Week 3 HW: Lab Automation
A group HW: Design cell-free RNA biosensors
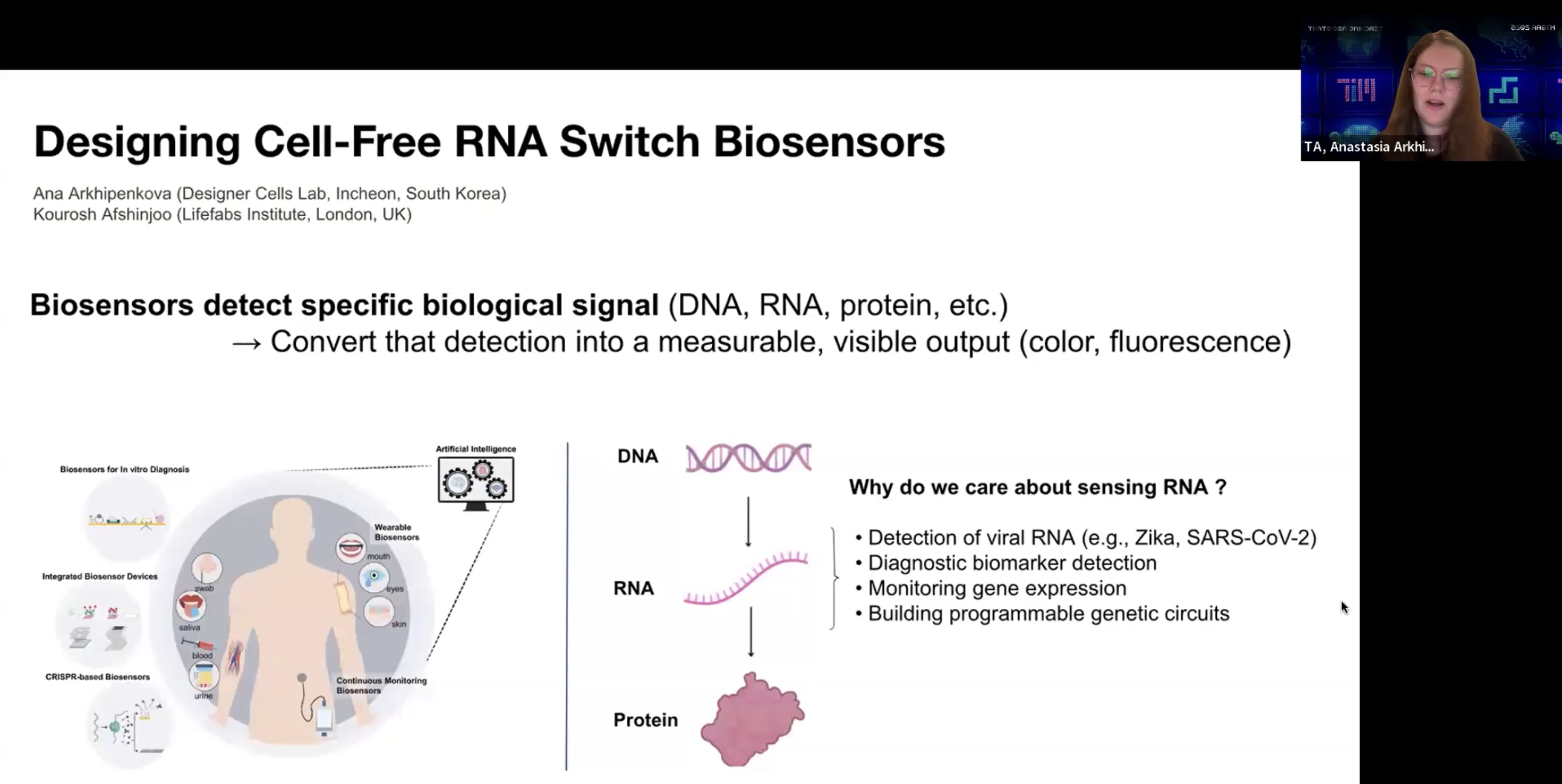
 RNA toehold switch가 작동하는 원리를 recitation에서 Ana가 설명해 주었는데,
나는 우선 mRNA가 단백질로 translation되는 기전을 이해하고나서 이런 biosensor의 기능을 이해할 수 있었다.
