Week 4 HW: Protein Design - Part 1
Part A: Conceptual Questions
1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
On average, if we assume that 25% of meat is protein, then we would be taking 500*0.25 = 125 g of protein intake. With an average of 100 Da per amino acid as it’s molecular weight, the total number of moles of amino acids become 1.25 moles. Multiplying with the Avagadro’s number, we get roughly seven hundred fifty-two sextillion (7.528 * 10^23) molecules !!
2. Why do humans eat beef but do not become a cow, eat fish but do not become fish?
When we eat beef or fish, our body does not absorb the protein in their native format. It degrades them to the lowest possible level: amino acids. These amino acids are what are absorbed and then reused to construct other new proteins that our body needs. Thus, we do not become what we eat, rather we become what we construct from what we eat.
3. Why are there only 20 natural amino acids?
These set of 20 amino acids are what are used by majority of the organisms on the planet. Though there isn’t a set rule, these 20 amino acids make possible all the proteins and their conformations that are required to sustain life by having & conferring suitable properties. Probably, we can argue that changing this “perfect set” of amino acids might be harmful/lethal to the organism, which would’ve been why only 20 amino acids are present.
4. Can you make other non-natural amino acids? Design some new amino acids.
We can! And this area of study is termed as Xenobiology.
For example, we can design amino acids with different side chain groups such as cyano (-CN) or a boron (-B) groups/atoms. These would enable the amino acids to bind to certain substrates more effectively, repel others, act as sensors, and open up a plethora of unique functions.
5. Where did amino acids come from before enzymes that make them, and before life started?
This is explained by the Miller-Urey experiment of abiotic synthesis. They showed that, in the “primordial soup” of early Earth, energy from lightnings, hydrothermal vents and other sources, reacted with gases such as methane, ammonia, etc. to form simple organic molecules, which could have been the earliest source of amino acids.
6. If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
D-amino acids are the “mirrored” versions of the L-amino acids. So, the steric hindrances present in the L-amino acid helices (which has a right handedness) would flip, conferring a left handedness to the D-amino acid helix.
7. Can you discover additional helices in proteins?
Yes of course! Some of them include: the 3(10)helix, which contains tighter and thinner helices; the pi helix, which is wider and shorter relative to the normal alpha helix.
8. Why are most molecular helices right-handed?
Most biological helices are right-handed due to the chirality of their L-amino acids (proteins), which cause steric clashes in their left-handed forms.The right-handed alpha-helices avoid side-chain and backbone repulsions, thus optimizing stability.
9. Why do β-sheets tend to aggregate? What is the driving force for β-sheet aggregation?
The primary driving force for aggregation of beta sheets are hydrogen bonding between the backbones and the hydrophobic effects involved. Since the edges of the sheets have available hydrogen bond donors and acceptors, they tend to form aggregates by grabbing onto other beta sheets.
Part B: Protein Analysis and Visualization
My protein of choice is the DnaA protein from Mycobacterium tuberculosis. It acts as a replication initiator, aiding in recognizing, binding and unwinding the oriC region to kick-start replication. It is absolutely essential for replication in M.tuberculosis, and without it, the bacterium will not be able to initiate replication.
I chose this particular protein, because I am currently working on it, especially with it’s DNA Binding domain (Domain 4), to screen and identify inhibitors against it. It is also a biologically sound target, since it is essential for replication and also has no eukaryotic homologs.
The amino acid sequence details:
Sequence:
sp|P9WNW3|DNAA_MYCTU Chromosomal replication initiator protein DnaA OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=dnaA PE=1 SV=1 MTDDPGSGFTTVWNAVVSELNGDPKVDDGPSSDANLSAPLTPQQRAWLNLVQPLTIVEGF ALLSVPSSFVQNEIERHLRAPITDALSRRLGHQIQLGVRIAPPATDEADDTTVPPSENPA TTSPDTTTDNDEIDDSAAARGDNQHSWPSYFTERPHNTDSATAGVTSLNRRYTFDTFVIG ASNRFAHAAALAIAEAPARAYNPLFIWGESGLGKTHLLHAAGNYAQRLFPGMRVKYVSTE EFTNDFINSLRDDRKVAFKRSYRDVDVLLVDDIQFIEGKEGIQEEFFHTFNTLHNANKQI VISSDRPPKQLATLEDRLRTRFEWGLITDVQPPELETRIAILRKKAQMERLAVPDDVLEL IASSIERNIRELEGALIRVTAFASLNKTPIDKALAEIVLRDLIADANTMQISAATIMAAT AEYFDTTVEELRGPGKTRALAQSRQIAMYLCRELTDLSLPKIGQAFGRDHTTVMYAQRKI LSEMAERREVFDHVKELTTRIRQRSKR
Length: 507 residues
The most frequent amino acid: Alanine (54 times), Leucine (46 times), threonine (42 times). (Using given colab notebook)
Number of protein sequence homologs: 250 homologs found, from using the Uniprot BLAST Tool.
