Week 1 HW: Principles and Practices
NiMap-S: An Engineered Streptomyces System for Mapping and Monitoring Nickel Contaminated Areas
In this project I propose NiMap-S, a synthetic microbial biosensor system designed to identify local patterns of nickel contamination in soil. The system is based on a genetically modified, soil native Streptomyces strain that functions as a biological reporter. Streptomyces is selected because it is already well adapted to soil environments and because its rich secondary metabolism allows sensing and reporting functions to be integrated without introducing a completely foreign organism into the soil ecosystem. The engineered bacteria are intended to be physically contained within biodegradable biobeads, allowing controlled interaction with the surrounding soil while limiting dispersal.
In NiMap-S, bioavailable nickel ions (Ni2+) are detected through a nickel-responsive genetic circuit in which metal sensing is coupled to regulated expression of a methyl salicylate biosynthesis pathway. As local nickel availability increases, the circuit drives a corresponding increase in the production and release of methyl salicylate (C8H8O3), a volatile compound commonly involved in plant signalling. This chemical output is chosen intentionally, as it enables nondestructive sensing and creates a functional link between microbial detection and existing plant-soil communication processes.
One of the main limitations of phytoremediation is limited information on how contaminants are distributed across an area. Soil contamination is spatially heterogeneous, yet existing analysis methods rely on destructive sampling, slow laboratory workflows, limited information, and high costs when applied at field scale. As a result, remediation strategies are often applied uniformly, even when contamination varies sharply across short distances.
NiMap-S is therefore conceived as a premapping system that operates before phytoremediation plants, such as sunflowers, are introduced. The system generates a chemical heat map of bioavailable nickel hotspots across the field. These signals can be detected using low-altitude drone platforms equipped with gas-sensing instruments that are already used in environmental monitoring. Importantly, the choice of methyl salicylate avoids reliance on speculative or custom built detection hardware, grounding the system by utilizing approaches that are already in practical use.
NiMap-S makes it possible to adjust remediation inputs with greater precision, for example by applying chelators only where and in the amounts needed, selecting appropriate plant varieties for specific locations, reducing unnecessary inputs, and overall remediation costs.
