Week 2 HW: DNA Read, Write and Edit
Part 0: Basics of Gel Electrophoresis
#S.S: 부트캠프와 Wk2 수업을 들은 후, 과제를 확인했지만 정확한 개념 정리가 안되어 있어서 뭘 진행하는지 이해하기 어려웠다. 따라서 ChatGPT를 활용, it became clearer once I understood the concept of restriction enzyme(molecular scissors) and gel electrophoresis(DNA size filter).
Prompt: "restriction enzyme biology 와 gel electrophoresis에 대해서 아주 간단히 설명해줘"
를 통해 다음과 같은 답변을 얻었다.
1. Restriction enzyme (제한효소) biology
제한효소는 특정한 DNA 염기서열을 인식해서 그 지점을 자르는 효소입니다.
원래는 박테리아의 방어 시스템으로, 침입한 바이러스 DNA를 잘라 파괴합니다.
각 효소는 고유한 인식 서열(예: GAATTC)을 가지며, 그 위치에서 DNA를 정확히 절단합니다.
이 특성 덕분에 분자생물학에서는 DNA를 원하는 조각으로 자르는 도구로 사용됩니다.
2. Gel electrophoresis (겔 전기영동)
겔 전기영동은 DNA 조각을 크기별로 분리하는 실험 방법입니다.
DNA는 음전하를 띠므로 전기를 걸면 양극 방향으로 이동합니다.
작은 DNA 조각일수록 겔 속을 더 빨리, 더 멀리 이동합니다.
결과적으로 DNA 조각들이 밴드 형태로 나타나며, 크기를 비교·확인할 수 있습니다.
One-line summary
-Restriction enzymes: molecular scissors that cut DNA at specific sequences
-Gel electrophoresis: a method to separate DNA fragments by size
Part 1: Benchling & In-silico Gel Art
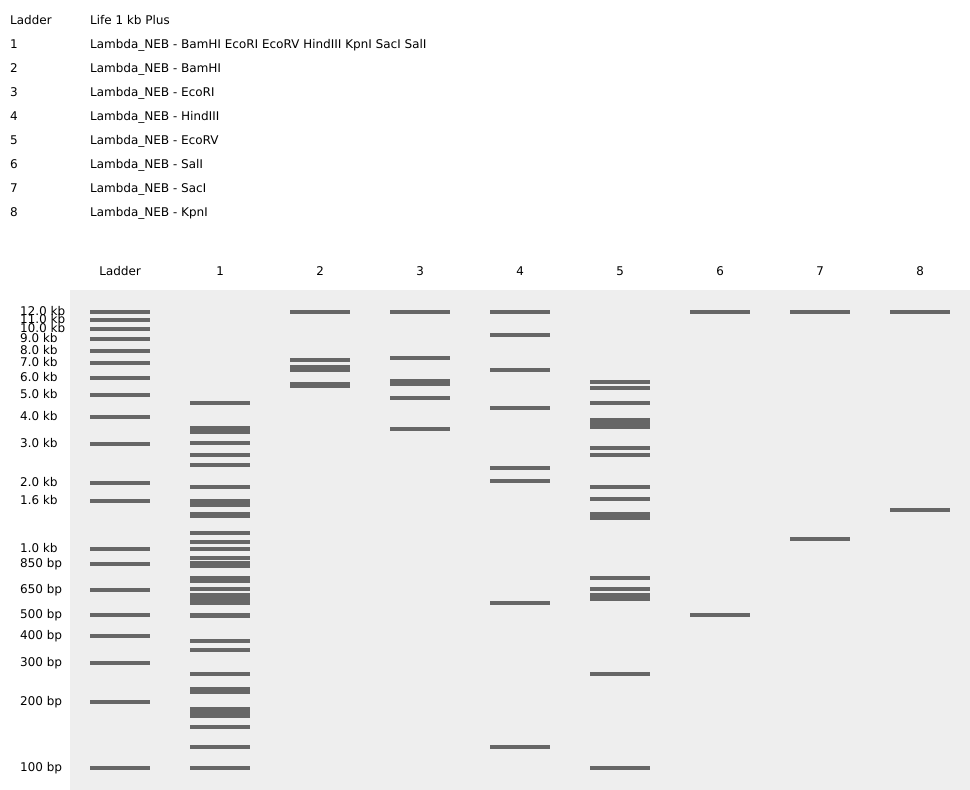


#S.S: Benchling is such a cool platform that we can edit DNA in-silico. I uploaded ‘Lambda DNA.txt’ and managed digest by choosing each enzyme that cuts DNA strand. My initial thought was to see the ladder that each enzyme makes to decide which figure/shape I’d like on my artwork, but the shown result were not a single bar but of multiple layers. — at this stage, I have no idea how to compose an gel art.