Belonging to a protein family: Yes, belongs to the DnaA family.
Protein Structure in RCSB:
PDB ID: 3PVV
Resolved in: 2011
Resolved by: Tsodikov, et al..
Link to the article: Here
Resolution: 2 Angstroms (It is a good quality structure.)
The solved structure contains the protein’s DNA Binding Domain (also called as the Domain 4), bound to a MtDnaA box, that is, bound to a DNA helix.
Note: The PDB structure only has the domain 4, and not the entire protein’s structure. However, the full structure of the protein is available in the Alphafold database.
Through the search in SCOP Database, I found that the protein indeed belongs to the structure classification family of “Chromosomal replication initiation factor DnaA C-terminal domain IV”. This family currently consists of DnaA proteins from three bacteria: E.coli, M.tuberculosis and A.aeolicus.
3D visualization:
Chosen Software: PyMol
Note: Only the PDB structure has been used in this homework for visualization.
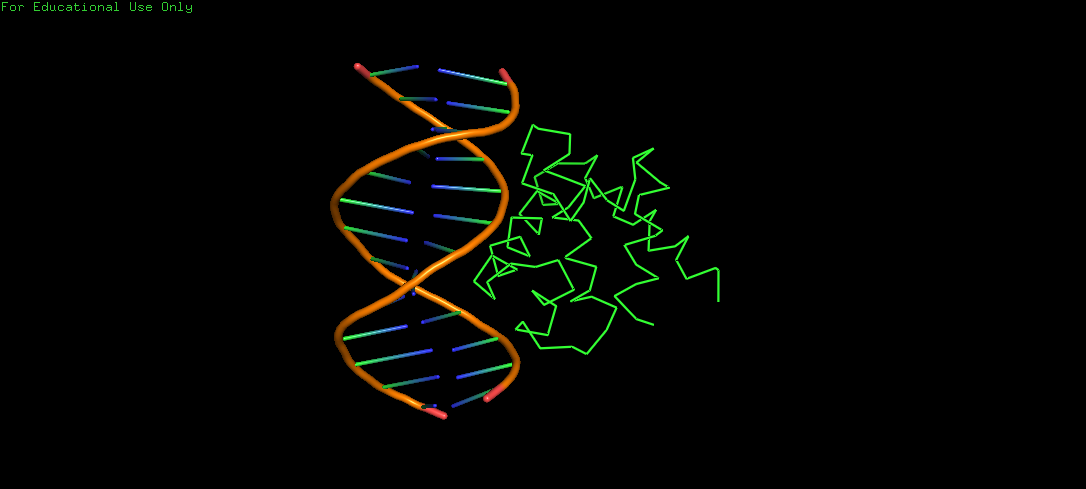
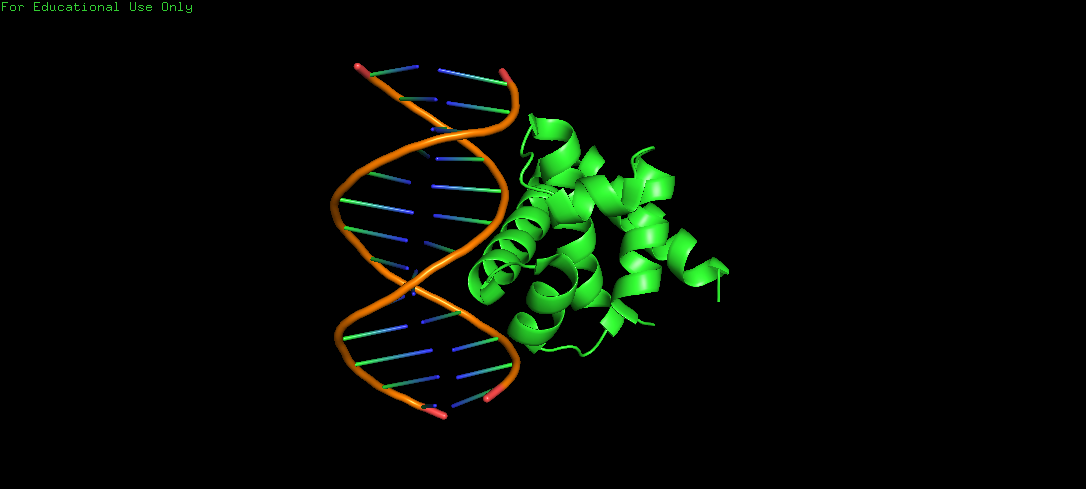
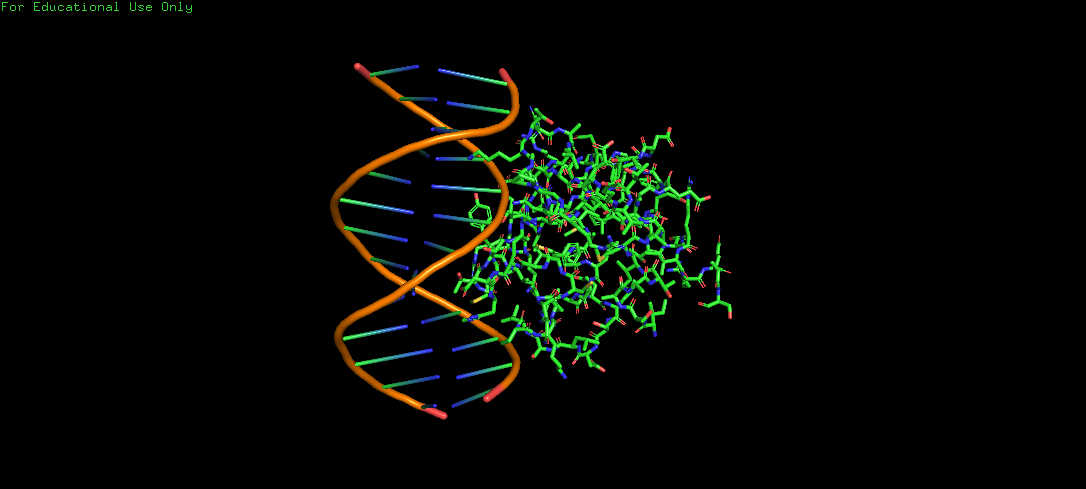
Protein Structure visualized as a Ribbon, Cartoon and Ball and Sticks.
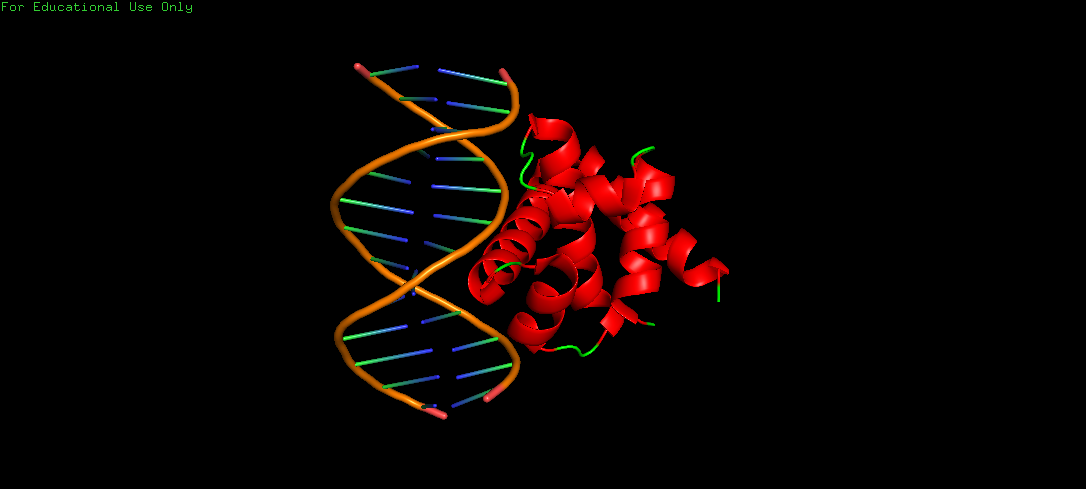

Upon colouring by secondary structure it is found to have a lot of helices. No beta sheets/strands were seen.