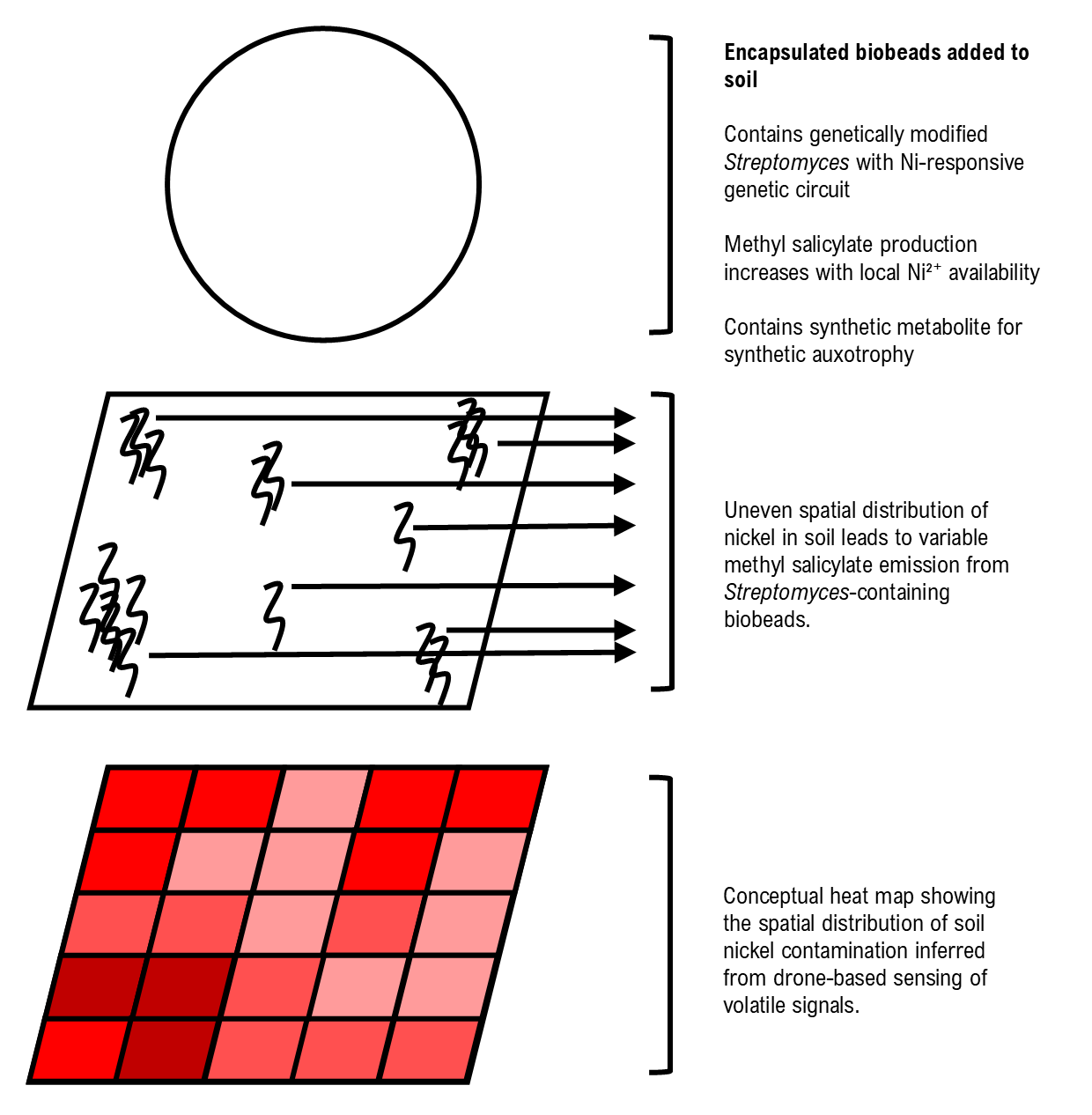
 Conceptual overview of the NiMap-S system
Conceptual overview of the NiMap-S system
Governance Goals
Goal 1: Providing Biosecurity
The first policy goal is to prevent biological and ecological harm resulting from the environmental use of engineered microorganisms. Since NiMap-S is intended to operate outside controlled laboratory settings, biosafety cannot depend solely on correct use or post application control. Instead, safety needs to be built directly into the biological design of the organism and the material design of the system.
Sub-goal 1.1: Limiting persistence
Engineered microorganisms should not remain active beyond their intended use. NiMap-S is therefore designed with the assumption that environmental presence should be temporary, so that microbial activity naturally ends once sensing objectives have been achieved.
Sub-goal 1.2: Preventing uncontrolled spread and evolution
The system should not spread beyond its area of application or adapt in ways that increase ecological risk. This subgoal focuses on containment strategies that restrict dispersal and reduce opportunities for long-term evolution in open environments.
Sub-goal 1.3: Ensuring reversibility
Environmental application should remain reversible. If unexpected effects occur, the system should allow for intervention.
Goal 2: Data Transparency and Responsible Governance
The second policy goal addresses how environmental data generated by NiMap-S is managed and used. Because such data can shape land-use application choices and remediation strategies, data governance is not only a technical issue but also an ethical one.
Sub-goal 2.1: Decision support with transparency
Data produced by NiMap-S should support informed actions, not surveillance or enforcement. Measurement goals, uncertainty, and technical limits should be communicated clearly to public institutions and local stakeholders.
Sub-goal 2.2: Shared access and control
Environmental data should remain accessible to public agencies, municipalities, and communities. Governance structures should avoid exclusive control by large corporate actors and instead support broader access to environmental knowledge.
Goal 3: Environmental Protection and Ecosystem Integrity
The third policy goal is to ensure that NiMap-S contributes directly to environmental recovery while respecting ecosystem integrity. Here, sensing is treated not as an end in itself, but as a tool to support remediation efforts that reduce contamination without creating additional ecological disturbance. Here, environmental protection is understood not just as avoiding harm, but as actively enabling soil systems to recover.
