Week 2-DNA Read, Write, & Edit
title: ‘Week 2’ weight: 10
Part 1: Benchling & In-silico Gel Art
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks. You might find Ronan’s website a helpful tool for quickly iterating on designs!


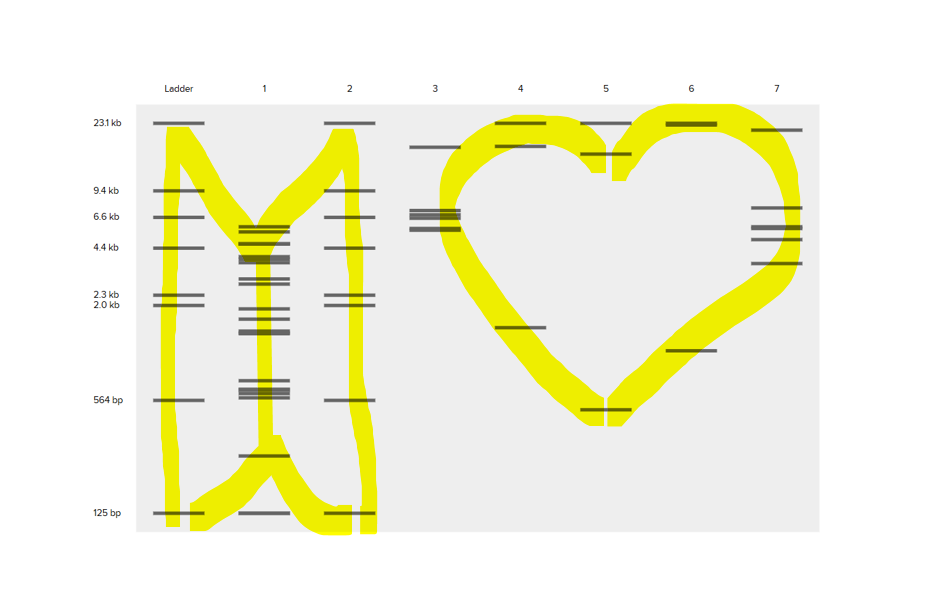

Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
I didn’t have acces to a lab ,so I studied the protocol.
Part 3: DNA Design Challenge
3.1. Choose your protein.
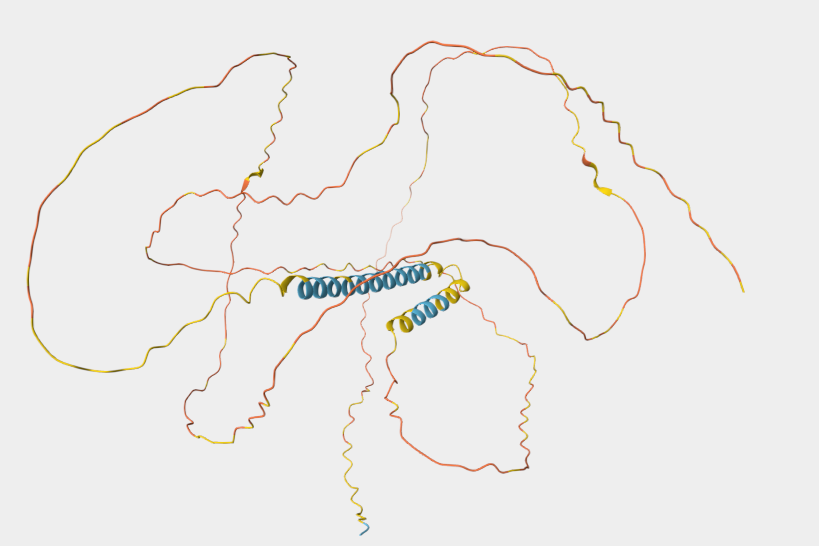

I choose Dsup-Damage suppressor protein from Ramazzottius varieornatus-tardigrade, because Dsup is a fascinating protein, unique to extremophile tardigrades, which binds to nucleosomes and protects DNA from damage caused by radiation , desiccation and reactive oxigen species.This makes it highly relevant for applications in biotechnology, such as enhancing radiation resistance in cells for space exploration, cancer therapy, or environmental remediation. It’s also an intrinsically disordered protein, offering insights into novel DNA-protection mechanisms beyond traditional repair pathways.