===
#S.S: Little update on my Gel Art, after attending a session from Designer Cells lab in Songdo, S. Korea.
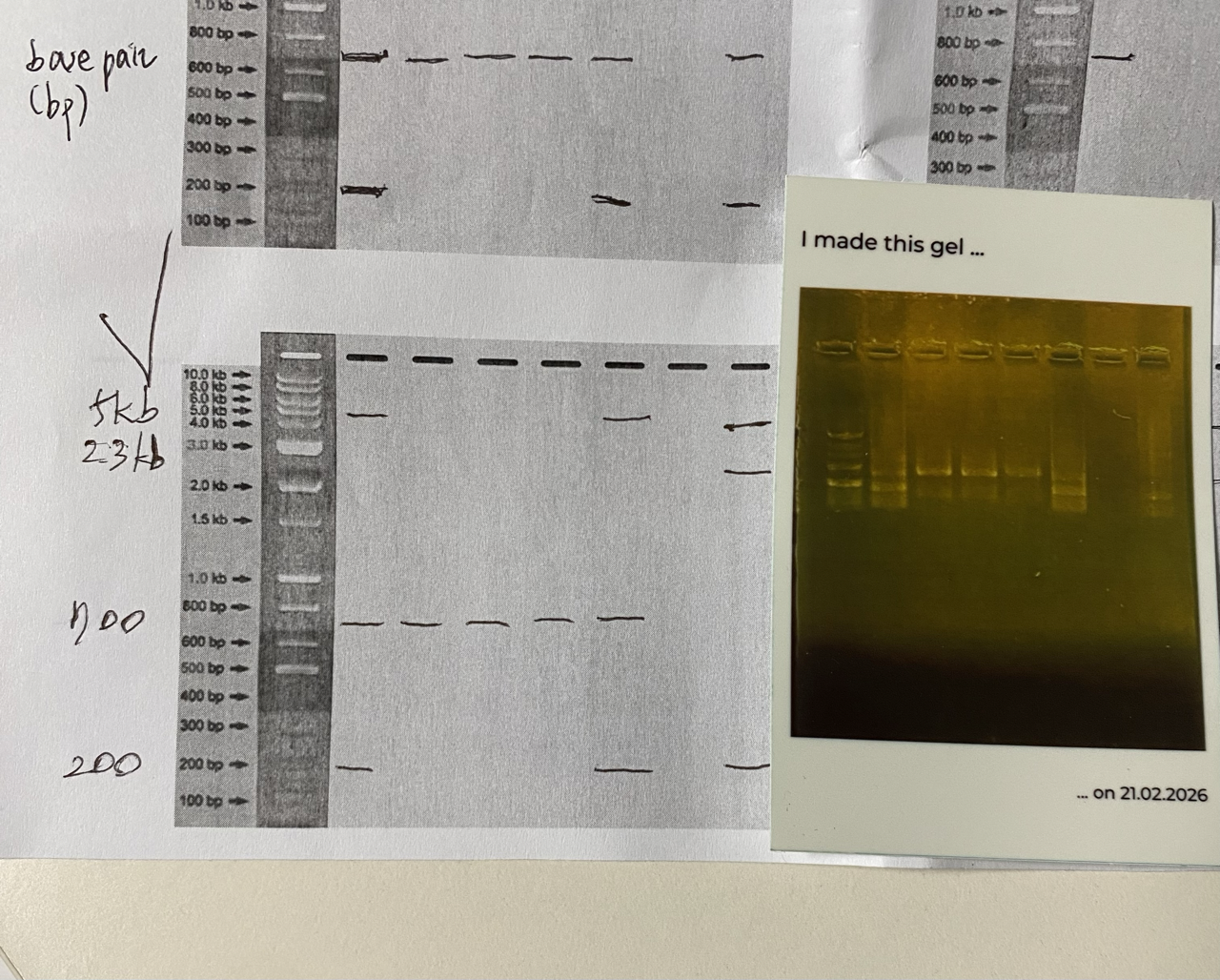
Ana, one of our TAs, printed out the Gel Electrophoresis result for us, using an instant photo printer. The photo on the right bottom. It also is a sticker !!!

So, my first ever wetlab experience started with how to use pipettes. Next, we moved on to the main dish, how to perform gel electrophoresis.
- mix TAE and agarose(powder) and the solution would be microwaved until it is clear, which means it is completely dissolved.
