Week 2 HW: DNA Read, Write & Edit
Part 1: Gel Art
I searched for the lambda DNA sequence registered in NCBI using its RefSeq accession number and clicked import.
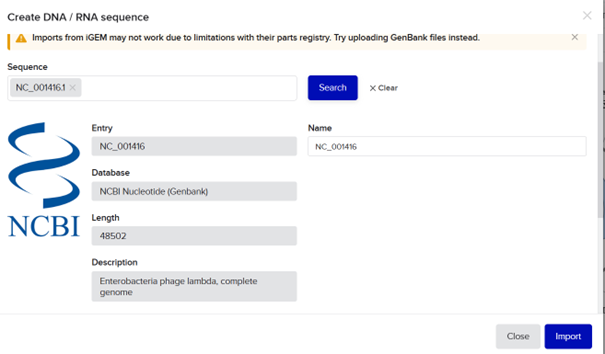

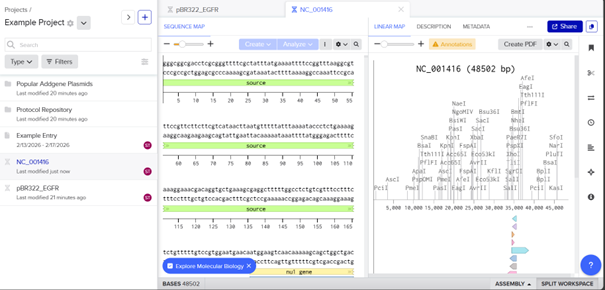

Then I selected the enzymes listed on the HTGAA site and added them to the tool.
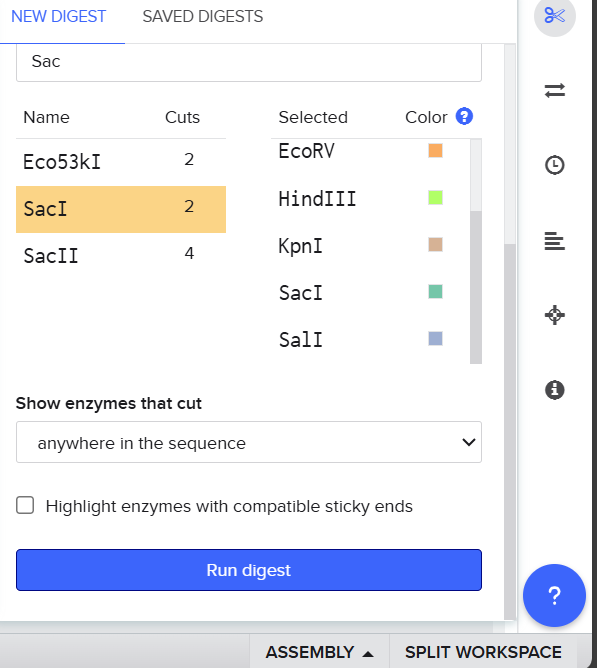

After clicking Run digest, a lot of restrictions appeared and the sequences were too short. I realized I was not supposed to select all enzymes at once because the system used all at the same time, so I ran each digestion separately.
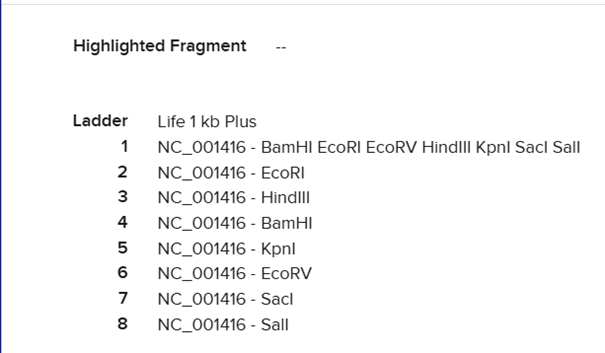