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
I used uniprot to find Dusp protein amino acid sequence. uniprot-P0DOW4
This is the protein sequence:MASTHQSSTEPSSTGKSEETKKDASQGSGQDSKNVTVTKGTGSS ATSAAIVKTGGSQGKDSSTTAGSSSTQGQKFSTTPTDPKTFSSDQKEKSKSPAKEVPS GGDSKSQGDTKSQSDAKSSGQSQGQSKDSGKSSSDSSKSHSVIGAVKDVVAGAKDVAG KAVEDAPSIMHTAVDAVKNAATTVKDVASSAASTVAEKVVDAYHSVVGDKTDDKKEGE HSGDKKDDSKAGSGSGQGGDNKKSEGETSGQAESSSGNEGAAPAKGRGRGRPPAAAKG VAKGAAKGAAASKGAKSGAESSKGGEQSSGDIEMADASSKGGSDQRDSAATVGEGGAS GSEGGAKKGRGRGAGKKADAGDTSAEPPRRSSRLTSSGTGAGSAPAAAKGGAKRAASS SSTPSNAKKQATGGAGKAAATKATAAKSAASKAPQNGAGAKKKGGKAGGRKRK
I reversed translated the amino acid sequnce into nucleotide sequence.
This is the DNA sequence: atggcatcca cacaccaatc atccacagaa ccctcttcca caggtaaatc tgaggaaacg 61 aagaaagatg cttcgcaagg gagcgggcaa gactccaaga acgtaaccgt taccaaaggt 121 accggttcct ccgccacctc agctgccatt gtcaagacag gaggatccca aggcaaagat 181 tcctctacta cagcgggctc ttctagtact cagggacaga agttcagtac tacacctacc 241 gacccgaaaa ctttcagctc tgaccaaaag gagaaatcca aaagcccagc caaagaagtc 301 ccgtctggtg gcgatagtaa gtcccaaggt gacaccaagt ctcaaagcga cgccaaatct 361 tctggacaaa gtcagggcca gtctaaagac agcggcaaat catcttccga cagtagcaag 421 agtcactctg tcatcggagc tgtcaaagac gtcgttgcag gcgccaaaga tgtcgcagga 481 aaagccgtcg aggatgctcc tagcatcatg catactgcag tcgatgctgt gaagaacgca 541 gccacgactg tgaaggatgt ggcatcgtcg gctgcatcga ctgtggcgga gaaggtagtc 601 gatgcttacc acagtgtggt gggagacaag acggacgaca agaaagaggg cgagcacagc 661 ggcgacaaga aggacgactc caaagctgga agtggctctg gacaaggtgg tgacaacaag 721 aagtctgaag gagagacttc tggccaagca gaatccagct ctggcaacga aggagctgct 781 ccagccaaag gccgtggtcg tggacggcct ccagcagctg ctaaaggagt tgctaagggt 841 gctgcaaagg gcgctgccgc ctccaaagga gccaagagcg gtgctgaatc ctccaaggga 901 ggagaacagt cgtcaggaga tatcgagatg gcagatgctt cctccaaggg aggctcggac 961 cagagggatt ccgcggcgac cgttggcgaa ggtggtgcat caggcagtga gggtggagct 1021 aagaaaggca gagggcgggg cgctggtaag aaagcggatg cgggtgatac gtccgctgag 1081 ccgcctcggc ggtcgtcccg cctgacgtct tcaggtacag gggcgggttc cgctccagct 1141 gcagcgaaag gcggagcgaa gcgtgctgct tcttcctcca gtacaccttc caacgctaag 1201 aagcaagcga ctggaggtgc tggcaaagct gctgccacca aagcaactgc tgccaaatcg 1261 gcagcctcta aagctcccca gaatggcgca ggtgccaaga agaagggagg aaaggctgga 1321 ggacggaaga ggaagtaa