- I missed the reasons- to add 1uL of SYBR Safe DNA stain, and ChatGPT explains as below:


- Place the comb(or we can put them later-it depends your way of doing it-just avoid forming bubbles) and pour the agarose gel.
- When the gel is solidified enough, remove the comb(which generated wells) and start loading the wells with your DNA stands.
There were 4 different sized DNA strands (200bp, 700bp, 2.3kb, 5kb) ready for us, thanks to one of our TAs, Hyunseo.
I designed my gel art saying “Hi”, but the result is shown rather like “HI”, which still, luckily conveys the intended meaning ;D Hyunseo guessed 5kb and 2.3kb might gone bad? and failed to show on our Gel.
I am presenting my artwork again below just to show off.

The most challenging part of wet lab was actually doing the math and figuring out volumn units. I would refer the summary from ChatGPT for my future references.
Basic Units and Calculations in Biology
1️⃣ Volume Units (Metric System)
All volume units are based on powers of 10.
- 1 L (liter) = base unit
- 1 mL = 10⁻³ L
- 1 µL = 10⁻⁶ L
- 1 nL = 10⁻⁹ L
Key rule: Each step (milli → micro → nano) differs by 1000×.
Examples:
- 1 mL = 1000 µL
- 1 µL = 1000 nL
2️⃣ Mole (mol)
A mole measures the amount of substance.
1 mol = 6.022 × 10²³ molecules (Avogadro’s number)
It is simply a very large counting unit for atoms or molecules.
3️⃣ Molarity (M)
Molarity is concentration.
[ M = \frac{\text{moles}}{\text{liters}} ]
Example: 1 M solution = 1 mole dissolved in 1 liter.
Important:
- mol = amount
- M = concentration
4️⃣ Dilution Formula
[ C_1 V_1 = C_2 V_2 ]
Used to prepare diluted solutions from a concentrated stock.
Example: To make 100 mL of 1 M solution from 10 M stock:
[ (10M)V_1 = (1M)(100mL) ]
V₁ = 10 mL
5️⃣ Key Takeaways
- milli, micro, nano differ by 1000× each step
- M means mol/L
- Most lab work uses µL volumes
- DNA amounts are often measured in ng (10⁻⁹ g)
Part 3: DNA Design Challenge
#S.S: now, this is going to be a fun exercise!
3.1. Choose your protein : Fibronectin type III domain-containing protein 10
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
'>sp|F2Z333|FND10_HUMAN Fibronectin type III domain-containing protein 10 OS=Homo sapiens OX=9606 GN=FNDC10 PE=1 SV=1 MRAPPLLLLLAACAPPPCAAAAPTPPGWEPTPDAPWCPYKVLPEGPEAGGGRLCFRSPARGFRCQAPGCVLHAPAGRSLRASVLRNRSVLLQWRLAPAAARRVRAFALNCSWRGAYTRFPCERVLLGASCRDYLLPDVHDSVLYRLCLQPLPLRAGPAAAAPETPEPAECVEFTAEPAGMQDIVVAMTAVGGSICVMLVVICLLVAYITENLMRPALARPGLRRHP'
'>reverse translation of sp|F2Z333|FND10_HUMAN Fibronectin type III domain-containing protein 10 OS=Homo sapiens OX=9606 GN=FNDC10 PE=1 SV=1 to a 678 base sequence of consensus codons.
atgmgngcnccnccnytnytnytnytnytngcngcntgygcnccnccnccntgygcngcn
gcngcnccnacnccnccnggntgggarccnacnccngaygcnccntggtgyccntayaar
gtnytnccngarggnccngargcnggnggnggnmgnytntgyttymgnwsnccngcnmgn
ggnttymgntgycargcnccnggntgygtnytncaygcnccngcnggnmgnwsnytnmgn
gcnwsngtnytnmgnaaymgnwsngtnytnytncartggmgnytngcnccngcngcngcn
mgnmgngtnmgngcnttygcnytnaaytgywsntggmgnggngcntayacnmgnttyccn
tgygarmgngtnytnytnggngcnwsntgymgngaytayytnytnccngaygtncaygay
wsngtnytntaymgnytntgyytncarccnytnccnytnmgngcnggnccngcngcngcn
gcnccngaracnccngarccngcngartgygtngarttyacngcngarccngcnggnatg
cargayathgtngtngcnatgacngcngtnggnggnwsnathtgygtnatgytngtngtn
athtgyytnytngtngcntayathacngaraayytnatgmgnccngcnytngcnmgnccn
ggnytnmgnmgncayccn'
3.3. Codon optimization.
The purpose of codon optimisation is to improve expression efficiency. It would not alter the encoded protein but it will redesign the DNA sequence to match a host organism.