For the trident design, I planned the order as: EcoRI on the left, then KpnI, then EcoRV, followed again by KpnI and EcoRI. After that I moved to Ronan Donovan’s website to construct the gel image.
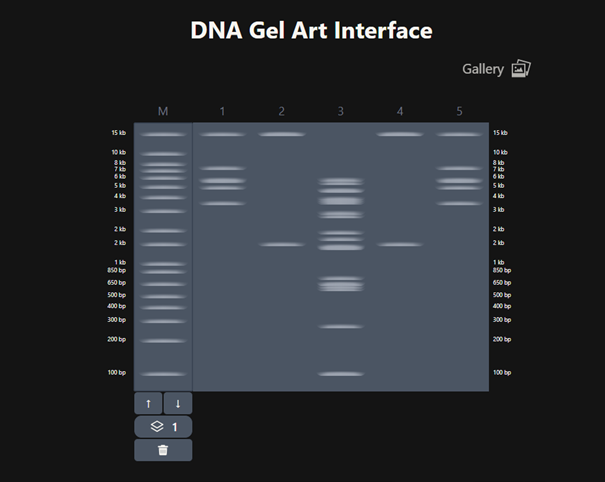


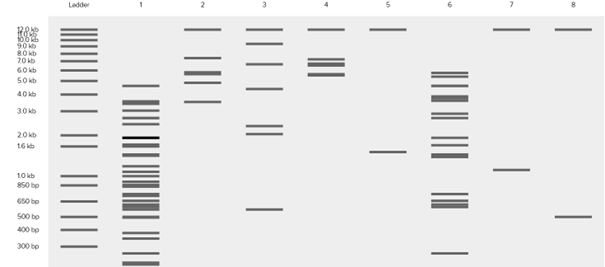
Based on these fragments, I want to create Poseidon’s trident. In Ronan Donovan’s website, I first added a ladder. According to the ladder, the gel ranges from 100 bp (bottom) to 15 kb (top). The pattern looked like this.
It does not look exactly like a trident at first, but considering that this is a gel image and that we can crop the upper part, we can say that it resembles a trident 🙂


Part 3: DNA Design Challenge
I selected NmtR (Nickel-Metal Transporter Regulator) from Streptomyces:
NmtR is a Ni/Co sensor and a transcriptional regulator (Kim et al., 2015),
In the absence of metal, it binds to DNA and represses transcription,
When Ni binds, it dissociates from DNA, allowing target genes to be expressed, with expression levels changing depending on Ni concentration,
I chose it because it is compatible with a Streptomyces chassis and minor adjustments are enough for optimization since my target organism is also a Streptomyces,
All these features make this protein suitable for the vector design I plan to build for Streptomyces coelicolor, and these selections are directly connected to the NiMap-S tool I designed in Week 1 HW
I performed my search in the NCBI database and decided to use the sequence from Streptomyces pseudoechinosporeus:

I copied the protein FASTA sequence
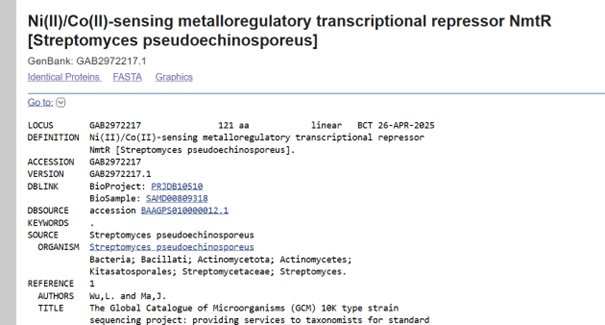




GAB2972217.1 Ni(II)/Co(II)-sensing metalloregulatory transcriptional repressor NmtR [Streptomyces pseudoechinosporeus] MHADEYAGGGGRGVLAAARLDEATAASVAATLQALATPSRLHILITLRHSPHPVGALAAAVGMEQSAVSHQLRLLRTLGLVTGRRDGRRIVYSLYDDHVAQLLDQAVHHIEHLRLGIHDPG
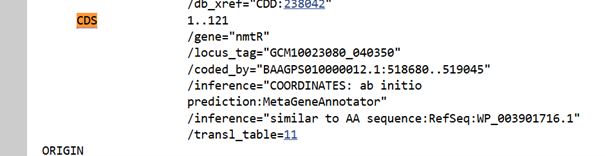



