Week 3 HW: Lab Automation
Assignment: Python Script for Opentrons Artwork
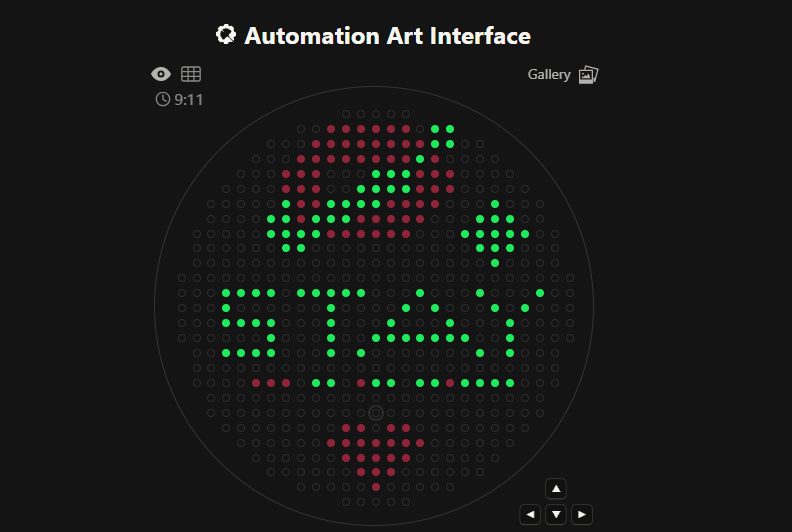

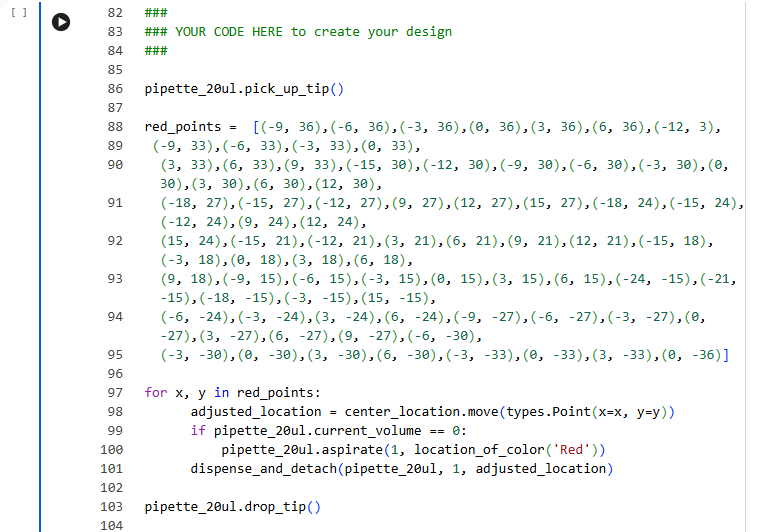
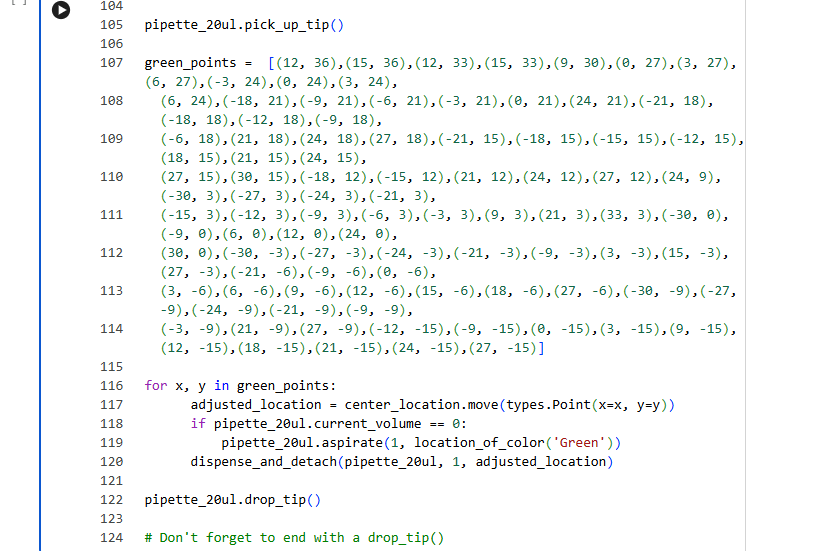
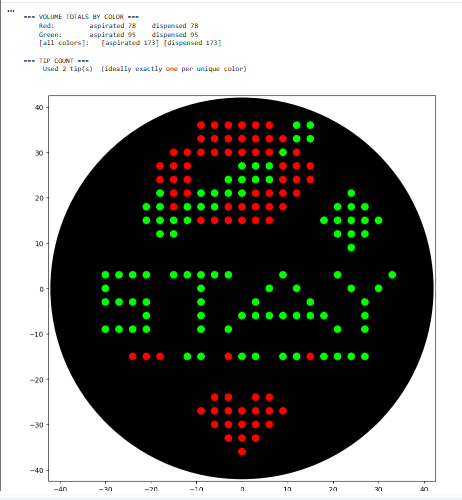
I used the GUI coordinates to prepare this code:
Post Lab Questions:
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely.
