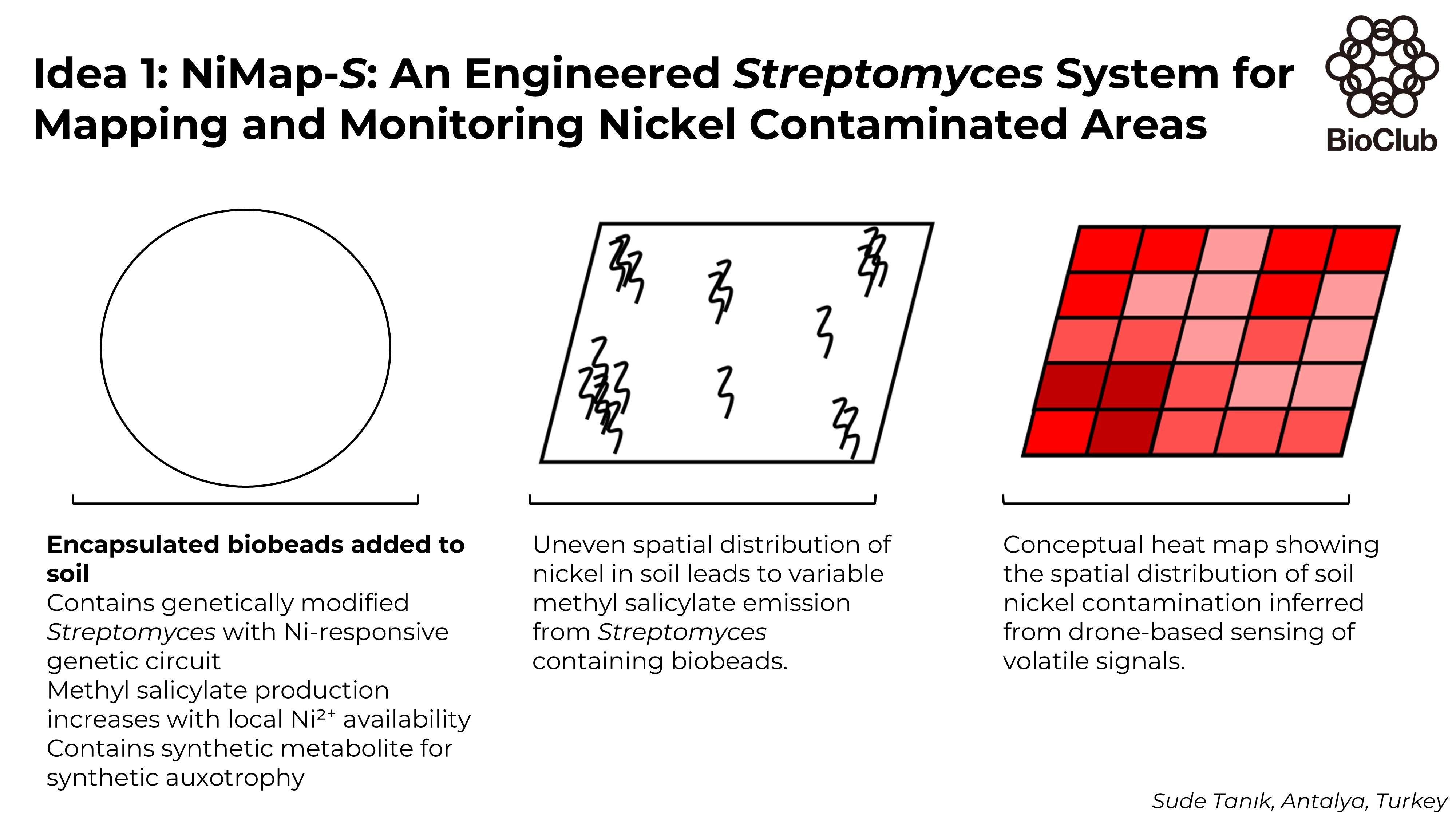
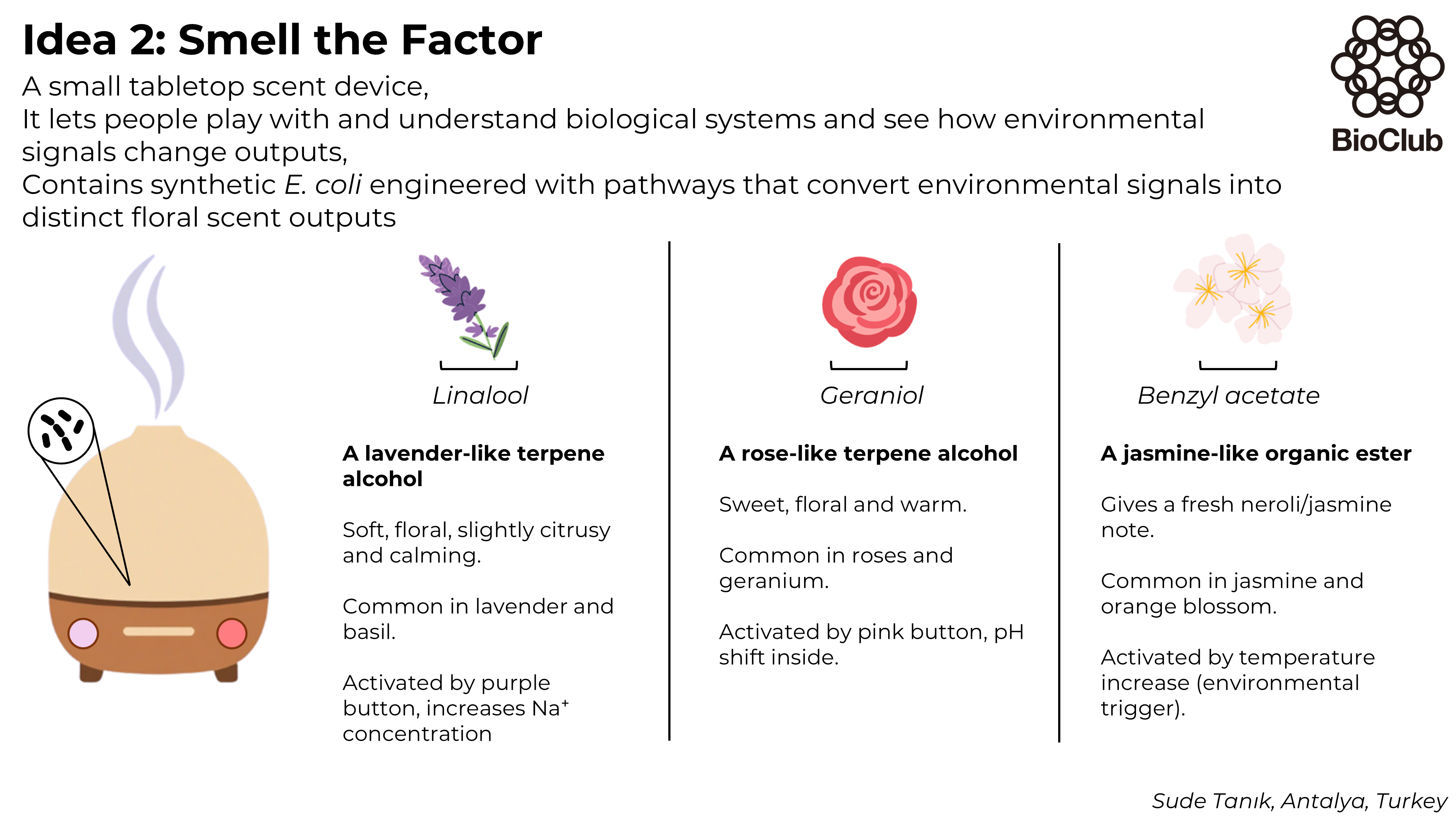