I did not need a reverse transcription tool, because I directly obtained the corresponding DNA sequence from NCBI:
BAAGPS010000012.1:518680–519045 Streptomyces pseudoechinosporeus JCM 3066 DNA, sequence012, whole genome shotgun sequence GTGCACGCCGATGAGTACGCCGGAGGCGGCGGCCGCGGCGTGCTCGCGGCCGCCCGCCTCGACGAGGCCACCGCCGCCTCCGTAGCCGCGACCCTCCAGGCCCTCGCCACCCCGTCCCGGCTGCACATCCTCATCACCCTGCGGCACTCTCCCCACCCGGTCGGCGCGCTCGCCGCCGCCGTCGGCATGGAACAGTCCGCCGTCTCCCACCAACTACGGCTTCTGCGCACCCTGGGCCTGGTCACCGGCCGACGCGACGGCCGCCGCATCGTCTACAGCCTCTACGACGACCACGTCGCCCAACTCCTCGACCAGGCCGTCCACCACATAGAACACCTGCGCCTCGGCATCCACGACCCCGGCTGA
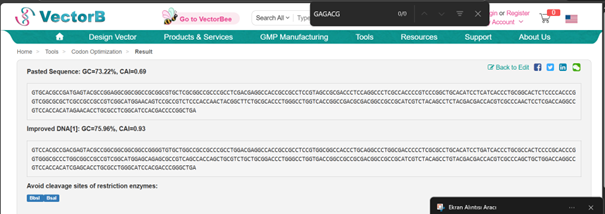

For optimization, I performed codon optimization for Streptomyces coelicolor A3(2), which is the model organism for the Streptomyces genus, using en.vectorbuilder.com. I used this platform because I forgot my IDT password and the reset email did not arrive, and I was also unable to complete the process on TWIST, I manually changed the start codon because the tool didnot recognize GTG as a start codon, which is common in Streptomyces
Optimized DNA sequence: ATGCACGCCGACGAGTACGCCGGCGGCGGCGGCCGGGGTGTGCTGGCCGCCGCCCGCCTGGACGAGGCCACCGCCGCCTCCGTGGCGGCCACCCTGCAGGCCCTGGCGACCCCCTCGCGCCTGCACATCCTGATCACCCTGCGCCACTCCCCGCACCCGGTGGGCGCCCTGGCGGCCGCCGTCGGCATGGAGCAGAGCGCCGTCAGCCACCAGCTGCGTCTGCTGCGGACCCTGGGCCTGGTGACCGGCCGCCGCGACGGCCGCCGCATCGTCTACAGCCTGTACGACGACCACGTCGCCCAGCTGCTGGACCAGGCCGTCCACCACATCGAGCACCTGCGCCTGGGCATCCACGACCCGGGCTGA
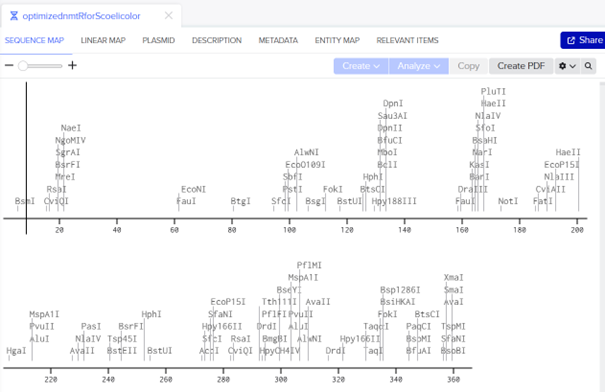