For this week, we’d like for you to do the following:
Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
While your description/project idea doesn’t need to be set in stone, we would like to see core details of what you would automate. This is due at the start of lecture and does not need to be tested on the Opentrons yet.
My chosen paper: https://doi.org/10.1016/j.slast.2023.08.007
Title: TidyTron: Reducing lab waste using validated wash-and-reuse protocols for common plasticware in Opentrons OT-2 lab robots.
Year of Publication: April 2024.
This paper introduces TidyTron, which is an open-source library of automated protocols that are designed for the Opentrons OT-2 lab robot. It’s aim is to decontaminate and reuse single-use plasticware, such as micropipette tips and microtiter plates. This system addresses the environmental and economic burden of biotechnology research, which generates approximately 5.5 million tons of plastic waste annually. By automating the cleaning process, TidyTron provides a high-throughput, consistent cleaning alternatives compared to manual decontamination, which is often impractical due to variable cleaning quality.
The researchers developed and validated these protocols by testing various cleaning solutions, contact times, and agitation methods to balance the cleaning stringency with time and cost efficiency. The study especially focused on removing common lab contaminants, such as DNA, E. coli, and S. cerevisiae (yeast). It was then validated by comparing the performance of “fresh” (first-time use) plasticware against the cleaned items using strict metrics: colony-forming units (CFU) for bacteria, qPCR for DNA concentration and amplification efficiency, and flow cytometry for yeast cell counts and debris.
Specific cleaning sequences, such as one bleach wash followed by two hydrogen peroxide rinses with a final water rinse, rendered cleaned tips indistinguishable from fresh ones in terms of bacterial carry-over. In addition, the protocols were shown to effectively remove DNA templates without causing significant inhibition of subsequent qPCR reactions. Economic and environmental modeling over a seven-year period suggests that implementing TidyTron (e.g., reusing tips upto four times) can lead to significant financial savings and drastic reductions in plastic consumption compared to standard single-use practices. The protocols and graphical user interface are also available as open-source resources.
I chose this paper because it addressess a major concern of mine: plastic wastage. This addresses and provides a cheap, sustainable and efficient solution to combat this “norm” in biotech, thereby pushing us, one step towards a pollution free world.
Automation Tools:
- Computational Design Automation
DNA construct design and management - Automated plasmid design, annotation, and version control using platforms such as Benchling.
Genetic logic compilation - Translation of Boolean circuit designs into DNA sequences using Cello.
- Robotic DNA Assembly
Automated liquid handling - Robotic setup of cloning reactions (PCR) using systems such as Opentrons OT-2.
Standardized reaction execution - Reduction of human variability through precise automated pipetting.
- Robotic Workflow
Automated transformation and plating - Programmed execution of transformation and colony processing steps.
Colony picking and culture inoculation - Robotic selection and transfer of clones for downstream processing.
- Software Control
Digital-to-robot execution - Direct export of design files into robotic instruction scripts.
Automated experimental planning - Software-driven scheduling and build instruction generation.
Iterative optimization - Data-informed redesign cycles enabling continuous Design–Build–Test–Learn refinement.
Thank You!