'>Improved DNA:
ATGGGCCCCCTGTTCGCCCTGGCCCCCCCCGGCGCCCGCCCCCCCCCCGT
GGGCCCCCCCGAGCCCTGGTGCCTGAAGTTCCGCGGCCGCGCCGGCGGCC
TGCTGCCCCGCGTGGTGCAGCCCGGCGGCAGCGCCCGCGGCTGGCGCTAA
CGCTTCCACGGCTGCCCCCGCGCCGGCGGCCTGCTGATGTGGGGCGCCAA
CGTGCCCGGCGGCCTGGGCTGGGACATCCCCAGCCAGAGCCTGGTGGTGA
CCAGCTGGCGCCCCCGCGCCCCCGAGCCCGACCGCGACGGCGACTACGCC
ACCGCCGACGCCGAGTGCGTGCACGACGCCTGGGGCCTGGTGTGCGTGTA
CTGCTGCGCCAACACCAAGTACGGCCGCTGCGCCGGCGGCACC'
{ref.} https://www.jcat.de/ --- codon optimisation tool, free online access
3.4. You have a sequence! Now what?
Using plasmid? The DNA could be used to produce protein.
3.5. How does it work in nature/biological systems?
#S.S(3.1.): Firstly, I had to recognise the differences between ‘gene vs. protein’. From my understanding, the central dogma goes like this : DNA(gene) -> RNA(transcription) -> Protein(translation) The problem is I am not familiar with names of proteins, thus come up with a gene that I’d encountered in the past! – such as, PCSK9
* PCSK9 - proprotein convertase subtilisin/kexin type 9
{ref. NCBI} https://www.ncbi.nlm.nih.gov/datasets/gene/255738/#transcripts-and-proteins
There are PCSK9 inhibitors in the market that are precsribed to regulated high LDL-C level of Dyslipidemia or FH(Familial Hyperlipidemia). But, I wasn’t happy with the NCBI lists of protein accession (NP_777596.2, etc.) since I don’t understand what they mean. In UniProtKB, however, I came across an interesting name, ‘Adnectin’ - I was curious, why does it have a special name on it? It is a protein therapeutics, according to below resources, and naturally I decided to look into Human Fibronectin deeper.