3.3. Codon optimization.
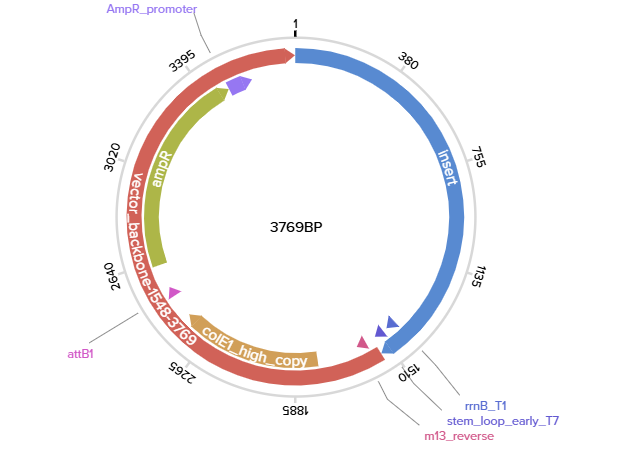

3.4. You have a sequence! Now what?
To produce the Dsup protein from this DNA, technologies leveraging the central dogma (DNA → RNA → protein) can be used. I'll describe both cell-dependent and cell-free methods.
Cell-dependent (in vivo): Insert the optimized DNA into an expression vector (e.g., pET-28a with T7 promoter). Transform the plasmid into E. coli (e.g., BL21(DE3) strain) via electroporation or heat shock. Induce expression with IPTG, which activates the T7 RNA polymerase to transcribe the DNA into mRNA. The mRNA is then translated by ribosomes using tRNAs to assemble the protein. Harvest cells, lyse them, and purify Dsup (e.g., via His-tag affinity chromatography if added). Cell-free (in vitro): Use a cell-free expression system (e.g., PURExpress or wheat germ extract). Transcribe the DNA into mRNA using T7 RNA polymerase, NTPs, and buffers. Then, add the mRNA to a translation mix containing ribosomes, tRNAs, amino acids, energy sources (ATP/GTP), and cofactors. Translation occurs directly, producing the protein without cells. This is faster for screening but lower yield. Both methods allow scalable production, with cell-dependent being cheaper for large quantities.
3.5.How does it work in nature/biological systems?
Describe how a single gene codes for multiple proteins at the transcriptional level. Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!! See example below.
In nature, the Dsup gene in tardigrades is transcribed into mRNA in the nucleus under stress conditions (e.g., radiation). The mRNA is exported to the cytoplasm, where it’s translated into the Dsup protein by ribosomes. Dsup then localizes to the nucleus via its nuclear localization signal, binding nucleosomes to shield DNA. For alignment (using Benchling-style visualization): DNA: atggcatccacacaccaatcatcc… (original sequence as in 3.2) RNA: auggcauccacacaccaaucaucc… (T → U) Protein: M A S T H Q S S T E P S… (each codon triplet → 1 AA) This shows the flow: DNA transcribed to RNA (complementary, T→U), RNA translated in triplets (e.g., AUG = M start). Failures like mutations could alter this, but optimization helps overcome expression issues in heterologous systems.