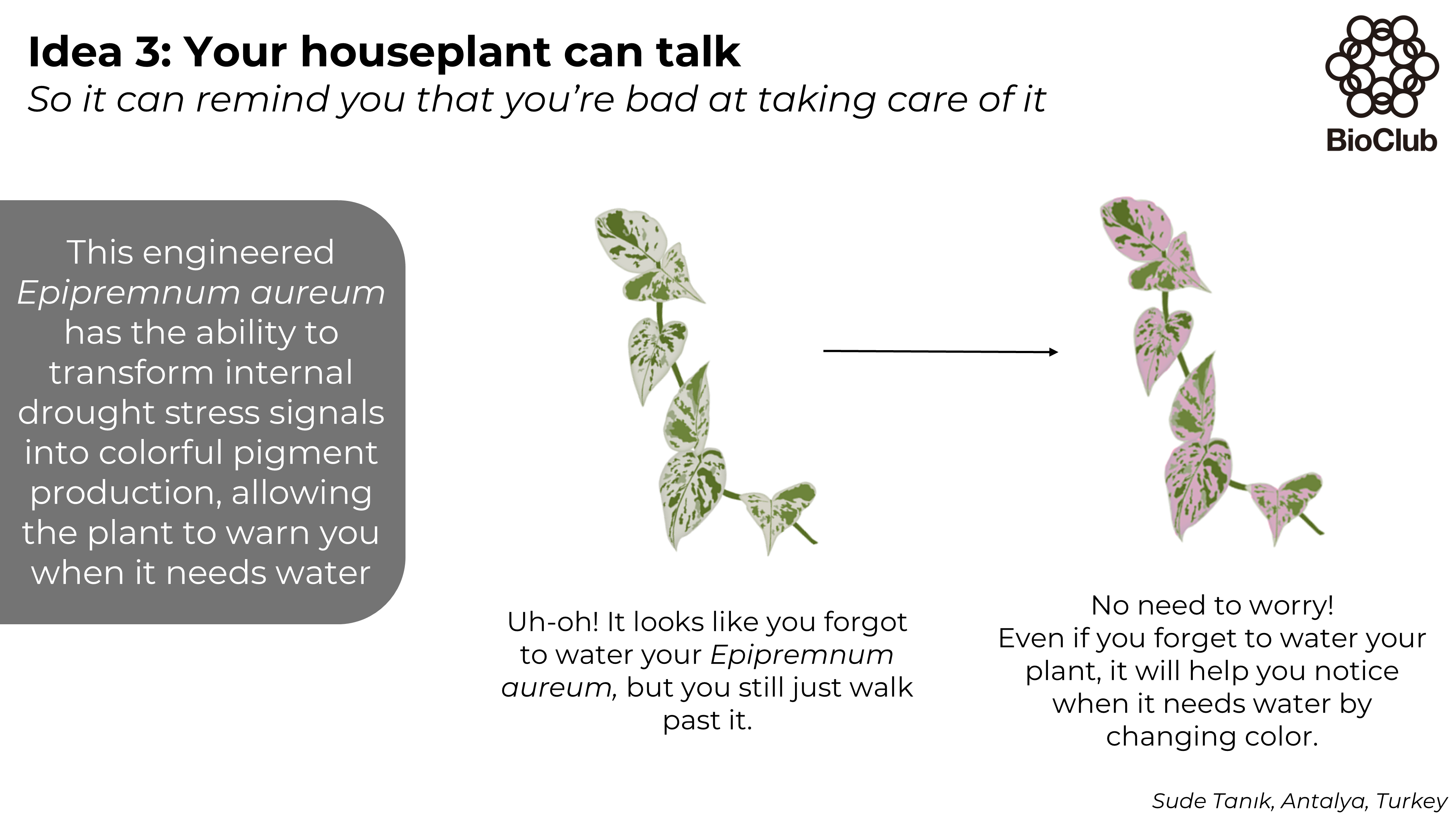
Week 3 HW: Lab Automation
Part 1: Python Script for Opentrons Artwork
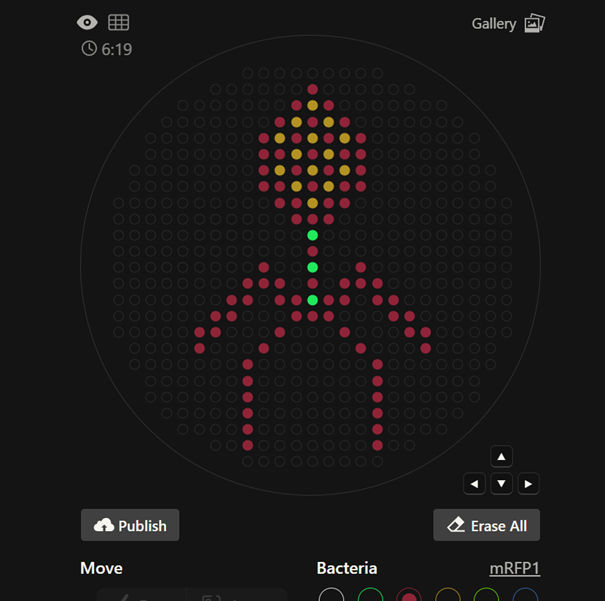