RNA toehold switch가 작동하는 원리를 recitation에서 Ana가 설명해 주었는데,
나는 우선 mRNA가 단백질로 translation되는 기전을 이해하고나서 이런 biosensor의 기능을 이해할 수 있었다.
mRNA에 ribosome이 와서 결합한다. 찾아다닌다기 보다는 cell내에서 자유롭게 떠다니던 ribosome이 충돌하여 결합하는 것에 가깝다고. Eukaryotes의 경우는 rough endoplasmic reticulum(RER, 세포소기관organelle)에 거칠어 보이도록 ribosome들이 붙어있어서 단백질을 생산, 반면에 Prokaryotes, 즉, bacteria의 경우 ribosome은 free-floating 상태로만 cytoplasm(세포질)에 존재.
- 요새 기본개념 학습에 도움을 받고 있는 biology podcast(https://podcasts.apple.com/kr/podcast/teach-me-biology/id1525760514?l=en-GB&i=1000486622621)의 기억을 떠올리면, Eukaryotes/animal cell에서 RER은 protein을 생산하고, smooth ER(w/o ribosome)은 lipid를 생산한다. free ribosome은 세포 내부에서 사용할 단백질을 생산한다고 한다. 추가로, mitochondria는 release energy & produce ATP. 그리고 risosome은 pathogen이나 dead cell, etc를 없애는 역할을 하는 것 같다.
mRNA에는 RBS(ribosome binding site)가 있어서 ribosome이 부착되고 번역을 시작할 수 있다. 이 RBS는 bacteria에서는 Shine–Dalgarno sequence라고 불린다고. 시작코돈 AUG 앞에 위치한다.