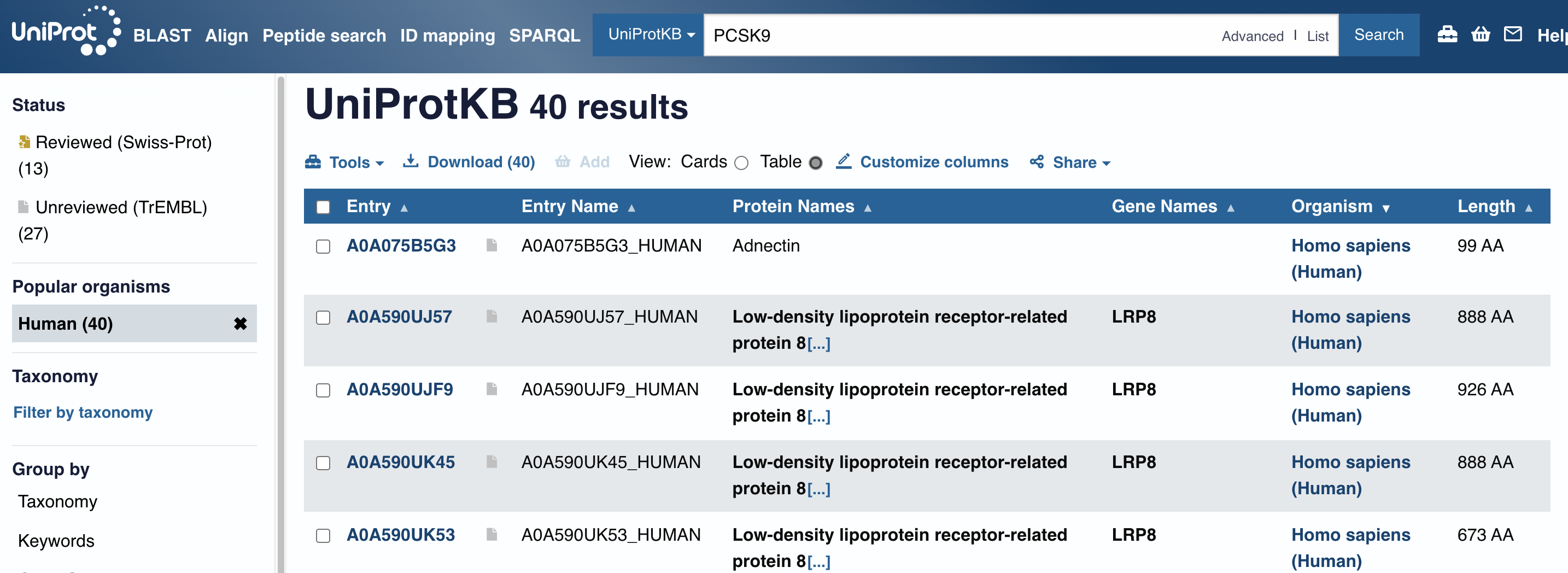
 {UniProt}
{UniProt}
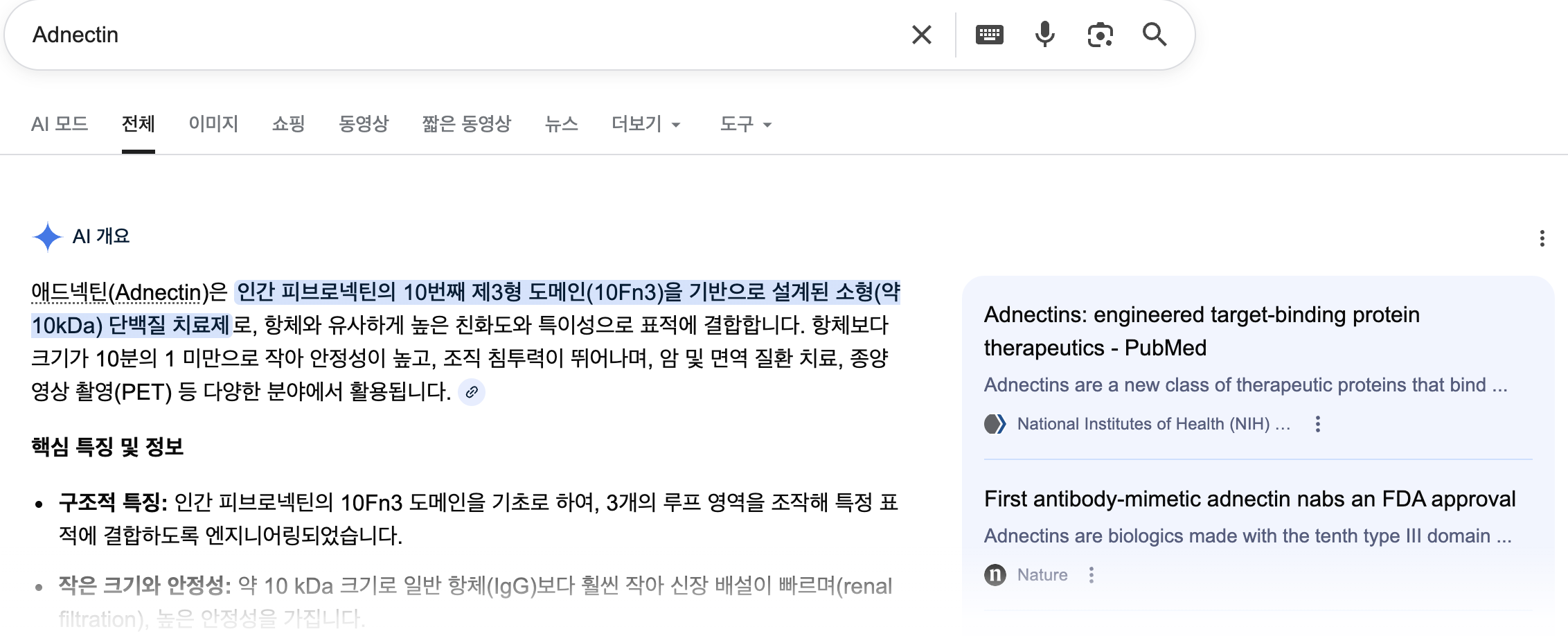
 {google}
{google}

 {PubMed. NCBI}
{PubMed. NCBI}
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank). I would like to sequence my own DNA, and especially check on the currently available cancer gene test.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Also answer the following questions:
Is your method first-, second- or third-generation or other? How so?
NGS(second) method would fit for my own DNA read, since it's fast enough and relatively cheaper that the third-gen method.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
I remember writing DNA of a bateriophage to used it as a therapeutic.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
Also answer the following questions:
- What are the essential steps of your chosen sequencing methods?
- What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
I'm more interested in utilizing RNA than editing DNA form.
(ii) What technology or technologies would you use to perform these DNA edits and why?
Also answer the following questions:
- How does your technology of choice edit DNA? What are the essential steps?
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?