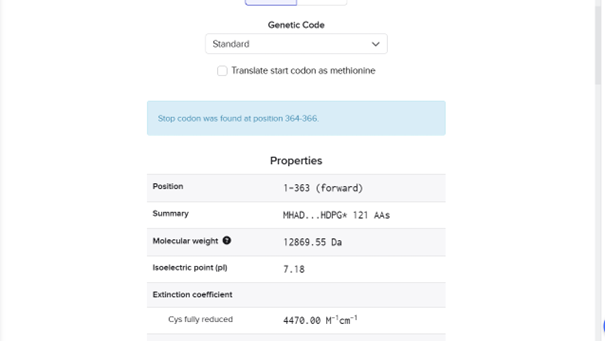

DNA sequence: ATGCACGCCGACGAGTACGCCGGCGGCGGCGGCCGGGGTGTGCTGGCCGCCGCCCGCCTGGACGAGGCCACCGCCGCCTCCGTGGCGGCCACCCTGCAGGCCCTGGCGACCCCCTCGCGCCTGCACATCCTGATCACCCTGCGCCACTCCCCGCACCCGGTGGGCGCCCTGGCGGCCGCCGTCGGCATGGAGCAGAGCGCCGTCAGCCACCAGCTGCGTCTGCTGCGGACCCTGGGCCTGGTGACCGGCCGCCGCGACGGCCGCCGCATCGTCTACAGCCTGTACGACGACCACGTCGCCCAGCTGCTGGACCAGGCCGTCCACCACATCGAGCACCTGCGCCTGGGCATCCACGACCCGGGCTGA
RNA sequence: AUGCACGCCGACGAGUACGCCGGCGGCGGCGGCCGGGGUGUGCUGGCCGCCGCCCGCCUGGACGAGGCCACCGCCGCCUCCGUGGCGGCCACCCUGCAGGCCCUGGCGACCCCCUCGCGCCUGCACAUCCUGAUCACCCUGCGCCACUCCCCGCACCCGGUGGGCGCCCUGGCGGCCGCCGUCGGCAUGGAGCAGAGCGCCGUCAGCCACCAGCUGCGUCUGCUGCGGACCCUGGGCCUGGUGACCGGCCGCCGCGACGGCCGCCGCAUCGUCUACAGCCUGUACGACGACCACGUCGCCCAGCUGCUGGACCAGGCCGUCCACCACAUCGAGCACCUGCGCCUGGGCAUCCACGACCCGGGCUGA
AA sequence: MHADEYAGGGGRGVLAAARLDEATAASVAATLQALATPSRLHILITLRHSPHPVGALAAAVGMEQSAVSHQLRLLRTLGLVTGRRDGRRIVYSLYDDHVAQLLDQAVHHIEHLRLGIHDPG
Part 4: Twist Vector Order
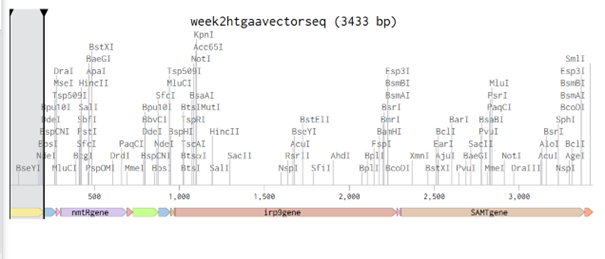
I frequently used NCBI, vectorbuilder, and the article by Bai et al. (2015) while designing this vector. The main idea is to build a vector for S. coelicolor so that it can produce increasing amounts of methyl salicylate (MeSA) according to different nickel levels in the soil when inoculated with the biobeads described in the Week 1 homework.
In plants, MeSA is synthesized from salicylic acid (SA) by the enzyme SAMT, which bacteria do not have. Therefore, the SAMT gene in the vector was taken from Clarkia breweri, because its SAMT is highly selective for SA (Ross et al., 1999; Ward et al., 2022). In addition, although some bacteria can produce SA (Bakker et al., 2014), Streptomyces can not, so the gene for salicylate synthase (Irp9) was obtained from Yersinia enterocolitica (Kerbarh et al., 2005). NCBI was used to retrieve the sequences, and they were optimized for Streptomyces coelicolor using vectorbuilder.