First, I created my design on the website and drew a bacteriophage
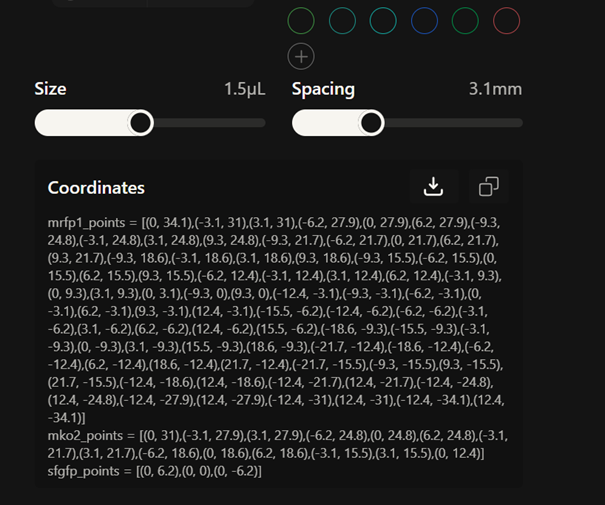

Then, I copied the coordinates of my design
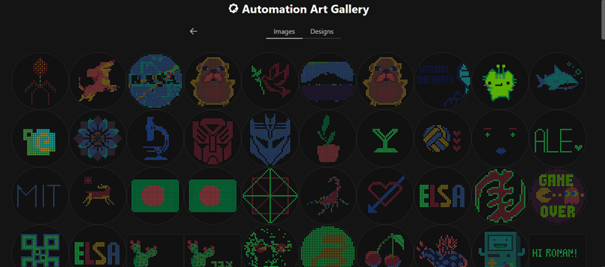