Sub-goal 3.1: Supporting effective remediation through phytoremediation
NiMap-S should strengthen phytoremediation by improving how contamination is identified and spatially mapped. By providing information on bioavailable nickel at appropriate scales, the system allows remediation efforts to be applied more efficiently, increasing cleanup effectiveness while reducing unnecessary use of water, chemicals, and land.
Sub-goal 3.2: Protecting ecosystem integrity during intervention
Remediation should reduce pollution without introducing new ecological risks. This includes limiting disturbance to surrounding soil, plants, and microbial communities, and ensuring that technological elements of the system remain compatible with ecosystem protection. In this sense, environmental protection overlaps with biosecurity, as containment and reversibility also help safeguard broader ecosystem functioning.
Goal 4: Equitable Access, Shared Benefits, and Collaborative Use
The fourth policy goal is to ensure that NiMap-S benefits not only the environment, but also the people whose land and livelihoods are affected by soil contamination. In agricultural contexts, pollution can reduce productivity in ways that are not always obvious, and these losses are often attributed to other causes. By improving access to environmental information, the system aims to support environmental recovery and agricultural viability together.
Sub-goal 4.1: Linking environmental recovery with agricultural productivity
NiMap-S should help farmers and local land managers recognise when contamination contributes to yield loss and use this information to guide remediation, crop choice, and land use. By enabling earlier and more targeted phytoremediation, the system supports environmental improvement while also helping maintain or restore agricultural productivity and local economic resilience.
Sub-goal 4.2: Supporting coordinated action in practice
NiMap-S is intended to operate within real world soil management and remediation practices involving researchers, farmers, local authorities, and oversight bodies. Rather than functioning as a stand alone technology, the system supports collaboration at the field level, where scientific knowledge, local experience, and regulatory oversight intersect. This subgoal emphasises shared use and responsibility.
Actions
Action 1: Physical Encapsulation via Biobeads for Environmental Application
Purpose
This action proposes requiring biodegradable biobead encapsulation as a condition for environmental use, limiting microbial dispersal while still allowing interaction with the surrounding soil.
Design
Biosafety regulators would define biobead encapsulation as part of environmental approval processes. Researchers and organisations applying NiMap-S would integrate bead-compatible designs early on, while materials scientists establish standards for bead composition, degradation behaviour, and diffusion properties.
Assumptions
This action assumes that encapsulation meaningfully limits dispersal under field conditions such as rainfall and soil movement, and that bead degradation can be reasonably predicted across environments. It also assumes that physical confinement does not substantially reduce sensing performance.
Risks of Failure and Success
Failure could occur if beads degrade faster than expected or if physical containment leads to reduced attention to genetic safeguards. At the same time, widespread success could normalise environmental microbial applications before broader governance frameworks fully mature.
Action 2: Synthetic Auxotrophy
Purpose
This action introduces synthetic auxotrophy as a licensing requirement, ensuring that engineered organisms cannot persist without an added metabolite.
Design
Regulatory bodies would define acceptable synthetic metabolites and testing standards, while academic laboratories and companies engineer dependencies into essential metabolic pathways. Funding agencies could reinforce this approach by prioritising projects that demonstrate stable auxotrophy.
Assumptions
This action assumes that evolutionary escape remains rare and that synthetic metabolites are not present in natural environments.
Risks of Failure and Success
Failure could occur through genetic bypass or unintended environmental availability of the metabolite. Successful implementation may also reduce incentives to develop additional containment strategies or disadvantage smaller research groups.
Action 3: Public Interest Data Stewardship
Purpose
Environmental biosensing data is often controlled by the organisation developing the system, which limits transparency, public oversight, broader access to environmental information. This action reframes NiMap-S data as a shared public tool that supports learning, wide implementation, transparency, and accountability.
Design
Environmental approval or public funding would be linked to compliance with open data standards. Researchers and organisations would document uncertainty, calibration assumptions, and known limitations of the measurements. Public agencies or municipalities would act as data stewards, ensuring that data remains accessible to other researchers, local authorities, and communities, and that it is presented in formats suitable for nonspecialist use as well as expert analysis.
Assumptions
This action assumes that wider access to environmental data helps reduce power imbalances, supports more informed decisions, and enables environmental protection beyond individual application sites. It also assumes that public institutions have, or can develop, the capacity to manage, contextualise, and communicate complex biological data responsibly.