One of the key features of NiMap-S from week 1 HW is increasing MeSA production in an analog manner, so a graded transcriptional response is required. This is why NmtR was selected (Busenlehner et al., 2003). For the protein itself, I had chosen another Streptomyces in Part 3, but I was not confident about its promoter. Therefore, I searched for a Streptomyces species with a divergent operon showing NmtR regulated expression and selected Streptomyces scabiei (Osman & Cavet, 2010). To place two separate expression cassettes, I included 200 bp from the ATG start sites of both oppositely oriented genes, covering the intergenic region, to avoid missing any promoter elements.
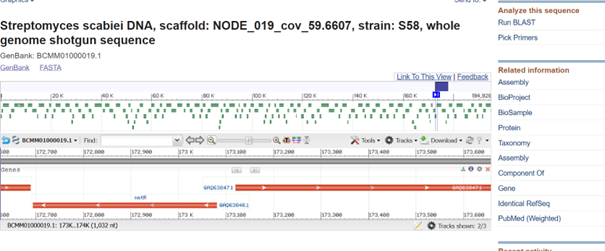

However, I have some concerns because the intergenic region in this divergent operon is very short, so the promoter regions overlap with the complementary sequences of the opposing CDS, which could lead to unintended expression from the opposite strand. Perhaps the intergenic region alone would be sufficient, but since NmtR is a repressor that binds as a homodimer and sigma factors will also bind in this region, it seemed too short. If I continue developing this project, I will investigate this further and may look for alternative promoters that provide more reliable, nickel dependent graded expression.
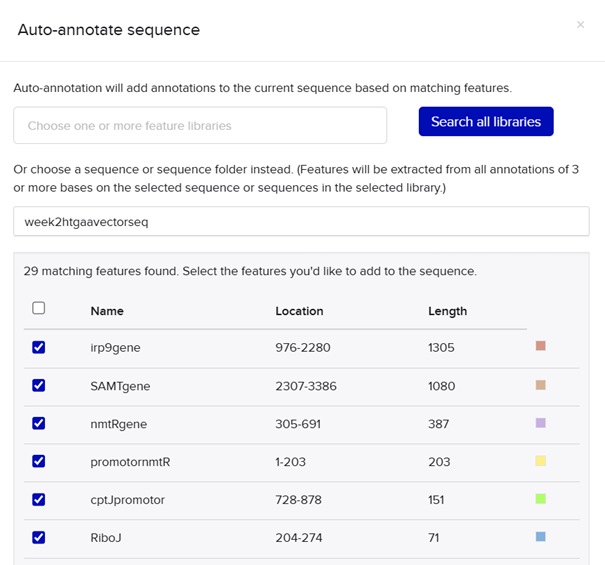
My RBS regions and RiboJ sequences were also chosen based on Bai et al. (2015). I used the R5, R7, and R9 RBS sequences from their supplementary data for nmtR, irp9, and SAMT, respectively, from weakest to strongest (Bai et al., 2015).
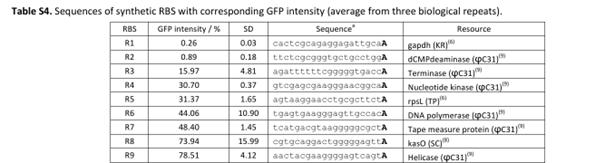