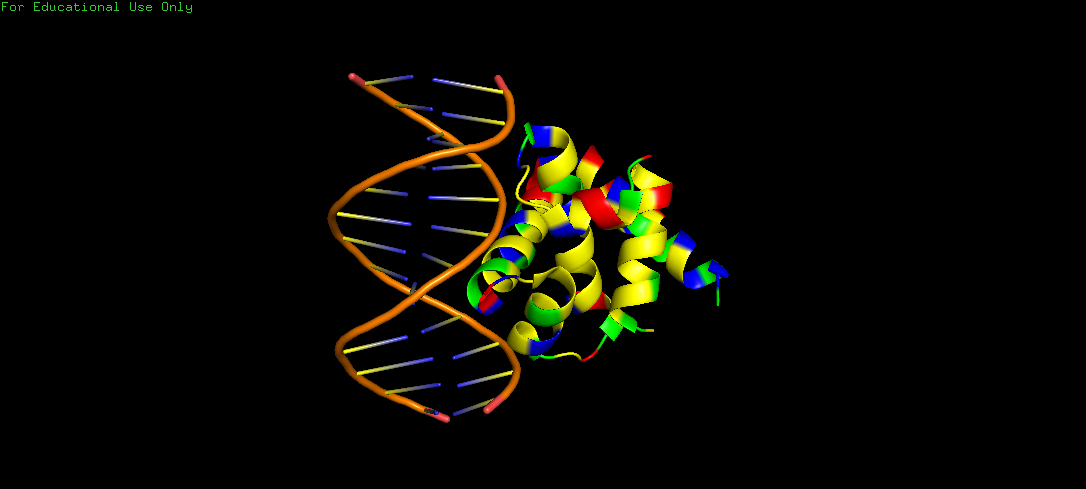

The DNA-interacting of the protein is likely enriched in hydrophilic/basic residues, enabling electrostatic attraction to the negatively charged phosphate backbone. These interactions are predominantly polar and ionic, not hydrophobic, since DNA is highly water-exposed and also charged. Whereas, hydrophobic residues are probably buried within the protein core, stabilizing its folded alpha helical structure.
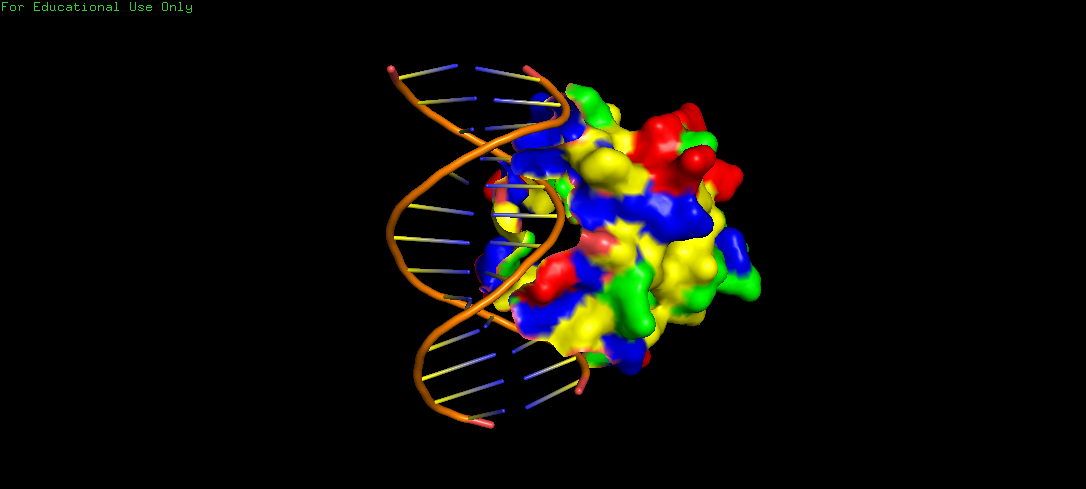

The surface representation shows visible grooves and cavities at the protein–DNA interface, suggesting potential binding pockets. These pockets appear partially recessed and contoured, which are consistent with nucleotide accommodation. Also, the clustering of varied residues around these areas indicate chemically diverse environments, suitable for specific binding to the DNA. However, visualizing binding pockets through the Alphafold structure might be more effective, since here, we can mistake the start and end of the protein sequence to be a binding pocket, when in fact, it is just the sequence and structure that has been determined.