Risks of Failure and Success
Failure could result in poorly documented or misinterpreted data, undermining trust or leading to inappropriate conclusions. At the same time, success may lead to data being overinterpreted, selectively reused, or applied outside its original context. These risks make clear metadata, uncertainty reporting, and active stewardship essential.
Action 4: Equity-Oriented Application and Incentives
Purpose
Environmental sensing technologies are often used where financial return is highest, not where environmental need is most urgent. In agricultural contexts, soil contamination can quietly reduce yields without farmers clearly identifying pollution as the cause. This action focuses on prioritising deployment in polluted or underserved areas, where both ecological damage and hidden productivity losses are likely, and where access to monitoring tools is limited.
Design
Public funding agencies and local authorities would steer application through targeted incentives. Municipalities and land managers would help identify relevant sites, while developers adapt the system to work under low-resource agricultural conditions. Direct involvement of farmers ensures that the system supports common implementation.
Assumptions
This action assumes that delayed intervention is often driven by lack of information, and that farmers and local actors are more likely to engage when monitoring is clearly connected to soil health and productivity.
Risks of Failure and Success
Incentives might not be strong enough to compete with profit goals, and experiments could be placed in sensitive areas without good supervision. However, if the system works properly, it can detect contamination earlier and help protect both the environment and farming.
Scoring of Each Governance Action Against Rubric of Policy Goals
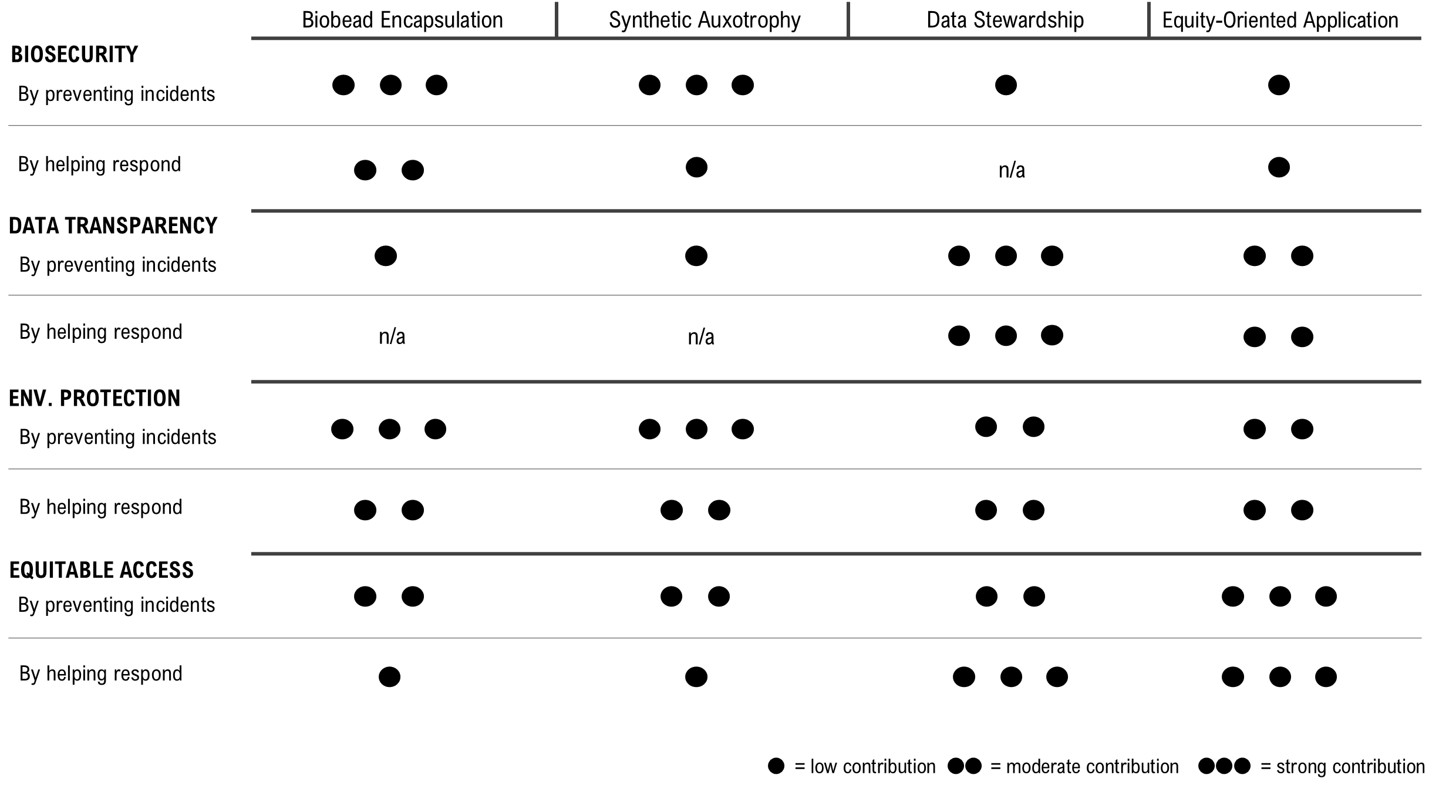

Prioritisation of Governance Actions
Based on my scoring matrix, I prioritise some governance actions as a package. NiMap-S only makes sense if biosafety and data practices are considered together. In this framing, Actions 1 and 2 define how the system can be used safely in the environment, while Action 3 shapes how the system and its data are made transparent and accountable beyond the laboratory.
Actions 1 and 2 should be implemented together from the outset. Biobead encapsulation limits physical spread, while synthetic auxotrophy reduces the risk of longterm persistence. Together, they lower the chance that a single failure leads to lasting ecological harm. The tradeoff is additional technical work and validation, which may slow early deployment but is necessary in open soil environments where engineered microorganisms raise legitimate concern. The key assumption here is that these biosafety features continue to work under real environmental conditions, including stable biobead encapsulation and effective auxotrophy, without rapid breakdown or unexpected behaviour in soil.