RBS sequences of varying strengths from Bai et al. (2015).
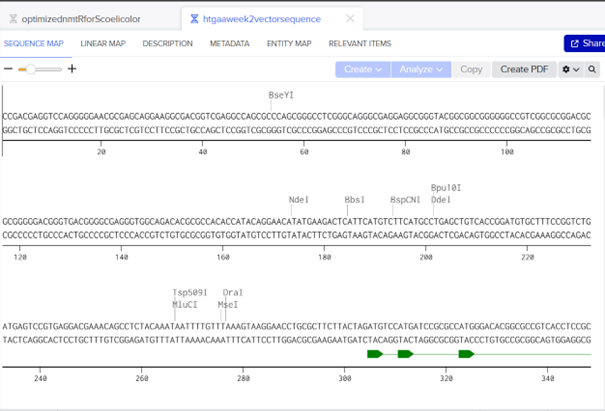

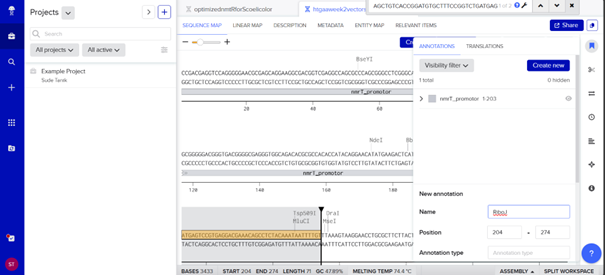
The link for my vector sequence with annotation is: https://benchling.com/s/seq-TgTZy6Lrs9s6JUGVv6Nz?m=slm-5ZdxPGhl1fFTWfiMyRpw







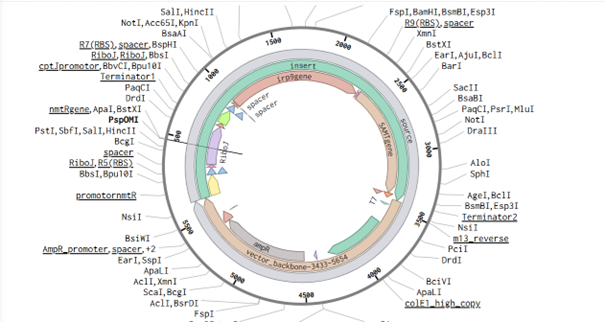

References
Ross, J. R., Nam, K. H., D’Auria, J. C., & Pichersky, E. (1999). S-Adenosyl-L methionine:salicylic acid carboxyl methyltransferase, an enzyme involved in floral scent production and plant defense. Archives of Biochemistry and Biophysics
Ward, L. C., McCue, H. V., & Carnell, A. J. (2020). Carboxyl methyltransferases: Natural functions and potential applications in industrial biotechnology. ChemCatChem
Bakker, P. A. H. M., Ran, L., & Mercado-Blanco, J. (2014). Rhizobacterial salicylate production provokes headaches! Plant and Soil
Kerbarh, O., Ciulli, A., Howard, N. I., & Abell, C. (2005). Salicylate biosynthesis: Overexpression, purification, and characterization of Irp9, a bifunctional salicylate synthase from Yersinia enterocolitica. Journal of Bacteriology
Kim, H. M., Ahn, B.-E., Lee, J.-H., & Roe, J.-H. (2015). Regulation of a nickel/cobalt efflux system and nickel homeostasis in Streptomyces coelicolor. Metallomics.
Busenlehner, L. S., Pennella, M. A., & Giedroc, D. P. (2003). The SmtB/ArsR family of metalloregulatory transcriptional repressors: structural insights into prokaryotic metal resistance. FEMS Microbiology Reviews
Bai, C., Zhang, Y., Zhao, X., Hu, Y., Xiang, S., Miao, J., Lou, C., & Zhang, L. (2015). Exploiting a precise design of universal synthetic modular regulatory elements to unlock microbial natural products in Streptomyces. Proceedings of the National Academy of Sciences (PNAS)