하나 더 몰랐던 개념, RNA는 folding landscape, 동적으로 접혔다 풀렸다 하는 구조집합을 보이는데, toehold switch에 나온 hairpin 구조는 rna 염기 가닥이 서로 연결되지 않은 loop, 그리고 일부가 상보적으로 연결된 stem으로 구성된 loop-stem 구조이다. 모든 RNA가 이렇게 모양이 생긴건 아니고 염기서열에 따라서 동적인듯 하다. 그렇지만 박테리아는 비교적 mRNA 구조가 단순하므로 의도적으로 hairpin구조로 만들어서 활용하는 듯 하다.
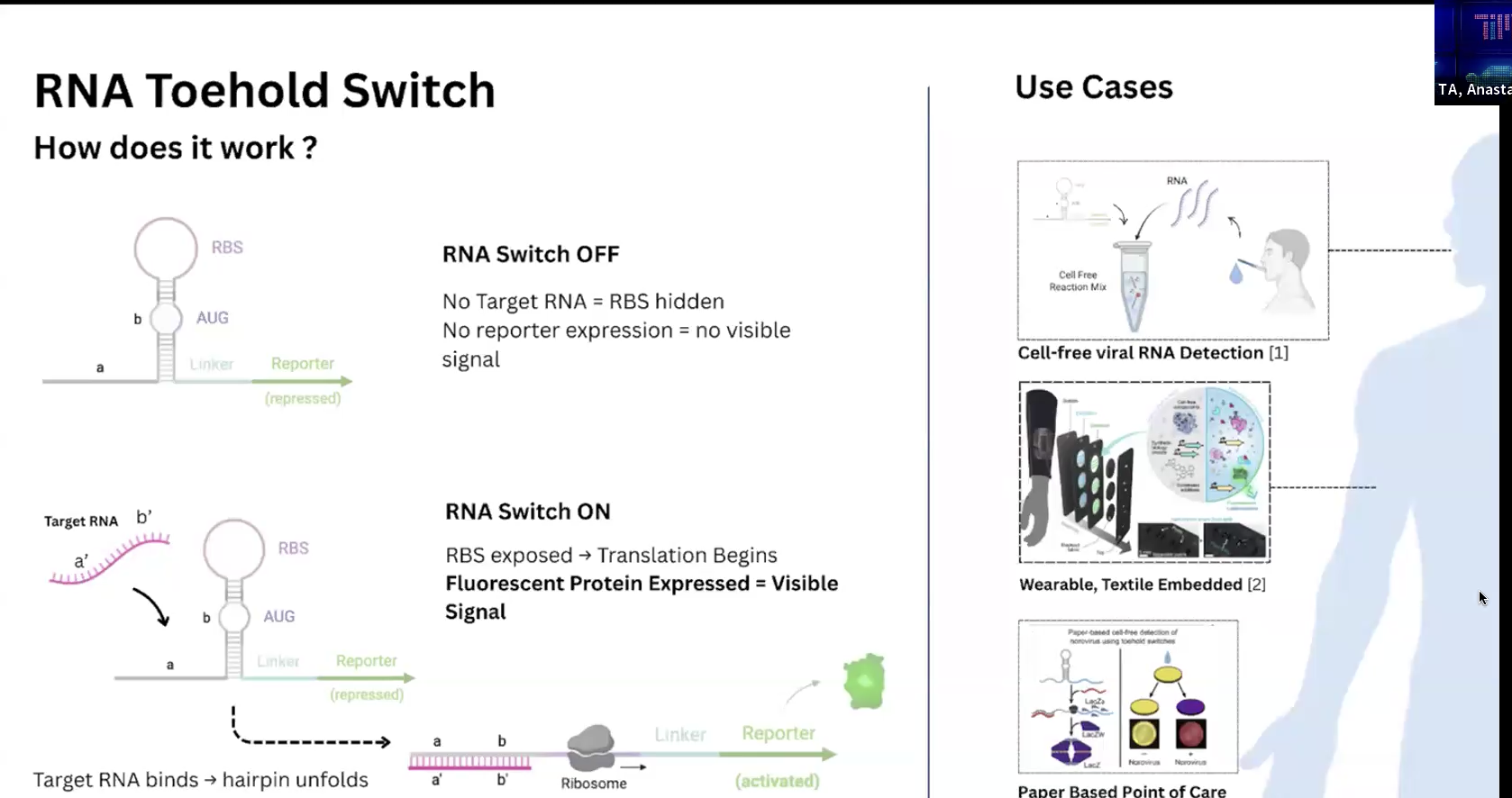
Finally, toehold switch에서 hairpin구조의 RNA switch는 평소(off status) RBS(is hidden within stem)와 AUG를 가림으로써 ribosome의 접근을 막아서 protein translation을 막는다. When switched-on(target RNA가 접근 시, it will bind and zip-open the stem followed by strand displacement(헤어핀 붕괴), in the end, ribosome can start translation process.
약간 이해가 안되었던 부분은 RNA가 지금 2종이 나온다는 거였는데, chatGPT에게 질문하려고 개념을 써보다 보니 얼추 아래와 같이 이해가 되었다.
toehold switch는 bacteria의 RNA가 표현되기를 hairpin 구조를 취하도록 re-engineered it. hairpin 상태에 있다가, target RNA(외부의 phage같은 것들의 input 등)가 들어오면 protein을 만들도록 설계.