Action 3 focuses less on biological design and more on how information produced by NiMap-S is handled. Being clear about how the system works, its limits, its outputs, and remaining uncertainties helps justify the environmental use of engineered bacteria, while open and shared access to information on nickel contamination and NiMap-S outputs supports wider use and informed decision making by regulators, implementers, local authorities, and the public. This level of transparency can also lower barriers to environmental approval. The tradeoff is coordination effort, as preparing data and documentation for public use requires time and institutional capacity. This action assumes that the use of NiMap-S is approved by relevant public institutions and that transparency helps maintain this approval. Transparency also requires careful data stewardship to avoid misuse or unintended consequences.
These recommendations can be directed to national environmental and agricultural agencies, together with municipal authorities in contaminated regions. These actors are best positioned to align biosafety rules, shared data practices, and incentive structures, and to turn NiMap-S from a laboratory-based conceptual system into an applicable environmental tool.
Thought Process
I initially considered using a luminescence-based reporter, since optical signals are common in microbial biosensing. However, I moved away from this option because soil depth and opacity make luminescence difficult to recover without disturbing the soil.
To overcome this, I briefly explored a retrievable bead system, where luminescent beads could be pulled out of the soil for external reading. While this addressed signal visibility, it raised practical concerns such as mechanical fragility, risk of damage during retrieval, and the need for repeated manual inspection. I therefore abandoned luminescence as a reporting strategy, while keeping biobeads as a containment and application element in the final system.
I also considered electrochemical reporting, but did not pursue it further because I could not clearly conceptualise a reliable setup for heterogeneous soil environments within the scope of this project.
Finally, I decided to use a volatile chemical signal. I first considered geosmin, a naturally produced Streptomyces compound that could be placed under nickel-dependent regulation. However, because many native soil bacteria already produce geosmin, I was concerned about background signal and ambiguity. I therefore selected methyl salicylate, which provides a stronger and more distinguishable signal and can be more clearly linked to nickel-induced stress. An additional reason for this choice was the possibility that methyl salicylate release could support systemic acquired resistance (SAR) in remediation plants during phytoremediation.