Part 4: Prepare a Twist DNA Synthesis Order
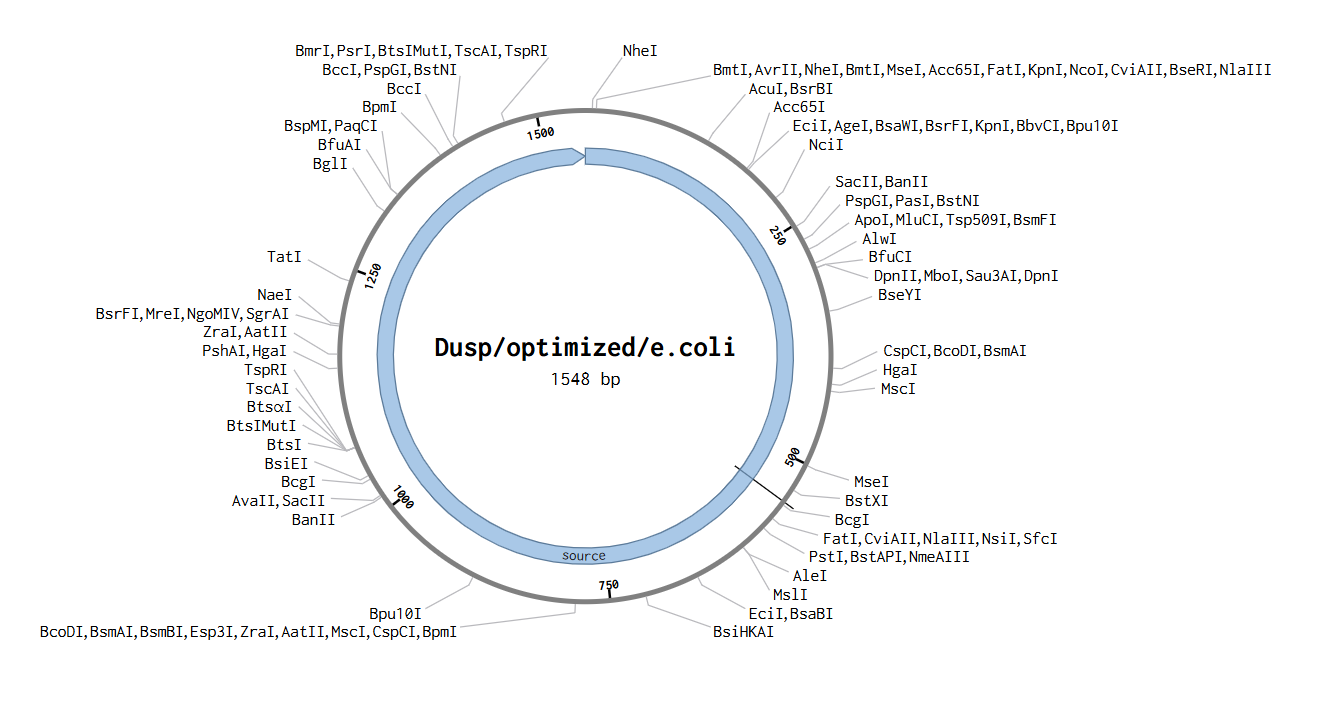

**5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would like to sequence the skin microbiome of individuals with atopic dermatitis to identify the imbalance between beneficial and harmful bacteria, which would help me in creating a personalized treatment.
(ii) What technology or technologies would you use to perform these DNA edits and why?
I would chose the third generation sequencing , because it’s essential for distinguishing between related bacterial strands in the microbiome .The essentials steps are that the DNA is passed through a nanopore and as it moves, it creates specific fluctuations in an electrical current, which are then decoded directly into the DNA sequence.The limitations are that it has a higher error rate per base compared to short-read sequencing, though high-coverage analysis successfully mitigates this. To complete this step, I must first collect and filter my raw input to remove errors. Next, categorize the remaining data into specific groups based on their shared traits. Finally, verify the results against your original requirements to ensure everything is accurate and ready for use.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
I would synthesize a Synthetic Antimicrobial Peptide (AMP) Genetic Circuit. This system uses a Quorum Sensing “switch”—a promoter that only activates when it detects high concentrations of S. aureus—linked to a gene for an Antimicrobial Peptide (like Omiganan). In patients with atopic dermatitis, the skin’s barrier is overwhelmed by harmful bacteria. By writing a circuit that only triggers in the presence of this specific pathogen, we create a personalized “search-and-destroy” therapeutic that restores microbial balance without the broad-spectrum damage of traditional antibiotics.
(ii) What technology or technologies would you use to perform this DNA synthesis and why? Also answer the following questions: What are the essential steps of your chosen sequencing methods? What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
For the synthesis, I would use Silicon-Based DNA Synthesis (Twist Bioscience). This technology uses a silicon chip to “write” thousands of different DNA sequences simultaneously, making it more scalable and cost-effective than old-fashioned plastic plate methods.
To verify that the synthesized DNA perfectly matches my design, I would use Next-Generation Sequencing (NGS). The essential steps include Library Prep , Cluster Generation , Sequencing by Synthesis (identifying bases via fluorescent flashes), and Data Analysis (reassembling the final code). While NGS is highly scalable, its main limitations are its total speed (taking 24–48 hours) and a slight dip in accuracy when encountering homopolymers.