ChapGPT의 도움을 받아 재정리해보면, Bacterial mRNA의 5’UTR (AUG가 붙은 앞쪽 부분으로 예상.)을 hairpin 구조로 합성한 것! (자연적으로 형성된 switch로 Riboswitch라는게 존재함)

#s.s: At this stage, I wondered what it means to be ‘cell-free’ and why it matters. Below is an examplary table from ChatGPT to help my understanding. It seems cell-free system could partially avoid GMO-regulatory risks or make things easier! (Which helps me narrow down the desired setting for my final project, i guess)
| Category (항목) | Cell-based (세포 기반) | Cell-free (세포 비의존) |
|---|---|---|
| Viability (생존) | Required (필요) | Not required (불필요) |
| Proliferation (증식) | Present (있음) | Absent (없음) |
| Regulation (규제) | GMO-regulated (GMO 규제) | Relatively relaxed (상대적 완화) |
| Response Speed (반응 속도) | Slow (느림) | Fast (빠름) |
| Circuit Complexity (회로 복잡성) | High (높음) | Low (낮음) |
| Field Deployability (현장 적용성) | Low (낮음) | High (높음) |
Brainstorming on RNA Biosensor, assisted by ChatGPT
- Target AMR(Antimicrobial Resistance) gene
Development of resistance for antibiotics seemed like a major issue in the medical field. It is routinely tested in hospital already with AST (antimicrobial susceptibility testing). Also, molecular testing for AMR gene is performed as an early guidance. Thus, RNA biosensor have potential to be the fastest detection tool replacing the current molecular test in urgent scenarios, although it might lack precision compared to AST.
- Microneedle patch for Vaccination
Could a biosensor detect humoral immunity after vaccine innoculation?
It is possible to check interstitial fluid (ISF) on one’s skin, but immunity is proved mostly by protein(as an antibody) and RNA sensor is not the best option to detect a vaccination result… so, I would dump this idea.
Virus RNA biosensor
Back to the familiar idea that I was introduced from the class, which is the detection of RNA viruses. RNA is THE genetic material of a pathogene and the sensor have potential to complement or partially replace PCR. In addition, it would have significant societal value - high impact on public health.
So, I’m thinking “broad coronavirus biosensor” as a front-line detector.
#GPT: “Detecting viral RNA directly is important because The presence of viral RNA typically indicates an active infection, rather than past exposure or residual immunity.
In regarding Coronaviruses, related viruses such as SARS-CoV (2003) and MERS-CoV (2012) had already caused outbreaks and the coronavirus family was already known. However, SARS-CoV-2(COVID-19) was a newly emerged zoonotic virus that humans had not previously encountered at population scale. So COVID-19 was not created “from nothing,” but it represented a new spillover event of a genetically distinct virus within a known viral family.
Coronaviruses belong to the family Coronaviridae.
Within it are multiple genera:
- Alphacoronavirus
- Betacoronavirus (includes SARS-CoV, MERS-CoV, SARS-CoV-2)
If you design a biosensor targeting highly conserved regions (e.g., RdRp gene)
You could potentially detect:
- SARS-like coronaviruses
- MERS-like coronaviruses
- Related zoonotic betacoronaviruses
This is because the RNA-dependent RNA polymerase (RdRp) region is relatively conserved across coronaviruses.
So in principle:
A conserved-region RNA biosensor could detect the presence of a coronavirus even before it is classified as a new human pathogen.
Would it detect a completely new zoonotic spillover?
Scenario 1: Spillover from a closely related SARS-like bat CoV
- Likely detectable with a conserved-region sensor.
Scenario 2: Spillover from a distantly related coronavirus lineage
- Detection depends on how conserved the target region is.
Scenario 3: Spillover from a completely different viral family
- ❌ A coronavirus-targeted sensor would not detect it.
Technical Considerations to design a broad CoV biosensor,
Multi-sequence alignment across:
- Bat SARS-like CoVs
- Pangolin CoVs
- Human SARS-CoV
- SARS-CoV-2
- MERS-CoV
Identification of:
- Highly conserved 25–35 nt regions
- Structurally accessible regions
- Minimal mutation hotspots
Possibly:
- Degenerate base tolerance in design
- Multiplexed switches covering multiple conserved motifs
— A biosensor targeting conserved coronavirus regions could potentially detect certain future zoonotic spillovers within the CoV family.
But:
It would not detect unrelated viral families.
It would not replace sequencing.
It must be part of an integrated surveillance system.