spatial constraints. In contrast, moPPIt peptides (GKTEKTYTDCCD, EEQNTCIQTTKA) are more polar and charged while balancing affinity, solubility and hemolysis objectives. moPPIt asks “what binds specifically near position 4, and is also therapeutically viable?”
Before advancing the studies of moPPIt peptides, the following next steps/ evaluations needed:
- SPR or ITC should be used to measure actual binding affinity
- AlphaFold3 rystallography to confirm binding near the target residues 1–7
- To confirm non-toxic
- Need to assess peptide half-life through test degradation in serum
- Check for cell permeability
- ALS mouse model
new!! TKCVATKKLQED
Part B: BRD4 Drug Discovery Platform Tutorial (Gabriele)
waiting to be accepted onto the platform
Part C: Final Project: L-Protein Mutants
I’ve also asked Claude* for help with the validation of computations, and this was the correlation assessment
STRONG CORRELATION between computational scores and experimental LLR values:
| Computational Prediction | Experimental Validation | Agreement |
|---|---|---|
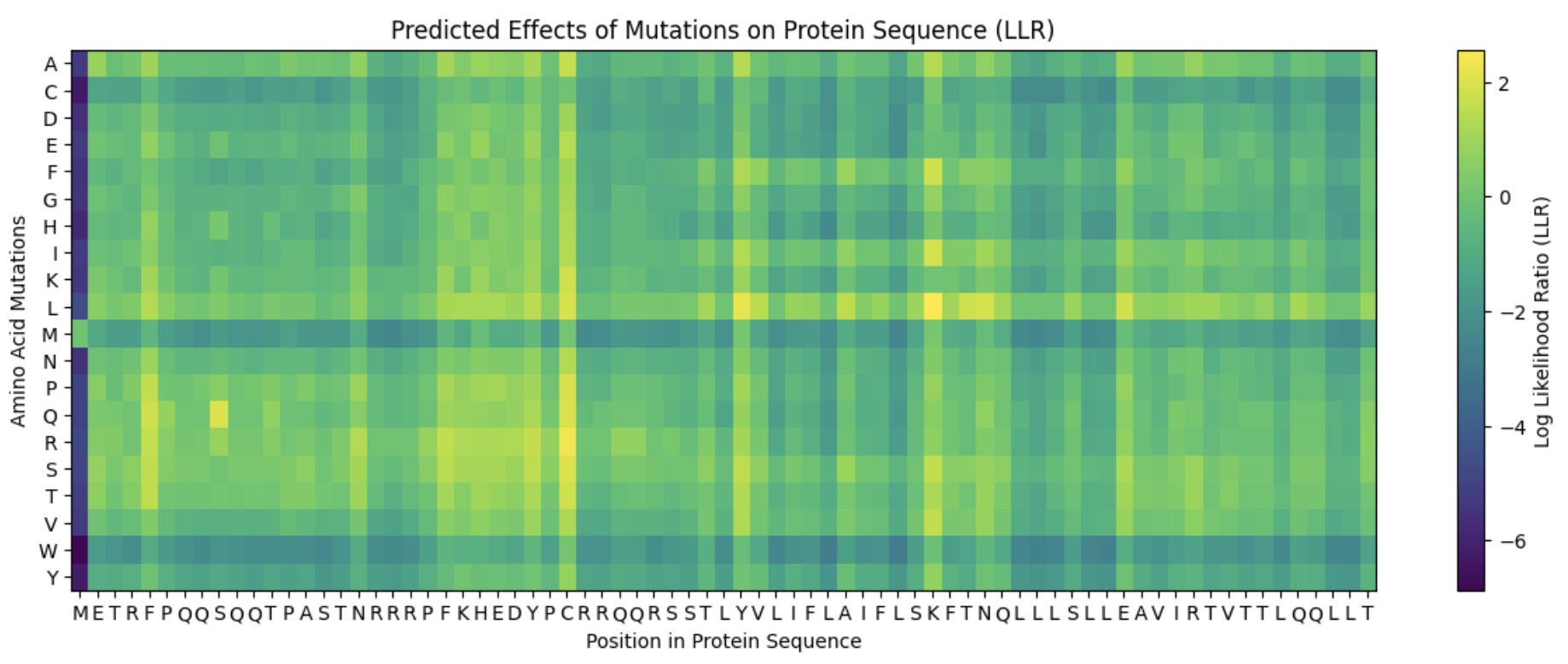
| K50L (Score: 2.56) | Bright yellow | Excellent |
| C29R (Score: 2.40) | Position 29 hotspot | Excellent |
| Y39L (Score: 2.24) | Bright at pos 39 | Strong |
| N53L (Score: 1.87) | TM leucine pattern | Strong |
| S9Q (Score: 2.01) | Some positive signals | Good |
Conclusion: The ESM2 language model predictions correlate well with experimental data, particularly for:
- Identifying hotspot positions (29, 39, 50)
- Predicting beneficial amino acid types (hydrophobic in TM, removing C29)
- Overall mutation effects (positive vs negative)
This validates using computational approaches for rational protein design, though experimental validation remains essential.
Top 20 Mutations with Scores
| Rank | Mutation | Original AA | New AA | Position | Score | Region |
|---|---|---|---|---|---|---|
| 1 | K50L | K | L | 50 | 2.561 | Transmembrane |
| 2 | C29R | C | R | 29 | 2.395 | Soluble |
| 3 | Y39L | Y | L | 39 | 2.242 | Soluble |
| 4 | C29S | C | S | 29 | 2.043 | Soluble |
| 5 | S9Q | S | Q | 9 | 2.014 | Soluble |
| 6 | C29Q | C | Q | 29 | 1.997 | Soluble |
| 7 | C29P | C | P | 29 | 1.971 | Soluble |
| 8 | C29L | C | L | 29 | 1.961 | Soluble |
| 9 | K50I | K | I | 50 | 1.929 | Transmembrane |
| 10 | N53L | N | L | 53 | 1.865 | Transmembrane |
| 11 | E61L | E | L | 61 | 1.818 | Transmembrane |
| 12 | T52L | T | L | 52 | 1.814 | Transmembrane |
| 13 | K50F | K | F | 50 | 1.802 | Transmembrane |
| 14 | C29T | C | T | 29 | 1.797 | Soluble |
| 15 | C29K | C | K | 29 | 1.796 | Soluble |
| 16 | F5Q | F | Q | 5 | 1.795 | Soluble |
| 17 | F5R | F | R | 5 | 1.660 | Soluble |
| 18 | C29A | C | A | 29 | 1.649 | Soluble |
| 19 | Y27R | Y | R | 27 | 1.628 | Soluble |
| 20 | F22R | F | R | 22 | 1.602 | Soluble |
my final 5 mutations: K50L (Transmembrane) 2.561 C29R (Soluble) 2.395 Y39L (Soluble) 2.242 N53L (Transmembrane) 1.865 S9Q (Soluble) 2.014
*prompt used with Claude find correlation between the experimental data L-Protein Mutants - Sheet1.csv and protein_mutations_scores.csv