I also published my design to the gallery.

After that, I went to colab

I sent my coordinates to ChatGPT 5.2 and asked it to generate code that would run my desired design
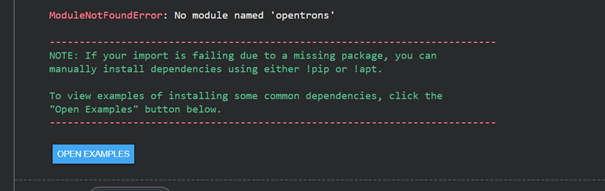

When I first tried to run it, I received an error

Then I executed the “Run this block once per runtime to set up your environment” section
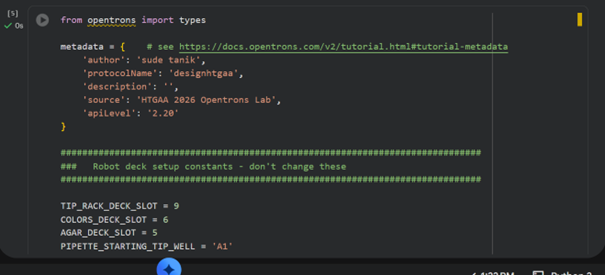

After that, I ran my code again
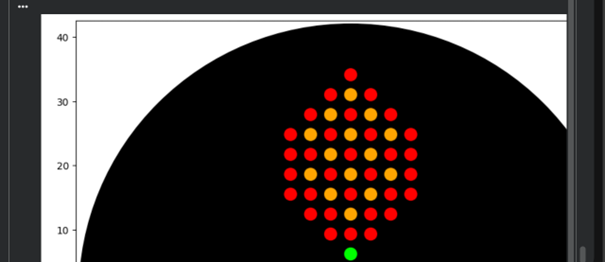

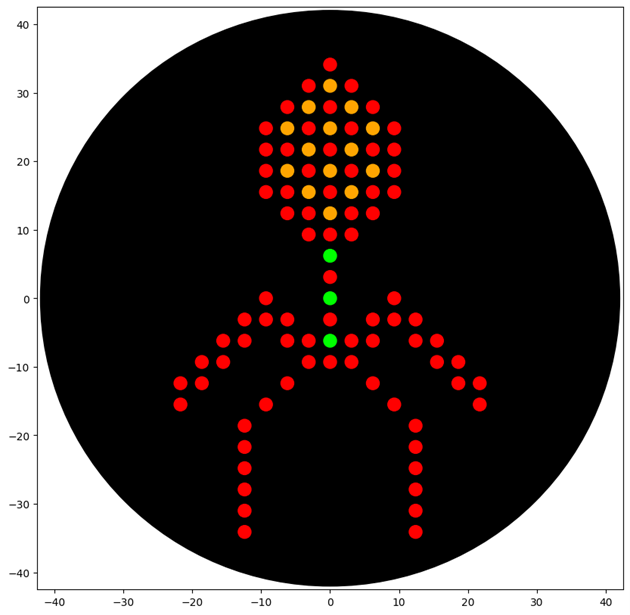

Finally, my designed shape appeared successfully
Access for code: https://colab.research.google.com/drive/1ppqogS0fGyVIS72rR_hJCog9bBsVNodc?usp=sharing
Part 2: Post-Lab Questions
1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications

 DOI: https://doi.org/10.1002/edn3.70074
DOI: https://doi.org/10.1002/edn3.70074
Rice’s whale (Balaenoptera ricei) is an endangered whale species living in the Gulf of Mexico. Because it is very rare and difficult to observe directly researchers in this article used environmental DNA (eDNA) from seawater samples to detect its presence. Since whales could occasionally be visually observed during surveys, these encounters were used to validate the eDNA detection method by collecting seawater samples shortly after a whale surfaced. Seawater samples were first filtered using different filter pore sizes to capture DNA fragments released by the whale. DNA was then extracted from the filters and analyzed using qPCR assay targeting a mitochondrial DNA region.
The assay was validated through in silico, in vitro, and in situ testing: computational analyses were used to ensure primer specificity, laboratory experiments tested the assay against whale DNA and non-target species, and environmental seawater samples collected in the field were used to confirm that the method could successfully detect Rice’s whale eDNA. An Opentrons OT-2 was used to automate the preparation of qPCR plates by pipetting reagents, standards etc. before the analysis was performed with a qPCR device.
2. How would I utilize opetrons for the final project
To test the feasibility of NiMap-S from the week 1 HW, I would first engineer E. coli with my nickel-responsive promoter before moving into Streptomyces. The promoter from cassette 2 (from week 2 HW) would drive GFP expression instead of the methyl salicylate (MeSA) pathway enzymes. Cassette 1 from week 2 HW would remain unchanged, expressing the regulatory protein nmrT to regulate the expression of cassette 2 depending on different Ni levels.
By measuring GFP fluorescence, I could evaluate two things: whether the nickel-responsive promoter is functioning at all (GFP vs. no GFP), and whether GFP expression changes quantitatively in response to different nickel concentrations. These preliminary experiments are important before transitioning into a more complex organism and a more complex genetic circuit. This allows me to verify that the promoter and sensing module (which uses nmrT as the regulatory protein for gradual Ni response) behave as expected before adding metabolic burden from the MeSA biosynthetic pathway. This strategy helps isolate potential problems early. If something fails, by implementing this preliminary research I can determine whether the issue lies in the promoter system or in the metabolic pathway.
So, I would use Opentrons robot in this experiment to prepare different nickel concentrations in a 96-well plate. The robot would create a nickel gradient using serial dilutions, add the engineered E. coli to each well, and make sure that all volumes are consistent. I would measure GFP fluorescence to see how the promoter responds to increasing nickel levels. This would allow me to generate a curve and understand whether the promoter response is gradual based on increasing nickel levels or switch like.
If the desired GFP expression patterns are observed in E. coli, I would then replace GFP with the methyl salicylate biosynthesis enzymes and test whether volatile production behaves as expected (I do not know if opentrons can be used to detect volatile signals). If that stage is successful, the validated circuit could then be transferred into Streptomyces for further development, since Streptomyces is a more complex organism to work with compared to E. coli. This gradual progression ensures that each layer of the system is validated before moving to the next level of biological complexity.
Final Project Ideas