AI Statement
I used Chat GPT 5.2 to check the technical feasibility of my ideas and to summarize some reading documents for the assignment. I used it to ask specific questions about the practicality of biobeads in soil, drone sensor sensitivity (like ppb-level detection of methyl salicylate), and wind interference. It was also helpful for reviewing my governance and actions framework, checking if my goals were consistent and if the roles for different actors were realistic. While using AI, I asked for reasoning or references when needed to check whether the answers made sense.
AI was used for grammar corrections, technical terminology, especially in the governance goals and actions sections, and sometimes as a writing support tool based on outlines and sentence fragments, which I prepared derived from my ideas.
DNA Read, Write, and Edit
Faculty Questions
Homework Questions from Professor Jacobson:
Q1: Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
Human genome length is 3.2 Gbp (Slide 10). Since 1 Gbp=109 bp, the human genome is 3.2×109 bp. Because DNA polymerase can make mistakes approximately every 106 bp (Slide 8), this makes the human genome prone to thousands of potential mistakes (3×109/106≃3000) during replication. In order to deal with this discrepancy, mismatch repair systems (Slide 14) detect and fix replication errors that escape polymerase proofreading.
Q2: How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
The number of average base pairs in a human protein is around 1036 bp (Slide 6), which corresponds to approximately 1036/3≃345 amino acids. Depending on the amino acid, each one can be coded by more than one codon (for example, leucine and serine have 6 codons, alanine and glycine have 4 codons, while methionine and tryptophan have only 1 codon). Because of this, depending on the amino acid composition of the protein, there can be many different DNA coding combinations for the same protein sequence. In practice, not all coding sequences work because different DNA sequences can have very different physical properties. Factors such as GC content (Slide 39) can interfere with DNA synthesis and proper expression by affecting secondary structure and minimum free energy. For this reason, coding sequences often need to be optimized for the host organism, since each organism has its own requirements in this respect.
Homework Questions from Dr. LeProust:
Q1: What’s the most commonly used method for oligo synthesis currently?
Modern oligonucleotide synthesis is most commonly based on phosphoramidite method (Slide 11; Hoose et al. 2023).
Q2: Why is it difficult to make oligos longer than 200nt via direct synthesis?
Since the elongation efficiency is not 100%, errors accumulate with each nucleotide addition cycle. Consequently, beyond 200 nt, the system becomes dominated by incorrectly synthesized DNA fragments. Moreover, as the chain length increases, the risks of depurination, strand breakage, and stabilized hairpin formation significantly reduce the final yield of pure DNA (Hoose et al., 2023).
Q3: Why can’t you make a 2000bp gene via direct oligo synthesis?
Even oligonucleotide fragments longer than 200 nt accumulate a high proportion of incorrectly synthesized DNA for the reasons discussed in Question 2. This means that synthesizing a 2000 bp sequence directly is even more difficult, as errors accumulate at a much higher rate and strand breakage and hairpin formation become much more likely. To build a 2000 bp gene, shorter oligonucleotide fragments must be assembled using methods such as Gibson assembly (Hoose et al., 2023)
Homework Question from George Church:
What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
Essential amino acids for all animals are: arginine, histidine, isoleucine, leucine, methionine, phenylalanine, threonine, tryptophan, valine, lysine. According to https://jurassicpark.fandom.com/wiki/Lysine_contingency, “lysine contingency” is a method of inserting a faulty enzyme gene in dinosaurs so that lysine metabolism is disrupted and dinosaurs are unable to biosynthesize the lysine they need. This was intended as a containment strategy, preventing them from surviving outside the island, because they are supposedly dependent on lysine rich plants found there. However, this logic is flawed, because in reality, like all animals, dinosaurs would not have been able to synthesize lysine in the first place. Instead, they would have obtained lysine from dietary sources. Thus, lysine dependency is a natural condition in animals, including humans, rather than an effective synthetic containment mechanism.
References
Hoose, A., et al. (2023). DNA synthesis technologies to close the gene writing gap. Nature Reviews Chemistry, 7, 144-161.
Week #2 slides from Prof. Jacobson, Dr. LeProust, and Prof. George Church (https://pages.htgaa.org/2026a/course-pages/weeks/week-01/index.html)
AI Statement
AI was used for grammatical corrections and paraphrasing ideas.